Differential Virulence of Clinical and
Bovine-Biased Enterohemorrhagic Escherichia coli O157:H7
Genotypes in Piglet and Dutch Belted Rabbit Models
The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters.Citation Shringi, S. et al. “Differential Virulence of Clinical and Bovine-Biased Enterohemorrhagic Escherichia Coli O157:H7 Genotypes in Piglet and Dutch Belted Rabbit Models.” Infection and Immunity 80.1 (2011): 369–380. © 2011 American Society for Microbiology
As Published http://dx.doi.org/10.1128/iai.05470-11
Publisher American Society for Microbiology
Version Author's final manuscript
Citable link http://hdl.handle.net/1721.1/78007
Terms of Use Creative Commons Attribution-Noncommercial-Share Alike 3.0
Differential virulence of clinical and bovine-biased enterohemorrahgic E. coli
1
O157:H7 genotypes in piglet and Dutch Belted rabbit models
2 3
Smriti Shringi1a, Alexis García2a, Kevin K. Lahmers1, Kathleen A. Potter1, Sureshkumar 4
Muthupalani2, Alton G. Swennes2, James G. Fox2, Carolyn J. Hovde3, Douglas R. Call1 5
and Thomas E. Besser1* 6
7
1
Department of Veterinary Microbiology and Pathology, Washington State University, 8
Pullman, WA 99164, USA 9
2
Division of Comparative Medicine, Massachusetts Institute of Technology, Cambridge, 10
MA 02139, USA 11
3
Department of Microbiology, Molecular Biology, and Biochemistry, University of 12
Idaho, Moscow, Idaho 83844, USA 13
a
Co-first author 14
15
Running title: Differential virulence of EHEC O157 genotypes
16 17
*
Corresponding Author: Thomas E. Besser
18
Mailing Address: Food and Waterborne Disease Research Program, Veterinary 19
Microbiology and Pathology, College of Veterinary Medicine, Washington State 20
University P.O. Box 647040, Pullman, WA 99164-7040. 21
Phone: 509-335-6075. 22
Fax: 509-335-8529. 23
E-mail: tbesser@vetmed.wsu.edu. 24
ABSTRACT:
25 26
Enterohemorrhagic E. coli O157:H7 (EHEC O157) is an important cause of food and 27
waterborne illness in the developed countries. Cattle are a reservoir host of EHEC O157 28
and a major source of human exposure through contaminated meat products. Shiga toxins 29
(Stx) are an important pathogenicity trait of EHEC O157. The insertion sites of the Stx-30
encoding bacteriophages differentiate EHEC O157 isolates into genogroups commonly 31
isolated from cattle but rarely from sick humans (bovine-biased genotypes, BBG) and 32
commonly isolated from both cattle and human patients (clinical genotypes, CG). Since 33
BBG and CG share the cardinal virulence factors of EHEC O157 and are carried by cattle 34
at similar prevalence, the infrequency of BBG among human disease isolates suggests 35
that they may be less virulent than CG. We compared the virulence potential of human 36
and bovine isolates of CG and BBG in newborn conventional pig and weaned Dutch 37
Belted rabbit models. CG-challenged piglets experienced more severe disease 38
accompanied by earlier and higher mortality than BBG-challenged piglets. Similarly, 39
CG-challenged rabbits were more likely to develop lesions in kidney and intestine when 40
compared with the BBG-challenged rabbits. The CG strains used in this study carried 41
stx2 and produced significantly higher amount of Stx, whereas the BBG strains carried 42
stx2c gene variant only. These results suggest that BBG are less virulent than CG and that 43
this difference in virulence potential is associated with the Stx 2 sub-type(s) carried 44
and/or the amount of Stx produced. 45
INTRODUCTION:
47 48
Enterohemorrhagic E. coli O157:H7 (EHEC O157) is a major cause of food- and 49
water-borne illnesses characterized by bloody diarrhea, hemorrhagic colitis (HC) and 50
life-threatening hemolytic uremic syndrome (HUS) in developed nations across the globe 51
(27, 38, 43). The lack of specific treatment leads to 2 to 5% mortality among HUS 52
patients. The annual incidence of EHEC O157 infection is estimated to be >70,000 cases 53
in the US (19). The CDC surveillance data shows that the number of EHEC O157 54
outbreaks increased recently from 27 in 2006 to 39 in 2007 (8, 9). In 2008, the CDC 55
reported 1.12 cases of foodborne EHEC O157 infections per 100,000 population in the 56
USA (7). The infectious dose of EHEC O157 is estimated to be very low (10-100 57
bacteria) (54), and transmission occurs primarily via contaminated food, water, or direct 58
contact with infected animals (18, 20, 25, 47). Cattle are considered a major reservoir of 59
EHEC O157 due to the frequent associations of EHEC O157 outbreaks with the 60
consumption of contaminated beef and dairy products, confirmed by molecular 61
epidemiologic evidence of identical strain types (3, 5). Cattle colonized with EHEC O157 62
remain asymptomatic while shedding up to 105 cfu / g of feces typically for periods of 63
days to weeks (51). EHEC O157 is defined as an adulterant when identified in foods, and 64
therefore, in addition to its public health significance, this organism is responsible for 65
significant economic losses to the food industry due to mandatory recalls of contaminated 66
meat and meat products. 67
Nearly all EHEC O157 isolates harbor two cardinal virulence factors: production 68
of one or more antigenically distinct Shiga toxins (Stxs) and the presence of a 69
chromosomal pathogenicity island referred to as the Locus of Enterocyte Effacement 70
(LEE) (34, 43). Stxs are encoded in late genes of one or more lambdoid bacteriophages 71
that are integrated into the EHEC O157 bacterial chromosome at specific sites (11, 22, 72
31, 32, 41, 45, 48). Shiga toxins are important virulence factors associated with the 73
systemic effects of EHEC infection including HC and HUS (28, 56). The LEE encodes a 74
type III secretion system (T3SS), including an effector molecule Tir (translocated intimin 75
receptor), and Intimin, an EHEC O157 surface expressed molecule that interacts with Tir 76
and that is also a putative adhesion factor (44). The LEE is required for EHEC O157 77
colonization and formation of attaching and effacing (A/E) lesions on intestinal 78
epithelium (15, 40). 79
Diverse genotyping systems for EHEC O157 strains (4, 29, 30, 36, 41, 52, 62, 63, 80
65) emphasize the role of bacteriophage, including Stx-encoding bacteriophages, in 81
generating genetic diversity in EHEC O157. Several of these methods identified 82
genotypes occurring at different frequencies in cattle and human host (4, 29, 36, 63). 83
Recent studies comparing some of these genotyping methods suggest that they result in 84
generally concordant classifications (33, 60). Using Stx-encoding bacteriophage insertion 85
(SBI) sites as a marker for strain differentiation, EHEC O157 isolates can be classified 86
into two genogroups: bovine-biased genotypes (BBG, predominantly SBI genotypes 5 87
and 6, found primarily in cattle and rarely isolated from sick humans) and clinical 88
genotypes (CG, predominantly SBI genotype 1-3, found in both cattle and humans) (4, 89
52, 61). Both BBG and CG carry and express the cardinal virulence factors of EHEC 90
O157 (4). Despite the similar prevalence of both genogroups in cattle, indicating 91
approximately equal human exposure, the relatively rare occurrence of BBG as a cause of 92
human disease suggests that BBG may be less virulent than CG. Therefore, this study 93
was designed to evaluate the virulence potential of human and bovine strains belonging 94
to BBG and CG, using a newborn conventional pig (NCP) and Dutch Belted rabbit 95
(DBR) models. We show that strains of EHEC O157 belonging to BBG are less virulent 96
and cause less severe disease in both animal models. This difference in virulence 97
potential is associated with the presence of different Stx 2 sub-type(s) and/or the quantity 98
of Stx produced by BBG strains. 99
100
MATERIALS AND METHODS
101 102
Piglet husbandry and experimental challenges
103
All animal experiments were conducted according to the protocols approved by the 104
WSU Institutional Animal Care and Use Committee (IACUC). Piglet challenges were 105
conducted as described previously (14) with a few modifications as follows. Piglets were 106
obtained from WSU swine center at ~4 h after birth, and confirmed EHEC O157-free by 107
testing fecal swabs obtained prior to challenge by using immunomagnetic separation 108
(Dynabeads anti-E. coli O157, Invitrogen Dynal AS, Oslo, Norway). Piglets were orally 109
treated with nalidixic acid (25 mg) followed by sodium bicarbonate (10 ml, 10% w/v) 1 h 110
later. Immediately after sodium bicarbonate treatment, piglets were orally challenged 111
with nalidixic acid resistant EHEC O157 (~1010 cfu, NalR). Piglets were fed Enfamil 112
infant milk formula (Mead Johnson & company, IN USA) three times daily beginning at 113
210 ml/day and gradually increasing to 450ml/day on day 6 and 7 post infection (PI). 114
Nalidixic acid (25 mg every 8 h) was orally administered to each piglet throughout the 115
experiment. To avoid confounding by potential genetic factors, each piglet cohort (4 – 6 116
piglets from single litter) was randomly allocated equally to CG and BBG strain 117
challenges. Piglets were observed daily for EHEC O157-associated clinical signs (Table 118
1) for up to 7 days PI. Fecal samples from each piglet were collected for quantification of 119
EHEC O157. Piglets were treated with oral fluids and electrolytes (Enterolyte H.E., Oral 120
Powder, Pfizer) and flunixin meglumine (Banamine 1 mg/lb BW IM q24 h) when 121
showing signs of diarrhea and moderate CNS disease (shivering, tremors or mild 122
seizures). Piglets exhibiting severe clinical signs (severe seizures, lateral recumbency, or 123
paresis) were humanely euthanized. At 7 days PI, all surviving piglets were humanely 124
euthanized. Complete necropsies were performed on all piglets, and internal organs were 125
collected and processed for bacteriological, histopathological and electron microscopic 126
examination. All clinical and pathologic assessments were conducted by veterinarians 127
blind to the challenge genotype. 128
129
Bacterial strains for NCP
130
Twenty EHEC O157 strains (Table 2) were randomly selected from the bank of SBI-131
genotypes of EHEC O157 at Washington State. EHEC O157 strain Sakai (CG-3, kindly 132
provided by Dr. Thomas S. Whittam, NFSTC-MSU) isolated from an outbreak in Japan 133
served as positive control. In order to generate spontaneous nalidixic acid resistant 134
variants (NalR), each EHEC O157 strain was inoculated into brain heart infusion (BHI) 135
broth, incubated at 37°C overnight (~16 h), and 1 ml was plated on Sorbitol MacConkey 136
agar plates (Hardy Diagnostics, Santa Maria, CA, USA) supplemented with nalidixic acid 137
(30 µg/ml) (SMAC-N). For animal challenge studies, NalR strains were inoculated in 200 138
ml of trypticase soy broth (TSB) and incubated at 37°C overnight with shaking at 200 139
rpm. Aliquots of these cultures were then stored in glycerol (10% v/v) stocks at -80°C 140
until use. 141
Bacteriological culture for piglet challenge
142
EHEC O157 were detected and enumerated in fecal samples and intestinal tissues of 143
each piglet by decimal dilution plating on SMAC-N agar plates using a spiral plater 144
(Whitley Automated Spiral Platter, Don Whitley Scientific Ltd, UK). Ten sorbitol non-145
fermenting colonies from each specimen were tested for lactose fermentation 146
(MacConkey’s agar, Becton, Dickinson and Company, Sparks, MD, USA) and for beta-147
glucuronidase activity (EC-MUG agar, Hardy Diagnostics, Santa Maria, CA, USA). 148
Presumptive EHEC O157 colonies (sorbitol negative, lactose positive and beta-149
glucuronidase negative) were confirmed by O157 latex agglutination test (E. coli PRO 150
O157, Hardy Diagnostics, USA). One confirmed EHEC O157 colony recovered from 151
each sample was stored (18% buffered glycerol in BHI, -80°C) for subsequent 152
comparisons with their respective challenge strains by SBI-genotyping (61) and by 153
pulsed field gel electrophoresis (PFGE) following XbaI restriction digestion (6, 12). 154
155
Histopathological examination of piglet tissues
156
Duodenum, jejunum, ileum, cecum, spiral colon, distal colon, recto-anal junction (RAJ), 157
kidney and brain collected at necropsy were immediately fixed in 10% neutral buffered 158
formalin, and subsequently paraffin-embedded, sectioned, and stained with hematoxylin 159
and eosin for histopathologic examination. Brain tissue sections were also stained with 160
Luxol-fast blue stain (59). Immunohistochemistry was performed on selected tissues 161
using a mouse IgG1 anti-O157 monoclonal antibody and the AEC detection kit 162
(SIGNET, Covance, CA). For EM, tissues were fixed in McDowell’s and Trump’s 4F:1G 163
(37). Histopathologic scoring was based on observed bacterial attachment, necrosis, 164
inflammation, vasculitis, and presence or absence of fibrin. Scoring was performed 165
independently by two ACVP board-certified pathologists (KP and KL) and by SS blinded 166
to the identity of the challenge strain. The histopathological lesions in brain and kidney 167
were scored by two observers (KP and KL), and in intestine by three observers (KP, KL, 168 and SS). 169 170 Rabbit experiments 171
All rabbit experiments were approved by the IACUC at Massachusetts Institute of 172
Technology. Three challenge experiments were performed using weaned 7 to 8-week-old 173
Dutch Belted rabbits (DBR). Each challenge experiment included five rabbits each for 174
positive controls inoculated with EHEC O157 strain EDL933 (CG-3) and four to five 175
rabbits each for negative controls inoculated with PBS, along with six rabbits each 176
inoculated with the CG or BBG challenge strain of EHEC O157. In each experiment, 177
rabbits were fasted overnight and sedated prior to blood collection and orogastric 178
intubation. Experimental and control rabbits were inoculated with 10 ml of 10% sterile 179
sodium bicarbonate to neutralize gastric acidity. Experimental rabbits received 1-2 × 109 180
colony-forming-units of EHEC O157 re-suspended in sterile PBS and control rabbits 181
received sterile PBS (vehicle only). Food and water were provided ad libitum following 182
inoculation. Rabbits were monitored closely for clinical signs and body weight 183
measurements were recorded. Fecal cultures were performed on days 1 and 3 PI. Rabbits 184
were euthanized on day 6 or 7 PI; however, rabbits exhibiting severe clinical signs were 185
euthanized before the end of the study (day 4 PI). Prior to euthanasia, blood samples were 186
collected for clinical pathological evaluation. Complete necropsies were performed on all 187
the rabbits and tissue samples from various organs including cecum and kidneys were 188
collected for histopathological evaluation. 189
190
Bacterial strains for DBR
191
Five challenge strains were selected for the rabbit infection studies (Table 2). Stock 192
inocula containing ~ 1010 cfu was prepared and stored in 1 ml aliquots at -80°C until use. 193
194
Bacteriological culture for DBR
195
Challenge strains were detected… 196
197
Histopathological examination of rabbit tissues
198
Tissues were fixed in formalin, embedded in paraffin, sectioned at 5 µm, and stained 199
with hematoxylin and eosin. Kidney sections were also stained with Carstairs’ to detect 200
fibrin deposition. Kidney and intestinal (cecal) sections were evaluated by a board-201
certified veterinary pathologist (SM) blinded to the identity of the challenge strain. 202
Scoring criteria for histopathological evaluation included renal glomerular (capillary 203
thickening, luminal constriction, fibrin thrombi, white blood cell infiltration, mesangial 204
deposits, changes in bowman’s capsule and space) and vascular (intimal swelling, mural 205
degeneration, perivascular edema, and red blood cell fragmentation) criteria as well as 206
assessment of intestinal vasculopathy (17). 207
C-reactive protein analysis
208
The concentration of C-reactive protein (CRP) was measured in serum samples from 209
rabbits in experiment 3 using a commercially available rabbit C-reactive protein ELISA 210
(Immunology Consultants Laboratory, Inc., Newberg, OR) following the manufacture’s 211
directions. Rabbit serum samples were diluted 1:400 in assay diluent buffer for testing. 212
Samples were adjusted to a 1:10,000 dilution and retested if they did not fall within the 213
working range of the assay’s standard curve. The normal range of rabbit CRP is 0-31 214 µg/ml (Gentry PA, 1999). 215 216 stx2 gene typing 217
The stx2 gene variants present in the challenge strains were determined using
PCR-218
RFLP (13). In this method, restriction patterns generated by HaeIII differentiate stx2 219
from stx2c while restriction pattern generated by PvuII differentiate stx2 and stx2c from 220
other stx2 variant types. 221
222
Stx Quantitation
223
Stx production in NalR challenge strains and their nalidixic acid susceptible wild-type 224
parents (WT-NalS) were assessed using ELISA (Premier EHEC test kit, Meridian 225
Bioscience, Cincinnati, OH). Briefly, for each strain, single colonies from blood agar 226
plates were inoculated into 10 ml of TSB, incubated at 37ºC overnight and used to 227
prepare fresh TSB subcultures by inoculating 500 µl of overnight culture into 10 ml of 228
fresh TSB. This subculture was processed for bacterial counts and three aliquots (1 ml 229
each) were transferred into three wells of 96-well culture plates followed by addition of 230
1) carbadox (0.5 µg/ml, Sigma-Aldrich, St. Louis, MO), 2) nalidixic acid (30 µg/ml) or 3) 231
no inducing agent. The plates were incubated overnight at 37°C with shaking at 250 rpm. 232
Overnight cultures were diluted 1:100 in sterile TSB immediately followed by 1:2 233
dilution in sample diluent provided in the ELISA kit and 100 µl aliquots of diluted 234
samples were tested by ELISA according to the manufacturer’s protocol. 235
236
Statistical analysis
237
Statistical analyses were performed using NCSS 2007 version 07.1.19 (23). For NCP 238
challenge study, the genogroups were compared using following statistical tests. The 239
clinical scores were analyzed using one- way ANOVA. The five clinical scoring 240
categories (diarrhea, CNS disease, appetite, vomiting and abnormal vocalization) were 241
analyzed using two-way generalized linear method (glm). Survival data was compared 242
using Kaplan-Meier survival analysis and the Log-Rank test. Bacterial load recovered 243
from five segments of intestine (ileum, cecum, spiral colon, distal colon and RAJ) of 244
piglets was compared using two-way glm. Scores for histopathological lesions in 245
intestinal tract were analyzed using two-way (genogroup and pathologist) glm. 246
Association of development of brain lesions in the piglets with the development of CNS 247
disease or with the genogroup of the EHEC O157 strain used to challenge the piglets was 248
assessed using Fisher’s exact test. In the DBR model, statistical analyses were performed 249
using GraphPad Prism. The Mann-Whitney two-tailed test was used to compare the 250
lesion scores of experimental versus control rabbits. All rabbits inoculated with EDL933 251
(positive controls) and those inoculated with PBS (negative controls) in the three 252
challenge experiments were grouped for the statistical analyses. Association between stx 253
gene typing and genogroup of the challenge strain was assessed using chi-square test. Stx 254
ELISA OD450 scores were analyzed using three-way (growth condition, nalidixic acid
255
susceptibility, and genogroup) glm. Multiple comparisons were only done when the 256
overall statistical model was significant using Tukey-Kramer test. Each statistical test 257
was conducted at the significance level of P≤0.05. 258 259 260 261 RESULTS 262 263 NCP Model 264 Clinical Signs 265
Piglets were scored for disease using a standard scoring system (Table 1). The piglets 266
cohorts did not differ significantly (P=0.138). Piglets challenged with the clinical 267
genotype strains (CG-1 and CG-3) exhibit higher clinical scores when compared with the 268
piglets challenged with bovine-biased genotype strains (BBG-5 and BBG-6) (P=0.005, 269
Table 3). Piglets challenged with CG strains and BBG strains had clinical scores that did 270
not differ significantly from the positive control Sakai strain and sham inoculated piglets, 271
respectively (P>0.439). In individual genotype comparisons, CG-3 challenged animals 272
had significantly higher clinical scores than BBG-6 inoculated piglets (P=0.023, Fig 1), 273
while CG-1 and BBG-5 challenged piglets showed intermediate clinical scores. Diarrhea, 274
CNS symptoms, appetite, vomiting, and abnormal vocalizations were scored separately. 275
Analyses of these categories suggested that CNS disease scores were significantly higher 276
for piglets challenged with CG strains when compared with the piglets challenged with 277
BBG strains (P=0.004, Table 3). No other scoring category differed significantly between 278
CG and BBG challenged piglets. 279
Mortality
280
Challenge with CG strains resulted in earlier and higher mortality than challenge with
281
BBG strains (P=0.002, Fig 2 and Table 3). Sham-inoculated piglets survived until the end 282
of experiment. CG-challenged piglets showed mortality similar to the piglets challenged 283
with positive control strain Sakai (P=0.852). Out of 10 piglets challenged with CG 284
strains only one piglet (challenged with strain E12064) survived until the end of the 285
experiment. On the contrary, out of 10 piglets challenged with BBG strains only two 286
piglets (challenged with strain E12053 and E12156) died on days 2 and 4 PI. 287
Magnitude of bacterial load
288
The mean±SEM log10 cfu/cm2 of EHEC O157 recovered from intestine (ileum, cecum,
289
spiral colon, distal colon and RAJ) were significantly higher for piglets challenged with 290
CG strains, when compared with the piglets challenged with BBG strains (P=0.003, 291
Table 3). The mean±SEM log10 cfu/cm2 of EHEC O157 recovered from ileum, cecum,
292
spiral colon, distal colon and RAJ were 3.86±0.40, 4.92±0.32, 4.56±0.22, 5.00±0.26 and 293
5.14±0.27, respectively. Although the number of bacteria recovered from large intestine 294
was higher when compared with small intestine, statistically significant differences were 295
only observed between RAJ and ileum. 296
Negative control piglets were housed in separate cages within the same room and 297
at the same time as EHEC challenged piglets. EHEC O157 was recovered from some 298
negative control piglets, albeit in small numbers, indicating the likely cross-transmission 299
of EHEC O157 between challenged and sham-inoculated animals. Therefore, SBI 300
genotypes and PFGE profiles of challenge strains were compared with the strains 301
recovered from each piglet to determine the level of cross-transmission, if any. In all but 302
two cases, only the challenged strain was recovered from each experimental piglet, 303
indicating that cross-transmission was uncommon. Both of the cross contaminated piglets 304
were challenged with BBG-6 strain. In one of these BBG-6 challenged piglets, one out of 305
6 recovered isolates belonged to a genotype (CG-3) that was different from the genotype 306
of the challenge strain. In other BBG-6 challenged piglet, all seven isolates recovered 307
belonged to non-BBG-6 genotypes (BBG-17, one isolate; BBG-5, one isolate; CG-1, five 308
isolates). However, the clinical scores of these two piglets were lower than the piglets 309
that were challenged with the cross-contaminating CG strains in the same cohort, 310
suggesting that this transmission had minimal effect on the clinical outcomes. Isolates 311
recovered from challenged animals occasionally differed in PFGE profiles by one or two 312
bands from the challenge or contaminating strain. Two genotypes that were not included 313
in the study (genotype 9, negative for all the six PCR fragments in SBI genotyping, and 314
genotype 17, positive for only yehV-left junction PCR fragment in SBI genotyping) were 315
recovered from two piglets. These piglets had been either challenged or contaminated by 316
BBG-5 strain (positive for stx2 gene and yehV-left junction PCR fragments in SBI 317
genotyping) suggesting loss of stx2 encoding bacteriophage or loss of stx2 gene either in-318
vivo or during the isolation process. However, the PFGE profiles of these recovered 319
isolates were closely related to the challenge or the contaminating BBG-5 strain, 320
confirming their origin. 321
Histopathology
The histopathological scores for intestinal tract in CG and Sakai challenged piglets were 323
higher than BBG and sham inoculated piglets but did not reach statistical significance 324
(P=0.071, Table 3). However when the two genogroups (CG and BBG) were compared 325
the histopathological scores of intestinal tract for CG-challenged piglets were 326
significantly higher than BBG-challenged piglets (P=0.02). Brain lesions were scored as 327
present or absent (Fig 3). The number of piglets in which brain lesions were observed by 328
at least one observer were 5/10, 2/10, 2/4, and 0/4 in CG, BBG, positive and negative 329
control groups, respectively. The number piglets showing brain lesions did not differ 330
significantly between CG and BBG challenged piglets (P=0.159). However, 7 out of 11 331
piglets that died or were euthanized following severe CNS signs did exhibit brain lesions 332
while none of 9 piglets that did not exhibit severe CNS signs or mortality developed brain 333
lesions (P<0.003). No kidney lesions were observed in any of the piglets. 334
335
DBR Model
336
Three challenge experiments were performed to compare the virulence potential of CG 337
and BBG strains in the DBR model. The most severe clinical signs were observed in 338
rabbits from experiment three. Rabbits in this experiment were challenged with EDL933, 339
E3046 and E5252. Three rabbits (one in each infection group) was euthanized early (at 4 340
days PI) due to the severity of diarrhea. The % change in body weight of these three 341
rabbits infected with EDL933, E3046, and E5252 at day 4 PI was -1.8%, -4.6%, and 342
11.5%, respectively. 343
Pre- and post-inoculation clinical pathological parameters including hematocrit, white 344
blood cell count, percent heterophils, platelet count, blood urea nitrogen, total protein, 345
and albumin were compared for rabbits in each group; however, analyses did not reveal 346
consistent changes associated with disease. In experiment three, increased serum 347
concentration of CRP was detected in rabbits that were euthanized on day 4 PI due to 348
severe disease: 451.4 µg/ml (EDL933), 162.1 µg/ml (E3046), and 399.5 µg/ml (E5252). 349
These values were approximately 4-fold higher than the group mean and were consistent 350
with published rabbit values (65-350 µg/ml) indicative of an acute phase response 351
(Gentry PA, 1999). 352
On gross necropsy, infected rabbits exhibited serosal hemorrhages near the cecocolic 353
junction (Fig. 4). Specifically, these hemorrhages were observed in approximately 33% 354
(5/15), 17% (1/6), and 83% (5/6) of rabbits infected with EDL933, E3046, and E5252, 355
respectively. Serosal hemorrhages were not observed in other infected groups or 356
uninfected controls. 357
Significant lesions in the cecum and kidneys of rabbits are summarized in Table 4. 358
Relative to sham-inoculated control rabbits, significant intestinal vasculopathy (Fig.5 B 359
and C) was observed in most O157:H7 infected rabbits except in those infected with 360
strain E5880. .Rabbits infected with E5252 developed the most renal lesions that were 361
significant and included both glomerular and vascular lesions such as glomerular 362
capillary thickening (Fig.5E), luminal constriction (Fig.5F), erythrocyte fragmentation 363
(Fig.5F), intimal swelling and perivascular edema. Rabbits infected with E3046 364
developed significant renal vascular lesions that included intimal swelling and 365 perivascular edema. . 366 367 stx2 gene typing 368
All the CG strains tested in this study carried stx2, and two of the five CG-1 strains also
369
carried stx2c. In contrast, all BBG strains carried stx2c alone (P<0.001). 370
371
Stx Quantitation
372
The number of bacteria in the initial subculture used for induction by nalidixic acid or 373
carbadox or left un-induced was ~107cfu/ml (range, 1.6x107cfu/ml to 7.8x107 cfu/ml). 374
There was no significant difference in the bacterial counts (mean±SEM log10 cfu/ml) of
375
BBG (7.54±0.06) when compared with CG (7.65±0.03) at the time of induction 376
(P=0.289). Induction with carbadox resulted in a significant increase in the Stx ELISA 377
OD450 scores irrespective of the genogroup (P=0.05), while induction with nalidixic acid
378
did not result in a significant increase in Stx ELISA OD450 scores by any group, probably
379
due to the NalR trait of all challenge strains. CG strains produced significantly higher Stx 380
ELISA OD450 scores than BBG strains under non-inducing and carbadox inducing
381
conditions (P=0.05). In individual genotype comparisons, challenge strains belonging to 382
CG-1 and CG-3 produced significantly higher Stx ELISA OD450 scores than BBG-5 and
383
BBG-6 under carbadox inducing conditions (P=0.05, Fig. 6). However, under non-384
inducing conditions, challenge strains belonging to CG-1 produced significantly higher 385
Stx ELISA OD450 scores than BBG-5 and BBG-6. Nalidixic acid induction resulted in no
386
significant difference between genotypes. There was no difference in production of Stx 387
between of NalS and NalR strains within the genotypes under any of the tested growth 388
conditions, suggesting that the use of nalidixic acid to suppress the normal flora of the 389
experimental piglets did not affect in vivo Stx production during these challenge 390
experiments. 391
392
DISCUSSION
393 394
The relatively decreased virulence of BBG strains support the hypothesis that not all 395
EHEC O157 SBI genotypes are equally pathogenic. This is evident based on the higher 396
clinical disease severity, earlier and higher mortality, and more severe histopathological 397
lesions observed for piglets challenged with CG strains. Similar results were obtained in 398
rabbits where animals challenged with CG strains exhibited significantly higher 399
histopathological scores. In contrast, the BBG strains generally showed little virulence in 400
either animal model. Our observations are consistent with the previously published study 401
where EHEC strains from healthy cattle were, on average, less virulent in gnotobiotic 402
piglets than EHEC isolated from human disease outbreaks (2). Here we can refine these 403
observations to show that virulence in animal models is associated with specific EHEC 404
O157 genotypes, and that these differences in virulence correlate with the frequency of 405
occurrence of these genotypes among clinical isolates rather than in the animal reservoir. 406
Further, we show that the CG and BBG genotypes tested in these animal models are 407
characterized by consistent differences in their Stx2 variant gene content, with stx2 408
consistently present in the CG strains (with or without stx2c) while stx2c is consistently 409
present and stx2 is consistently absent in the BBG strains. 410
Our piglet model differed in several ways from that of previously published 411
reports (2). In contrast to the gnotobiotic piglet model, in which ~24 h old piglets were 412
inoculated with~109 cfu of EHEC O157, we used ~4 h old nalidixic acid treated 413
conventional piglets that were pretreated with 10% sodium bicarbonate before challenge 414
with ~1010 cfu of EHEC O157. The piglet model closely reproduces some aspects of 415
EHEC O157-induced systemic disease of human infection. However, one of the 416
important clinical manifestations of EHEC O157 infection in piglets is CNS disease, 417
which is often associated with brain lesions (2). In this case, the animal model 418
demonstrates lesions in a tissue (brain) typically unaffected in infected humans; however, 419
the brain lesions and CNS disease are arguably analogous to Stx-mediated renal disease 420
in humans as they are the result of Stx being absorbed into circulation, damaging Stx-421
receptor-expressing endothelial cells resulting in microangiopathy and target tissue 422
damage (14, 56). CG strains induced more severe CNS symptoms (seizures, convulsions, 423
paresis, and lateral recumbency) as compared with BBG strains. In addition, there was a 424
significant association between CNS disease and the presence of histopathological lesions 425
in the brain. 426
Several recent studies have shown that EHEC O157 infected piglets also develop 427
kidney lesions that appear to be associated with Stx production (2, 21, 46). However, 428
kidney lesions were not observed in the piglets infected with either BBG or CG strains in 429
this study. Perhaps due to the rapid onset of severe CNS disease in piglets, the renal 430
disease is only minimally expressed before death (2). Genetic differences (the breed of 431
pigs), the age at the time of infection and time of euthanasia after the appearance of first 432
CNS signs may have contributed to the observed differences in the severity of brain 433
lesions and lack of kidney lesions in our study compared with the previous studies using 434
piglet model. 435
We present direct evidence of some cross-transmission in our piglet studies; 436
however the direction and degree of cross-transmission were not sufficient to confound 437
the strong bacterial genotype effects on piglet virulence observed here. Cross-438
transmission of EHEC O157 from infected to uninfected piglets has been described 439
earlier by Cornick and Vukhac (10), suggesting that EHEC O157 was readily transmitted 440
among swine via contaminated aerosols and suggesting that a very low dose is required 441
for piglet infection. Similarly, minor differences in bacteriophage content and PFGE 442
pattern between challenge strains and fecal isolates as reported here have been reported 443
previously, for example in experimental bovine infections (64). The infrequency of these 444
observations in our study demonstrates the overall stability of genotypes during piglet 445
infections. 446
The DBR model for EHEC O157 exhibits renal lesions that mimic HUS in 447
humans (17). Therefore, we used this model to evaluate the relative virulence of BBG 448
and CG strains and their capacity to induce intestinal and renal lesions. Rabbits 449
experimentally infected with different EHEC O157 genotypes developed a wide range of 450
intestinal and renal histopathological changes suggesting that the virulence of EHEC 451
O157 varies depending on the strain and/or genotype. More specifically, CG-3 strain 452
E5252 appeared to be the most pathogenic based on the extent and severity of the renal 453
and intestinal lesions. Furthermore, the increased serum concentration of CRP in the most 454
clinically affected rabbits suggested that this acute phase protein may represent a 455
potential biomarker of severe EHEC infection and disease. CRP has been considered a 456
predictive factor for hemolytic uremic syndrome development in humans with E. coli 457
O157:H7 infection (Ikeda et al, 2000). 458
Several studies have associated the amount of Stx produced and/or type of stx 459
gene present to the virulence of EHEC O157 strains (2, 16, 39, 42). Therefore, we 460
compared the stx2 gene subtype present and the quantity of Stx produced by the EHEC 461
O157 strains used in this study. CG strains carry stx2 alone or in combination with stx1 or 462
stx2c genes while BBG strains carry only stx2c gene. Moreover, under bacteriophage 463
inducing conditions using carbadox, CG strains produced higher Stx ELISA OD450 scores
464
when compared with the BBG strains. These results suggest that the presence of stx2 465
gene and/or production of higher amounts of Stx may be contributing to the increased 466
virulence of CG strains. Although, similar relationship between amount of Stx produced 467
and virulence has been reported earlier (2, 16, 39, 42), the quantification of Stx depends 468
on the specificity of antibody used in the kit, which detects Stx1 or Stx2 and Stx2c with 469
different sensitivity (26, 42). In our study, we observed higher Stx ELISA scores for the 470
CG strains when compared with the BBG strains. However, it was not possible for us to 471
attribute this difference solely to the amounts of toxin produced by these strains as they 472
also differ in the type of stx2 gene content. Therefore, a careful interpretation is needed 473
while using ELISA to compare the amount of Stx produced among EHEC O157 strains. 474
HUS/severe disease in humans have been linked to the strains that carry stx2, stx2c 475
or in combination (1, 42). 9 out of 10 piglets challenged with strains carrying stx2, 476
whether alone or in combination with stx1 or stx2c, died. However, only 2 out of 10 477
piglets challenged with the strains carrying stx2c alone died. These results suggest that 478
the EHEC O157 strains that carry stx2c alone are likely to be less pathogenic when 479
compared with the strains that carry stx2 alone or in combination with stx2c or stx1. The 480
higher amount of Stx produced suggested by higher Stx ELISA OD450 scores of CG
481
strains carrying stx2 gene could explain the severe CNS signs that were observed in CG 482
challenged piglets when compared with the BBG challenged piglets. Other researchers 483
have observed the histopathological lesions in the brain and kidney and have attributed 484
these to the systemic effect of Stx (2, 21, 46). In addition to the amount of Stx produced, 485
differences in stx2 gene content could result in differences in the affinity of the toxin to 486
the receptors in the brain and eventual differences in their ability to cause disease or Stx-487
induced damage to the brain (42). 488
Besides inducing systemic disease, recent studies suggest that Stx2 production in 489
the intestine can also promote colonization, by enhancing the expression of host cell 490
intimin receptors, using invitro cell culture and mouse model (35, 50). In contrast to this, 491
similar studies conducted in cattle and rabbit suggested no effect of Stx on colonization in 492
intestine (49, 53). Thus the effect of stx2 gene and more specifically its subtype on 493
colonization in intestine needs more investigation. 494
CG strains induced more severe intestinal histopathological lesions when compared 495
with the BBG strains in the piglet and rabbit models, suggesting differences in their 496
ability to attach to and efface the intestinal mucosa. Recently, whole genome expression 497
microarray has been used to compare the differential gene expression between BBG and 498
CG strains (58). The important virulence factors, including the genes encoded on 499
chromosomal LEE, and several genes encoded on pO157 plasmid (the enterohemolysin 500
ehxA, toxB, and etp genes necessary for T2SS) showed increased expression in the CG 501
when compared with BBG strains. In contrast, the genes essential for survival in the 502
environment, such as acid resistance and stress response, were up regulated in the BBG 503
when compared with the CG strains (24, 55, 57, 58). This study supports our observations 504
suggesting that differential expression of LEE-encoded virulence genes in vivo have led 505
to the differences in the magnitude of intestinal bacterial load and histopathological 506
lesions in the intestines between the piglets and rabbits challenged with CG and BBG 507
strains. This initial damage to the intestinal epithelium may facilitate the absorption of 508
Stx into the blood stream where it may cause more severe systemic damage. 509
In conclusion, we have demonstrated that BBG and CG EHEC O157 differ in 510
virulence potential using two animal models, consistent with the hypothesis that not all 511
EHEC O157 strains are equally pathogenic. The amount of Stx and/or type of stx2 gene 512
present are correlated to the virulence differences of EHEC O157 genotypes and more 513
studies are needed in order to clarify the role of these bacterial factors in pathogenesis 514
and virulence of specific EHEC O157 genotypes. 515
516
ACKNOWLEDGEMENTS
517
This work was funded in part by NIAID NIH contract N01-AI-30055, R21AI073803, and 518
by the Agricultural Animal Health Program, Washington State University College of 519
Veterinary Medicine, Pullman, WA. The authors thank Rebecca A. King, Melissa W. 520
Mobley and Amanda Potter for their technical assistance with studies involving the rabbit 521 model. 522 523 REFERENCES 524
1. Abu-Ali, G. S., L. M. Ouellette, S. T. Henderson, D. W. Lacher, J. T.
525
Riordan, T. S. Whittam, and S. D. Manning. 2010. Increased adherence and
526
expression of virulence genes in a lineage of Escherichia coli O157:H7 527
commonly associated with human infections. PLoS One 5:e10167. 528
2. Baker, D. R., R. A. Moxley, M. B. Steele, J. T. Lejeune, J.
Christopher-529
Hennings, D. G. Chen, P. R. Hardwidge, and D. H. Francis. 2007. Differences
530
in Virulence among Escherichia coli O157:H7 Strains Isolated from Humans 531
during Disease Outbreaks and from Healthy Cattle. Appl Environ Microbiol 532
73:7338-7346.
533
3. Besser, R. E., P. M. Griffin, and L. Slutsker. 1999. Escherichia coli O157:H7
534
gastroenteritis and the hemolytic uremic syndrome: an emerging infectious 535
disease. Annu Rev Med 50:355-367. 536
4. Besser, T. E., N. Shaikh, N. J. Holt, P. I. Tarr, M. E. Konkel, P. Malik-Kale,
537
C. W. Walsh, T. S. Whittam, and J. L. Bono. 2007. Greater diversity of Shiga
538
toxin-encoding bacteriophage insertion sites among Escherichia coli O157:H7 539
isolates from cattle than in those from humans. Appl Environ Microbiol 73:671-540
679. 541
5. Borczyk, A. A., M. A. Karmali, H. Lior, and L. M. Duncan. 1987. Bovine
542
reservoir for verotoxin-producing Escherichia coli O157:H7. Lancet 1:98. 543
6. Centers for Disease Control and Prevention. 1999. One-day (24–28 h)
544
standardized laboratory protocol for molecular subtyping of Escherichia coli 545
serotype O157 by pulsed field gel electrophoresis (PFGE). In t. N. M. S. N. f. F. 546
D. S. PulseNet and A. Centers for Disease Control and Prevention, Ga. (ed.). 547
7. Centers for Disease Control and Prevention. 2009. Preliminary FoodNet Data
548
on the incidence of infection with pathogens transmitted commonly through food-549
-10 States, 2008. MMWR Morb Mortal Wkly Rep 58:333-337. 550
8. Centers for Disease Control and Prevention. 2010. Surveillance for foodborne
551
disease outbreaks --- United States, 2007. MMWR Morb Mortal Wkly Rep 552
59:973-979.
553
9. Centers for Disease Control and Prevention. 2009. Surveillance for foodborne
554
disease outbreaks - United States, 2006. MMWR Morb Mortal Wkly Rep 58:609-555
615. 556
10. Cornick, N. A., and H. Vukhac. 2008. Indirect transmission of Escherichia coli
557
O157:H7 occurs readily among swine but not among sheep. Appl Environ 558
Microbiol 74:2488-2491. 559
11. Creuzburg, K., B. Kohler, H. Hempel, P. Schreier, E. Jacobs, and H.
560
Schmidt. 2005. Genetic structure and chromosomal integration site of the cryptic
561
prophage CP-1639 encoding Shiga toxin 1. Microbiology 151:941-950. 562
12. Davis, M. A., D. D. Hancock, T. E. Besser, and D. R. Call. 2003. Evaluation of
563
pulsed-field gel electrophoresis as a tool for determining the degree of genetic 564
relatedness between strains of Escherichia coli O157:H7. J Clin Microbiol 565
41:1843-1849.
566
13. De Baets, L., I. Van der Taelen, M. De Filette, D. Pierard, L. Allison, H. De
567
Greve, J. P. Hernalsteens, and H. Imberechts. 2004. Genetic typing of shiga
568
toxin 2 variants of Escherichia coli by PCR-restriction fragment length 569
polymorphism analysis. Appl Environ Microbiol 70:6309-6314. 570
14. Dean-Nystrom, E. A., J. F. Pohlenz, H. W. Moon, and A. D. O'Brien. 2000.
571
Escherichia coli O157:H7 causes more-severe systemic disease in suckling 572
piglets than in colostrum-deprived neonatal piglets. Infect Immun 68:2356-2358. 573
15. Donnenberg, M. S., S. Tzipori, M. L. McKee, A. D. O'Brien, J. Alroy, and J.
574
B. Kaper. 1993. The role of the eae gene of enterohemorrhagic Escherichia coli
575
in intimate attachment in vitro and in a porcine model. J Clin Invest 92:1418-576
1424. 577
16. Eklund, M., K. Leino, and A. Siitonen. 2002. Clinical Escherichia coli strains
578
carrying stx genes: stx variants and stx-positive virulence profiles. J Clin 579
Microbiol 40:4585-4593. 580
17. Garcia, A., C. J. Bosques, J. S. Wishnok, Y. Feng, B. J. Karalius, J. R.
581
Butterton, D. B. Schauer, A. B. Rogers, and J. G. Fox. 2006. Renal injury is a
582
consistent finding in Dutch Belted rabbits experimentally infected with 583
enterohemorrhagic Escherichia coli. J Infect Dis 193:1125-1134. 584
18. García, A., J. G. Fox, and T. E. Besser. 2010. Zoonotic Enterohemorrhagic
585
Escherichia coli: A One Health Perspective. ILAR J. 51:221-232. 586
19. Gould, L. H., C. Bopp, N. Strockbine, R. Atkinson, V. Baselski, B. Body, R.
587
Carey, C. Crandall, S. Hurd, R. Kaplan, M. Neill, S. Shea, P. Somsel, M.
588
Tobin-D'Angelo, P. M. Griffin, and P. Gerner-Smidt. 2009. Recommendations
589
for diagnosis of shiga toxin--producing Escherichia coli infections by clinical 590
laboratories. MMWR Recomm Rep 58:1-14. 591
20. Griffin, P. M., and R. V. Tauxe. 1991. The epidemiology of infections caused
592
by Escherichia coli O157:H7, other enterohemorrhagic E. coli, and the associated 593
hemolytic uremic syndrome. Epidemiol Rev 13:60-98. 594
21. Gunzer, F., I. Hennig-Pauka, K. H. Waldmann, R. Sandhoff, H. J. Grone, H.
595
H. Kreipe, A. Matussek, and M. Mengel. 2002. Gnotobiotic piglets develop
thrombotic microangiopathy after oral infection with enterohemorrhagic 597
Escherichia coli. Am J Clin Pathol 118:364-375. 598
22. Hayashi, T., K. Makino, M. Ohnishi, K. Kurokawa, K. Ishii, K. Yokoyama,
599
C. G. Han, E. Ohtsubo, K. Nakayama, T. Murata, M. Tanaka, T. Tobe, T.
600
Iida, H. Takami, T. Honda, C. Sasakawa, N. Ogasawara, T. Yasunaga, S.
601
Kuhara, T. Shiba, M. Hattori, and H. Shinagawa. 2001. Complete genome
602
sequence of enterohemorrhagic Escherichia coli O157:H7 and genomic 603
comparison with a laboratory strain K-12. DNA Res 8:11-22. 604
23. Hintzee, J. 2007. NCSS 2007. NCSS, LLC. Kaysville, Utah, USA.
605
www.ncss.com. 606
24. Kailasan Vanaja, S., T. M. Bergholz, and T. S. Whittam. 2009.
607
Characterization of the Escherichia coli O157:H7 Sakai GadE regulon. J Bacteriol 608
191:1868-1877.
609
25. Karmali, M. A. 2004. Infection by Shiga toxin-producing Escherichia coli: an
610
overview. Mol Biotechnol 26:117-122. 611
26. Karmali, M. A., M. Petric, and M. Bielaszewska. 1999. Evaluation of a
612
microplate latex agglutination method (Verotox-F assay) for detecting and 613
characterizing verotoxins (Shiga toxins) in Escherichia coli. J Clin Microbiol 614
37:396-399.
615
27. Karmali, M. A., M. Petric, C. Lim, P. C. Fleming, G. S. Arbus, and H. Lior.
616
1985. The association between idiopathic hemolytic uremic syndrome and 617
infection by verotoxin-producing Escherichia coli. J Infect Dis 151:775-782. 618
28. Karmali, M. A., B. T. Steele, M. Petric, and C. Lim. 1983. Sporadic cases of
619
haemolytic-uraemic syndrome associated with faecal cytotoxin and cytotoxin-620
producing Escherichia coli in stools. Lancet 1:619-620. 621
29. Kim, J., J. Nietfeldt, and A. K. Benson. 1999. Octamer-based genome scanning
622
distinguishes a unique subpopulation of Escherichia coli O157:H7 strains in 623
cattle. Proc Natl Acad Sci U S A 96:13288-13293. 624
30. Kim, J., J. Nietfeldt, J. Ju, J. Wise, N. Fegan, P. Desmarchelier, and A. K.
625
Benson. 2001. Ancestral divergence, genome diversification, and
626
phylogeographic variation in subpopulations of sorbitol-negative, beta-627
glucuronidase-negative enterohemorrhagic Escherichia coli O157. J Bacteriol 628
183:6885-6897.
629
31. Koch, C., S. Hertwig, and B. Appel. 2003. Nucleotide sequence of the
630
integration site of the temperate bacteriophage 6220, which carries the Shiga toxin 631
gene stx(1ox3). J Bacteriol 185:6463-6466. 632
32. Kotewicz, M. L., M. K. Mammel, J. E. LeClerc, and T. A. Cebula. 2008.
633
Optical mapping and 454 sequencing of Escherichia coli O157 : H7 isolates 634
linked to the US 2006 spinach-associated outbreak. Microbiology 154:3518-3528. 635
33. Laing, C. R., C. Buchanan, E. N. Taboada, Y. Zhang, M. A. Karmali, J. E.
636
Thomas, and V. P. Gannon. 2009. In silico genomic analyses reveal three
637
distinct lineages of Escherichia coli O157:H7, one of which is associated with 638
hyper-virulence. BMC Genomics 10:287. 639
34. Law, D. 2000. Virulence factors of Escherichia coli O157 and other Shiga
toxin-640
producing E. coli. J Appl Microbiol 88:729-745. 641
35. Liu, B., X. Yin, Y. Feng, J. R. Chambers, A. Guo, J. Gong, J. Zhu, and C. L.
642
Gyles. 2010. Verotoxin 2 enhances adherence of enterohemorrhagic Escherichia
643
coli O157:H7 to intestinal epithelial cells and expression of {beta}1-integrin by 644
IPEC-J2 cells. Appl Environ Microbiol 76:4461-4468. 645
36. Manning, S. D., A. S. Motiwala, A. C. Springman, W. Qi, D. W. Lacher, L.
646
M. Ouellette, J. M. Mladonicky, P. Somsel, J. T. Rudrik, S. E. Dietrich, W.
647
Zhang, B. Swaminathan, D. Alland, and T. S. Whittam. 2008. Variation in
648
virulence among clades of Escherichia coli O157:H7 associated with disease 649
outbreaks. Proc Natl Acad Sci U S A 105:4868-4873. 650
37. McDowell, E. M., and B. F. Trump. 1976. Histologic fixatives suitable for
651
diagnostic light and electron microscopy. Arch Pathol Lab Med 100:405-414. 652
38. Mead, P. S., and P. M. Griffin. 1998. Escherichia coli O157:H7. Lancet
653
352:1207-1212.
654
39. Muniesa, M., M. de Simon, G. Prats, D. Ferrer, H. Panella, and J. Jofre.
655
2003. Shiga toxin 2-converting bacteriophages associated with clonal variability 656
in Escherichia coli O157:H7 strains of human origin isolated from a single 657
outbreak. Infect Immun 71:4554-4562. 658
40. Naylor, S. W., A. J. Roe, P. Nart, K. Spears, D. G. Smith, J. C. Low, and D.
659
L. Gally. 2005. Escherichia coli O157 : H7 forms attaching and effacing lesions
660
at the terminal rectum of cattle and colonization requires the LEE4 operon. 661
Microbiology 151:2773-2781. 662
41. Ohnishi, M., J. Terajima, K. Kurokawa, K. Nakayama, T. Murata, K.
663
Tamura, Y. Ogura, H. Watanabe, and T. Hayashi. 2002. Genomic diversity of
enterohemorrhagic Escherichia coli O157 revealed by whole genome PCR 665
scanning. Proc Natl Acad Sci U S A 99:17043-17048. 666
42. Orth, D., K. Grif, A. B. Khan, A. Naim, M. P. Dierich, and R. Wurzner. 2007.
667
The Shiga toxin genotype rather than the amount of Shiga toxin or the 668
cytotoxicity of Shiga toxin in vitro correlates with the appearance of the 669
hemolytic uremic syndrome. Diagn Microbiol Infect Dis 59:235-242. 670
43. Paton, J. C., and A. W. Paton. 1998. Pathogenesis and diagnosis of Shiga
toxin-671
producing Escherichia coli infections. Clin Microbiol Rev 11:450-479. 672
44. Perna, N. T., G. F. Mayhew, G. Posfai, S. Elliott, M. S. Donnenberg, J. B.
673
Kaper, and F. R. Blattner. 1998. Molecular evolution of a pathogenicity island
674
from enterohemorrhagic Escherichia coli O157:H7. Infect Immun 66:3810-3817. 675
45. Perna, N. T., G. Plunkett, 3rd, V. Burland, B. Mau, J. D. Glasner, D. J. Rose,
676
G. F. Mayhew, P. S. Evans, J. Gregor, H. A. Kirkpatrick, G. Posfai, J.
677
Hackett, S. Klink, A. Boutin, Y. Shao, L. Miller, E. J. Grotbeck, N. W. Davis,
678
A. Lim, E. T. Dimalanta, K. D. Potamousis, J. Apodaca, T. S.
679
Anantharaman, J. Lin, G. Yen, D. C. Schwartz, R. A. Welch, and F. R.
680
Blattner. 2001. Genome sequence of enterohaemorrhagic Escherichia coli
681
O157:H7. Nature 409:529-533. 682
46. Pohlenz, J. F., K. R. Winter, and E. A. Dean-Nystrom. 2005. Shiga-toxigenic
683
Escherichia coli-inoculated neonatal piglets develop kidney lesions that are 684
comparable to those in humans with hemolytic-uremic syndrome. Infect Immun 685
73:612-616.
47. Rangel, J. M., P. H. Sparling, C. Crowe, P. M. Griffin, and D. L. Swerdlow.
687
2005. Epidemiology of Escherichia coli O157:H7 outbreaks, United States, 1982-688
2002. Emerg Infect Dis 11:603-609. 689
48. Recktenwald, J., and H. Schmidt. 2002. The nucleotide sequence of Shiga toxin
690
(Stx) 2e-encoding phage phiP27 is not related to other Stx phage genomes, but the 691
modular genetic structure is conserved. Infect Immun 70:1896-1908. 692
49. Ritchie, J. M., C. M. Thorpe, A. B. Rogers, and M. K. Waldor. 2003. Critical
693
roles for stx2, eae, and tir in enterohemorrhagic Escherichia coli-induced diarrhea 694
and intestinal inflammation in infant rabbits. Infect Immun 71:7129-7139. 695
50. Robinson, C. M., J. F. Sinclair, M. J. Smith, and A. D. O'Brien. 2006. Shiga
696
toxin of enterohemorrhagic Escherichia coli type O157:H7 promotes intestinal 697
colonization. Proc Natl Acad Sci U S A 103:9667-9672. 698
51. S.J. Bach, T. A. M., D.M. Veira, V.P.J. Gannon and R.A. Holley. 2002.
699
Transmission and control of Escherichia coli O157:H7—a review. Can. J. Anim. 700
Sci. 82:475–490. 701
52. Shaikh, N., and P. I. Tarr. 2003. Escherichia coli O157:H7 Shiga
toxin-702
encoding bacteriophages: integrations, excisions, truncations, and evolutionary 703
implications. J Bacteriol 185:3596-3605. 704
53. Sheng, H., J. Y. Lim, H. J. Knecht, J. Li, and C. J. Hovde. 2006. Role of
705
Escherichia coli O157:H7 virulence factors in colonization at the bovine terminal 706
rectal mucosa. Infect Immun 74:4685-4693. 707
54. Shimizu, K., T. Asahara, K. Nomoto, R. Tanaka, T. Hamabata, A. Ozawa,
708
and Y. Takeda. 2003. Development of a lethal Shiga toxin-producing
Escherichia coli-infection mouse model using multiple mitomycin C treatment. 710
Microb Pathog 35:1-9. 711
55. Shin, S., M. P. Castanie-Cornet, J. W. Foster, J. A. Crawford, C. Brinkley,
712
and J. B. Kaper. 2001. An activator of glutamate decarboxylase genes regulates
713
the expression of enteropathogenic Escherichia coli virulence genes through 714
control of the plasmid-encoded regulator, Per. Mol Microbiol 41:1133-1150. 715
56. Tarr, P. I., C. A. Gordon, and W. L. Chandler. 2005. Shiga-toxin-producing
716
Escherichia coli and haemolytic uraemic syndrome. Lancet 365:1073-1086. 717
57. Tatsuno, I., K. Nagano, K. Taguchi, L. Rong, H. Mori, and C. Sasakawa.
718
2003. Increased adherence to Caco-2 cells caused by disruption of the yhiE and 719
yhiF genes in enterohemorrhagic Escherichia coli O157:H7. Infect Immun 720
71:2598-2606.
721
58. Vanaja, S. K., A. C. Springman, T. E. Besser, T. S. Whittam, and S. D.
722
Manning. 2010. Differential expression of virulence and stress fitness genes
723
between Escherichia coli O157:H7 strains with clinical or bovine-biased 724
genotypes. Appl Environ Microbiol 76:60-68. 725
59. Watanabe, M., K. Matsuoka, E. Kita, K. Igai, N. Higashi, A. Miyagawa, T.
726
Watanabe, R. Yanoshita, Y. Samejima, D. Terunuma, Y. Natori, and K.
727
Nishikawa. 2004. Oral therapeutic agents with highly clustered globotriose for
728
treatment of Shiga toxigenic Escherichia coli infections. J Infect Dis 189:360-729
368. 730
60. Whitworth, J., Y. Zhang, J. Bono, E. Pleydell, N. French, and T. Besser.
731
2009. Diverse genetic markers concordantly identify bovine origin Escherichia 732
coli O157 genotypes underrepresented in human disease. Appl Environ Microbiol 733
76:361-365.
734
61. Whitworth, J. H., N. Fegan, J. Keller, K. S. Gobius, J. L. Bono, D. R. Call, D.
735
D. Hancock, and T. E. Besser. 2008. International comparison of clinical,
736
bovine, and environmental Escherichia coli O157 isolates on the basis of Shiga 737
toxin-encoding bacteriophage insertion site genotypes. Appl Environ Microbiol 738
74:7447-7450.
739
62. Wick, L. M., W. Qi, D. W. Lacher, and T. S. Whittam. 2005. Evolution of
740
genomic content in the stepwise emergence of Escherichia coli O157:H7. J 741
Bacteriol 187:1783-1791. 742
63. Yang, Z., J. Kovar, J. Kim, J. Nietfeldt, D. R. Smith, R. A. Moxley, M. E.
743
Olson, P. D. Fey, and A. K. Benson. 2004. Identification of common
744
subpopulations of non-sorbitol-fermenting, beta-glucuronidase-negative 745
Escherichia coli O157:H7 from bovine production environments and human 746
clinical samples. Appl Environ Microbiol 70:6846-6854. 747
64. Yoshii, N., Y. Ogura, T. Hayashi, T. Ajiro, T. Sameshima, M. Nakazawa, M.
748
Kusumoto, T. Iwata, and M. Akiba. 2009. Pulsed-field gel electrophoresis
749
profile changes resulting from spontaneous chromosomal deletions in 750
enterohemorrhagic Escherichia coli O157:H7 during passage in cattle. Appl 751
Environ Microbiol 75:5719-5726. 752
65. Zhang, Y., C. Laing, M. Steele, K. Ziebell, R. Johnson, A. K. Benson, E.
753
Taboada, and V. P. Gannon. 2007. Genome evolution in major Escherichia coli
754
O157:H7 lineages. BMC Genomics 8:121. 755
756 757
Table 1. Scoring system for clinical signs applied to neonatal piglets.
758
Diarrhea (consistency, frequency, straining ) (0-3) CNS signs (0-5) Vomiting (0-1) Loss of appetite (0-2) Abnormal Vocalization (0-1)
None (formed feces) (0) None (0) None (0) None (0) None (0)
Loose (not formed), no straining or blood (1)
Lethargy (1) Vomiting (1) Slight (1) Present (1)
Thick liquid, more than twice, mild straining and staining of hind limbs with feces (2)
Splayleg, hind limb weakness, thinner appearance (2)
Complete (2)
Runny watery- with/without blood, severe straining and staining of hind limbs with feces (3)
Shivering, tremors (3)
Seizures, convulsions (4) Grand mal seizures or lateral recumbency or paralysis of legs (5) 759
Table 2. EHEC O157 strains used in this study and their origin.
760
EHEC Bank # SBI Genotypea Yearb Source and Location of collectionc Cohortd
CDC EDL 933 CG-3 Raw hamburger meat implicated in
hemorrhagic colitis outbreak NA
E2325 CG-1 1995 Bovine fecal, WA USA R1
E3046 CG-3 1995 Bovine fecal, WA USA R3
E5252 CG-3 1998 Bovine fecal, OR, USA R3
E5880 BBG-5 1999 Water, USA R2
E6996 BBG-6 2000 Bovine fecal, WA, USA R2
E12000 (Sakai) CG-3 1996 Sakai City, Osaka Pref. Japan NA
E12053 BBG-5 1993 Bovine fecal, USA P3
E12056 BBG-5 1999 Water, USA P1
E12057 BBG-6 1999 Bovine fecal USA P2
E12058 BBG-6 2000 Bovine fecal, WA, USA P1
E12061 CG-1 2004 Human Clinical WADOH, WA USA P2
E12062 CG-1 2004 Human Clinical WADOH, WA USA P5
E12063 CG-3 2004 Human Clinical WADOH, WA USA P2
E12064 CG-1 2005 Human Clinical WADOH, WA USA P1
E12065 CG-3 2005 Human Clinical WADOH, WA USA P3
E12066 CG-1 2006 Human Clinical WADOH, WA USA P3
E12067 CG-3 2006 Human Clinical WADOH, WA USA P1
E12068 BBG-5 1996 Bovine fecal, TX, USA P5
E12149 BBG-6 1995 Bovine fecal, WA USA P4
E12153 BBG-6 2001 Bovine fecal, WA, USA P4
E12154 CG-1 2004 Human Clinical WADOH, WA USA P5
E12155 CG-3 2006 Human Clinical WADOH, WA USA P5
E12156 BBG-5 1994 Bovine fecal, WA, USA P5
E12377 CG-3 2004 Human Clinical WADOH, WA USA P5
a
CG=Clinical genotypes, BBG=Bovine-biased genotypes 761
b
WADOH=Washington State Department Of Health 762
c
Year of the strain isolation or time deposited in WSU EHEC bank. 763
d
P=Litter cohorts of piglets, R= Experimental cohorts of Rabbits, NA= Not Assigned 764 765 766 767 768 769 770 771 772 773 774 775
Table 3. Clinical scores, clinical scores for CNS disease, survival data, intestinal bacterial load and histopathological scores 777
for intestine in the piglets challenged with BBG strains, CG strains, positive (Sakai) and negative (sham-inoculated) control 778
piglets. 779
Groups not sharing superscript letters differed significantly (P≤0.05). 780
Disease Outcome Groups of piglets (mean ± SEM)
BBG CG Negative Positive
Clinical scores 1.5 ± 0.45 a 4.12 ± 0.66 b 0.98 ± 0.34 a 3.96 ± 0.26 ab CNS disease Scores 0.69 ± 0.25 a 2.39 ± 0.46 b 0.07 ± 0.07 a 2.35 ± 0.28 b Survival time (days) 6.20 ± 0.55 a 3.70 ± 0.06 b 7.00 ± 0.00 a 4.25 ± 0.63 ab Intestinal bacterial load (cfu) 4.30 ± 0.18 a 5.09 ± 0.19 b 1.13 ± 0.47 c 5.85 ± 0.17 b Histopathological scores of intestine 1.08 ± 0.23 2.01 ± 0.30 1.00 ± 0.52 2.88 ± 0.58
Table 4. Intestinal and renal lesions in rabbits infected with various Escherichia coli 781 O157:H7 strains* 782 O157:H7 strain Intestinal vasculopathy Capillary thickening Luminal constriction Erythrocyte fragmentation Intimal swelling Perivascular edema EDL933 P<0.002 NS NS NS NS NS E5252 P<0.003 P<0.03 P<0.008 (P<0.006) P<0.008 P<0.003 (P<0.0003) P<0.02 (P<0.003) E3046 P<0.01 NS NS NS P<0.03 (P<0.004) P<0.04 (P<0.007) E2325 P<0.007 NS NS NS NS NS E6996 P<0.04 NS NS NS NS NS E5880 NS NS NS P<0.01 NS NS
*, P values are relative to sham-inoculated controls; NS, not significant; P values in 783
parenthesis are relative to EDL933-infected rabbits. 784 785 786 787 788 789 790 791 792 793 794 795 796 797
Figure 1. Clinical scores of piglets challenged with EHEC strains belonging to different
798
genotypes. Groups not sharing specific letters differed significantly (P=0.023). 799
Genotype CG-1 CG-3 BBG-5 BBG-6 0 1 2 3 4 5 6 800 801 802 803 804 805 806
Figure 2. Kaplan-Meier survival curves for piglet groups challenged with different
807
EHEC O157 genogroups. 808 Time (Days) 0 1 2 3 4 5 6 7 0.0 0.2 0.4 0.6 0.8 1.0 BBG CG Negative (Sham-inoculated) Positive (Sakai) 809 810 811 812 813 814
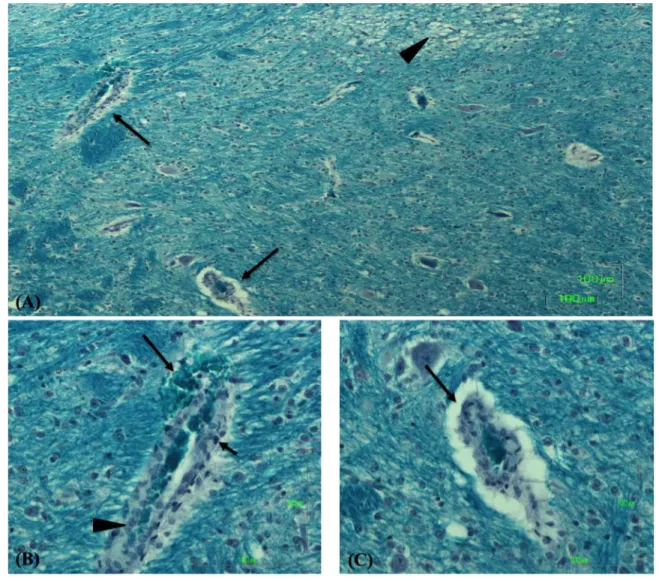
Figure 3. Piglet challenged with positive control Sakai strain. (A) Section of the brain
815
stained with LFB stain showing affected blood vessel (arrows, magnified in fig. 3B and 816
3C) with vacuolation of adjacent tissues (arrow head) caused by focal myelin 817
degeneration. (B) Affected blood vessel in brain showing microhemorrhage (long arrow), 818
pyknosis (short arrow) and hyperplasia (arrow head). (C) Affected blood vessel in brain 819
showing perivascular edema. Bar=100µm. 820
821 822 823 824
Figure 4. Serosal hemorrhage in the area of the distal cecum adjacent to the junction
825
with the proximal colon in a Dutch Belted rabbit infected with Escherichia coli O157:H7 826 strain EDL933. 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842