Autonomous replication of integrative and conjugative elements
Laurel D. Wright
A.B. Biology Brown University
2009
Submitted to the Department of Biology in partial fulfillment of the requirements for the degree of
Doctor of Philosophy in Biology Massachusetts Institute of Technology
September 2016
C 2016 Laurel D. Wright. All rights reserved.
The author hereby grants to MIT permission to reproduce and to distribute publicly paper and electronic copies of this thesis document in whole or in part
in any medium known or hereafter created. /11
Signature of the author:
Certified by: Certified by: MASSACHUSETTS INSTITUTE OF TECHNOLOGY
SEP
017
2016
LIBRARIES
ARCHIVES
Signature redacted
Department of Biology July 15, 2016Signature redacted..
) Alan D. Grossman Professor of Biology Thesis SupervisorSignature redacted
Michael Hemann Associate Professor of Biology Chair, Committee for Graduate StudentsAutonomous replication of integrative and conjugative elements
byLaurel D. Wright
Submitted to the Department of Biology on July 15, 2016 in partial fulfillment of the requirements for the degree of
Doctor of Philosophy in Biology
Abstract
Mobile genetic elements facilitate movement of genes, including those conferring antibiotic resistance and other traits, between bacteria. Integrative and conjugative elements (ICEs), also known as conjugative transposons, are a large family of mobile genetic elements that can transfer between neighboring cells. ICEs are found integrated in the chromosome of their host bacterium, where they are transmitted to daughter cells by chromosomal replication and cell division. Under certain conditions, ICE DNA will excise and form a circular plasmid-like intermediate. It was previously thought that ICEs were incapable of autonomous replication. However, my research, along with the work of others, shows that ICEs can replicate
autonomously, and that many ICEs utilize a rolling circle replication mechanism.
Plasmids and phages that use rolling circle replication encode a single strand origin (sso) that enhances priming of DNA synthesis. We identified a functional single strand origin, ssol, in the integrative and conjugative element ICEBs1 of Bacillus subtilis. Genetic analyses indicated that ICEBs1 uses ssol and at least one other region for second strand DNA synthesis. Sso activity was important for autonomous, rolling circle replication of ICEBs1 in host cells, and for stable acquisition of the element in new host cells.
I also showed that the broad-host range ICE Tn916 replicates autonomously by a rolling
circle mechanism. Replication of Tn916 was dependent on the relaxase encoded by Tn916 orf2O. The origin of transfer of Tn916, oriT(916), also functioned as an origin of replication. I found that the relaxase (Orf2O) and the two putative helicase processivity factors (Ort22 and Orf23)
encoded by Tn916 likely interact in a complex to facilitate replication. Lastly, I identified a functional single strand origin of replication (sso) in Tn916 that I predict primes second strand synthesis during rolling circle replication.
The importance of autonomous replication by rolling circle in the ICE lifecycle and horizontal gene transfer processes is discussed.
Thesis Supervisor: Alan D. Grossman Title: Professor of Biology
Acknowledgements
I am tremendously grateful for Alan's mentorship and the opportunity to work in his lab. Alan's
ability to balance hard-ass-ness and compassion is amazing. He pushed me when I needed to be pushed (which was basically always), and coddled me when I needed to be coddled (which was also basically always). Also he is very smart, which was helpful.
Thank you to Mike and Steve for always being enthusiastic about my science and for offering critical feedback and insight.
I will miss the Grossman lab tremendously. I got to share my room with two of the most
hilarious people I know, Mary and Tyler. Thankfully they are also good scientists, so I
sometimes got work done. Thank you also to Tyler for his mentorship during my early years. Lu, Moni and Mary are a girl squad that rivals T Swift's. Thank you to C. Lee for showing me the alchemy of microbial genetics, and to Chris for his scientific mentorship. C Lee and Chris' scientific rigor and unwavering Zen-ness amaze me, and they were tremendous role models.
I'm grateful to Mark Johnson for giving me my first shot at bench science, and to Juan Rivera and Ana Olivera for taking on a still-novice. I had wonderful colleagues and unofficial mentors in the Johnson and Rivera labs. Thanks especially to Alex Leydon, Ryo Suzuki and Jennifer Sargent for being such cool big kids.
Thank you to David for being an amazing colleague from my first days in a lab, and a loving partner during grad school.
Above all, thank you to my parents, for driving me all over the city of Rochester and beyond for music lessons and theater and nerd camps, for always making me bring a book, and for
Table of Contents
Abstract 2 Acknowledgements 3 List of Tables 5 List of Figures 6 Chapter I Introduction 7Chapter 2 Identification of a single strand origin 43
of replication in the integrative and conjugative element ICEBsJ of Bacillus subtilis
Chapter 3 Autonomous replication of the 87
conjugative transposon Tn916
List of Tables
Chapter 3 Table 1. ICEBs gene function 17-18
Table 2. Tn916 gene function 21
Chapter 2 Table 1. ICEs with regions identical or similar to 69 known single strand origins from plasmids.
Table 2. Bacillus subtilis strains used 82 Chapter 3 Table 1. Mass spectrometry of affinity-purified 106
Orf2O-his shows that the HelP homologues are associated with the relaxase
Table 2. Complementation of the Tn916 106 replication defects of relaxase mutants AorJ20
and orJ2O-3UAA
Table 3. sso916 is conserved in other Tn916-like 111 ICEs
List of Figures
Chapter I Fig. 1. ICE lifecycle 14
Fig. 2. Genetic maps of ICEBs] and Tn916 16
Fig. 3. ICEBs] gene regulation 19
Fig. 4. Tn916 gene regulation 24-25 Fig. 5. Rolling circle replication of ICEBs] 27 Fig. 6. Relaxse-mediated DNA nicking and 30-31 ligation
Fig. 7 Termination of DNA strand transfer by 32-33 relaxase in the donor or recipient
Fig. 8. The single strand origin of M13 recruits 34 host RNA polymerase
Chapter 2 Fig. 1. Map of ICEBsJ and mutants 49
Fig. 2. A region of ICEBsJ (ssol) is similar to 51
sso's from RCR plasmids
Fig. 3. Visualization of Sso function in living 54 cells.
Fig. 4. The amount of plasmid DNA associated 55 with Ssb-GFP was decreased in the plasmid with
ssol.
Fig. 5. Southern blot analysis demonstrates that 57 ssol decreases the amount of plasmid ssDNA.
Fig. 6. ssol contributes to stable acquisition of 61 ICEBs] by recipients.
Fig. 7. Sso activity is important for maintenance 65 of ICEBsJ after excision in growing cells.
Fig. 8. Model of ICE integration with an sso 72 downstream (A) and upstream (B) of oriT.
Chapter 3 Figure 1. Genetic map of Tn916 and schematic 93 for detecting excision products.
Figure 2. Products generated following excision 98-99 of Tn916 from the chromosome.
Figure 3. The relaxase encoded by or/20 contains 104 a conserved N-terminal helix-turn-helix region.
Figure 4. Alignment of several relaxase 105 homologues.
Figure 5. Ssb-GFP to visualize ssDNA and single 109 strand origin activity.
Figure 6. sso916 reduces the amount of Ssb-GFP 110 bound to plasmid DNA.
Chapter 1
Bacteria can acquire DNA from extracellular sources via horizontal gene transfer (HGT)
processes. HGT diversifies bacterial genomes and facilitates the spread of novel traits, including
resistance to antibiotics, pathogenicity functions, and novel metabolic pathways. Knowledge of
the mechanisms of HGT is relevant to understanding and preventing the spread of enhanced
virulence and drug resistance in many pathogenic bacteria resistances(Amita et al., 2003; Palmer
et al., 2010; Paulsen, I. T. et al., 2003; Sebaihia et al., 2006), and can aid efforts to engineer
bacteria with beneficial activities (Gelvin, 2003; Rauch and De Vos, 1992). My research has
better defined the mechanism, conservation and importance of autonomous replication of
integrative and conjugative elements, a class of horizontally-acquired DNA.
Relevance of HGT
Significant proportions of most bacterial genomes contain horizontally-acquired DNA
(Ochman et al., 2000). Genes on horizontal elements often confers novel phenotypes that allow
bacteria to adapt to certain intracellular and environmental stresses, and to colonize new niches,
including human hosts (Coburn et al., 2007; Li et al., 2012). Importantly, horizontal elements can
mobilize or incorporate non-self-transferable DNA, leading to element diversification and
plasticity.
Virulence and host range capacities of pathogenic bacteria are often associated with
horizontal elements, and transfer of genes conferring antibiotic resistance or other virulence
factors can convert non-pathogenic bacteria to disease-causing agents. The Ti plasmid of
Agrobacterium tumefaciens allows the bacteria to sense and infect damaged plant cells, leading crown gall disease (Gohlke and Deeken, 2014). pINV plasmids of Shigella spp. encode virulence
genes, including toxins and functions that drive epithelial cell invasion and induction of
macrophage apoptosis (Lan et al., 2006; Schroeder and Hilbi, 2008). Horizontal transfer of pINV
plasmid pWR100 to Escherichia coli led to development of pathogenic and invasive phenotypes
in E. coli (Sansonetti et al., 1983). Another intracellular pathogen, Salmonella enterica, harbors a
pathogenicity island derived from HGT that encodes, among other virulence factors, Mg2 and
proton transporters to regulate intracellular homeostasis and promote survival within
macrophages (Groisman et al., 2013). Large proportions of the genomes of vancomycin-resistant
Enterococcusfaecalis strain V583 and multi-drug resistant Clostridium dificile strain 630 are derived from mobile DNA and horizontal elements that encode not only drug resistance but other
virulence phenotypes (Paulsen, I. T. et al., 2003; Sebaihia et al., 2006). Along with the
aforementioned genomic and molecular studies, recent epidemiological data support a direct link
between HGT processes and disease spread (Amita et al., 2003; Li et al., 2012; Palmer et al.,
2010)
Mechanisms of HGT
There are three primary modes of HGT in bacteria (reviewed in (Frost et al., 2005;
Ochman et al., 2000; Thomas and Nielsen, 2005)): transformation, transduction, and
conjugation.
Transformation describes uptake of DNA from the environment (Thomas and Nielsen,
2005). Many bacterial species, including Bacillus subtilis, Streptococcus pneumonia,
Haemophilus influenza, and Neisseria gonorrhoeae are naturally competent (Johnsborg et al., 2007); that is, these species have DNA uptake systems that enable transformation. Transduction
phage, host DNA may be packaged along with the phage genome into the infective phage
particle and delivered to new hosts through subsequent infections. There are two types of
transduction: generalized and specialized. Generalized transduction describes the incorporation
of random parts of the host genome into the phage. Lysogenic phages that integrate into the host
genome can carry out specialized transduction, the packaging of DNA adjacent to the phage
integration site. DNA acquired via transformation or transduction may recombine into the host
genome if there is sufficient sequence homology (Frost et al., 2005; Mell and Redfield, 2014).
Conjugation is the contact-dependent transfer of multi-gene elements between bacteria.
There are two classes of conjugative elements: conjugative plasmids and integrative and
conjugative elements (ICEs; also known as conjugative transposons) (Frost et al., 2005).
Whereas conjugative plasmids are independent replicons, ICEs integrate into the host genome.
Both conjugative plasmids and ICEs transfer as a single-stranded DNA (ssDNA) species via a
conserved Type IV secretion system (TIVSS) that is encoded by the elements (reviewed in
(Alvarez-Martinez and Christie, 2009b)). The lifecycle of ICEs from actinomycetes species is
distinct (Hagege et al., 1993; te Poele et al., 2008); notably, these elements transfer as
double-stranded DNA via a FstK-like ATPase pump (te Poele et al., 2008). For the purposes of my
thesis, I will exclude analysis of ICEs from actinomycetes.
Along with phages and conjugative elements, bacterial genomes contain
non-self-transferable mobile genetic elements (MGEs), including plasmids (Frost et al., 2005),
transposons and insertion sequences (Craig, 1996), integrons (Mazel, 2006), and integrative
mobilizable elements (IMEs) (Boyd et al., 2009; Hentschel and Hacker, 2001). IMEs are discrete
genomic regions derived from HGT that have lost the ability to transfer. Many plasmids and
bacteria, a phenomenon known as mobilization (Bellanger et al., 2011; Pavlovic et al., 2004;
Smillie et al., 2010). Genetic rearrangements between elements, including transposition;
homologous recombination-driven insertions and deletions; and site-specific recombination via
phage and conjugative element recombinases, diversify both host genomes and MGEs, and can
lead to dissemination of otherwise transfer-deficient MGEs (Bellanger et al., 2011; Burrus and
Waldor, 2004; Draper et al., 2005; Johnson and Grossman, 2015; Pavlovic et al., 2004).
Introduction to integrative and conjugative elements (ICEs)
Integrative and conjugative elements represent a diverse and abundant family of bacterial
mobile genetic elements. ICEs range in size from -18 kb to 500 kb, and are found integrated in
the genomes of both Gram-positive and Gram-negative hosts (Burrus and Waldor, 2004; Johnson
and Grossman, 2015). ICEs often reside in a host tRNA gene (Auchtung et al., 2005;
Dimopoulou et al., 2002; Sullivan and Ronson, 1998) or another preferred site in the host
genome (Burrus et al., 2000; Hochhut and Waldor, 1999), but some elements will integrate in
multiple sites (Bedzyk et al., 1992; Mullany et al., 2012; Norgren and Scott, 1991; Rice et al.,
2010). ICEs are modular, consisting of domains encoding recombination genes; regulatory
genes; and transfer genes, including a conjugative relaxase and type IV secretion system
(TIVSS) (Johnson and Grossman, 2015).
Like other horizontally-acquired elements, ICEs often endow selective advantages to
their hosts (reviewed in (Burrus and Waldor, 2004)), including clinically-relevant phenotypes.
ICEs from the SXT/R391family from Vibrio cholerae can carry resistances to multiple drugs
(Ahmed et al., 2005; Constins et al., 2001) and heavy metals (Song et al., 2013), and the spread
is a pathogenicity island and ICE from Pseudomonas aeruginosa that encodes virulence factors, including cup genes used for biofilm formation (Carter et al., 2010; He et al., 2004). ICEs may
also encode functions that benefit industry, agriculture and human health. Lactococcus lactis
strains with Tn5276 are used for production of the antimicrobial nisin and sucrose fermentation
(Rauch and De Vos, 1992). ICEMlsymR7
A from Mesorhizobium loti is a symbiosis island that
facilitates nodule formation in legumes and nitrogen fixation for the host plant (Sullivan and
Ronson, 1998). Genomic studies of other conserved ICE features has identified cryptic ICEs that
may encode novel host benefit functions (Johnson and Grossman, 2015).
Similar to a lysogenic phage, many ICE genes are repressed when the element is
integrated in the chromosome, including the genes required for conjugative transfer. ICE gene
expression is induced in response to specific signals (Johnson and Grossman, 2015), or
stochastically (Minoia et al., 2008), leading to excision of the element from the chromosome.
Many of the signals that induce expression are conserved among multiple ICEs, and ICEs may
respond to multiple signals. ICE induction leads to activation of a recombinase that mediates
excision of the ICE from the chromosome and produces a circular plasmid species. Many ICE
recombinases are site-specific tyrosine integrases, similar to lamda int (Rajeev et al., 2009), but
some ICEs employ serine and DDE recombinases (Johnson and Grossman, 2015). The signals
that induce recombinase expression also lead to activation of the genes required for ICE DNA
processing and transfer, including the relaxase and TIVSS components.
DNA damage can induce gene expression of ICEs, including SXT (Beaber et al., 2004)
and ICEBsJ from B. subtilis (Auchtung et al., 2005). During DNA damage, ssDNA accumulation
activates RecA, which can bind and stimulate autocleavage of proteins such as LexA (reviewed
thermophilus encode lamda cl-like repressors with conserved autocleavage sites (Beaber et al., 2004; Bellanger et al., 2007). In contrast, recA-dependent degradation of the ICEBs1 repressor is
mediated by an element-encoded protease (Bose et al., 2008) (described below).
ICE induction can also occur under high cell density. For example, excision of ICEBsJ,
ICESt3, and ICEMlsymR7
A from Mesorhizobium loti increases during stationary phase (Auchtung
et al., 2005; Carraro et al., 2011; Ramsay et al., 2006; Reinhard et al., 2013). Expression of the
integrase of ICEclc from Pseudomonas knackmussii B 13 is induced during stationary phase
(Reinhard et al., 2013; Sentchilo et al., 2003), but excision and mating predominantly occur upon
renewed nutrient availability (Delavat et al., 2016). Many ICEs also encode signaling molecules
and cognate response regulators that, under high population densities, can sense the presence of
potential recipients (Johnson and Grossman, 2015). ICEBsJ encodes the raplphrI signaling
system (Auchtung et al., 2005) (discussed below). ICEMlsymR7A encodes traR and traIl, which
constititute a N-acyl-homoserine lactone (AHL)-sensing system similar to the quorum sensing
LuxRI system (Ramsay et al., 2009). Lastly, ICE induction can occur in the presence of cargo
gene substrates. ICEclc allows Pseudomonas hosts to utilize 3-chlorobenzoate (3CBA) as a sole
carbon source (Dorn et al., 1974), and 3CBA enhances ICEclc induction (Sentchilo et al., 2003).
Some ICEs that encode resistance to the antibiotic tetracycline are induced in the presence of
drug, including conjugative transposons CTnDOT and CTn341 in Bacteroides species (Parker
and Smith, 2004; Shoemaker and Salyers, 1988; Stevens et al., 1990; Wang et al., 2004) and
Tn916 (Celli and Trieu-Cuot, 1998; Su et al., 1992).
Despite the diversity of ICE auxiliary phenotypes and gene expression regulatory
systems, ICEs share a conserved lifecycle that is summarized in Fig. 1. ICEs integrate in host
transfer and will excise from the host chromosome and form a circular, plasmid species. 2) A
relaxase expressed by the ICE nicks at an origin of transfer oriT in the ICE DNA. We
hypothesize that the relaxase and oriT of most, if not all, ICEs are replicative, and excised ICEs
undergo autonomous rolling circle replication. 3) Relaxase-bound ssDNA is transferred from a
donor to recipient cell via a TIVSS (gray cylinder). 4) ICE ssDNA re-circularizes and replicates.
Double-stranded ICE DNA integrates in the genome of the recipient cell, forming a stable
transconjugant.
recipient transconjugant
00
chromosome
relaxaseICEjL
donor
ocell 1. Excision; 2.Nicking; 3. Transfer 4. Replication;
Gene Replication Integration
expression
Fig. 1. ICE lifecycle. See text for description.
ICEBsJ biology and regulation
ICEBs1 is ~20 kb (see map, Fig. 2A, and Table 1, gene functions) and integrates at a specific site in the tRNA gene trnS-leu2 in the B. subtilis chromosome. ICEBsI is capable of
transfer to other Gram-positive species, including Listeria monocytogenes, Bacillus anthracis,
and Bacillus licheniformis (Auchtung et al., 2005). ICEBsJ encodes a cell-cell sensory system composed of rapI and phrI(Auchtung et al., 2005). Overproduction of rapI leads to induction
and excision of ICEBs1 in over 90% of cells (Auchtung et al., 2005; Lee et al., 2007), facilitating
robust population- and single-cell-level experiments that are infeasible or cumbersome in most
ICE model systems.
A
ICEBsI
coupling
immA ydzL protein yddA hydrolase ydd
mRyd conQ relaxase ydc conC __wT _phr_
imrnR / ydcO c~lQ nicK cw1T ________
intI /xis /7e/P S T / conB 1conD conE yddF conG I J K rap! I yddM
attL attR oriT ssol B Tn916 24 xis \23 22 21 20 19 18 17 16 15 14 13 12 tet(M) 6 910 7 8 5 / int
attL (helPs) coupling (conD) (conE) (conG)
/
(conB) atIRprotein orIT \ sso916 hydrolase
(conQ) relaxase
Fig. 2. Genetic maps of ICEBsI and Tn96. Arrows correspond to open reading frames (ORFs), and the direction of the arrows indicates the direction of transcription. ORF names are written above the
corresponding gene. The ICEBsI homologue of Tn9/6 TIVSS genes is shown in parentheses under the
corresponding Tn9I6 gene. Black rectangles denote attL and attR, direct repeats recognized by the
element's cognate site-specific-recombinase int. oriTs and ssos are indicated with thick black lines under
the corresponding sequence. Functional oriTs were mapped by cloning sequences into a shuttle vector
(Jaworski and Clewell, 1995) or into a mini-ICEBsI backbone (Lee and Grossman, 2007) and
determining the minimal region required for mobilization in trans. Functional ssos were mapped as
described in Ch. 2 and Ch.3. The gene maps are not absolutely to scale. Green: T4SS; red: DNA transfer
and replication proteins; purple: recombinase int and directionality factor xis; orange: cargo genes; blue:
Table 1. ICEBsJ gene function
ORF Functiona Referencesf
Site-specific recombinase
Excision and integration (Lee et al., 2007)
Anti-repressor protease (Bose and Grossman, 2011; Bose et al.,
ImmR degradation 2008)
immR Transcriptional repressor (Auchtung et al., 2007)
Immunity repressor
Recombination-directionality-factor (Lee et al., 2007) Promotes excision via int
Regulation of transcriptionb
ydzL DNA bindingb
ydcO Unknown
Helicase processivity factor
helP Processive unwinding of DNA for (Thomas et al., 2013) replication and mating
conQ Coupling protein ATPase( Conjugation and mobilization
nicK Relaxase for replication and mating (Lee and Grossman, 2007; Lee et al., 2010)
ydcS Unknown
ydcT Unknown
yddA Unknown
Pore formation (VirB8-like)c Conjugation and mobilization
conC Conjugation and mobilization (Leonetti et al., 2015)
conD ConE ATPase localization (VirB3-like) (Berkmen et al., 2010; Leonetti et al., Conjugation and mobilization 2015)
conE ATPase (VirB4-like) (Berkmen et al., 2010; Leonetti et al.,
Conjugation and mobilization 2015) yddF Unknownd
conG Pore (VirB6-like)c
conG Poe(iB-ie'(Leonetti et al., 2015)
Conjugation and mobilization
cwlT Cell wall hydrolase (DeWitt and Grossman, 2014; Fukushima
Conjugation et al., 2008)
yddI Unknowne
yddJ Unknown yddK Unknown
rapI Cell-cell signaling (Auchtung et al., 2005; Bose et al., 2008)
Activates ImmA
phrl Cell-cell signaling (Auchtung et al., 2005)
Inhibits RapI yddM Unknown
Table 1 continued. a Function has been shown experimentally unless otherwise marked with b-f Genes required for ICEBsJ conjugation and plasmid or mini-ICEBsJ mobilization are part of the TIVSS of ICEBsJ. b Gene ontology prediction from (Berkmen et al., 2011). TIVSS gene as predicted in (Alvarez-Martinez and Christie, 2009a). Homologues or analogues from the Ti plasmid of Agrobacterium tumefaciens are indicated in parentheses. d Highly conserved with a domain of unknown function (DUF 1874) found in hypothetical viral and bacterial
proteins(Berkmen et al., 2011). ' May affect conjugation and interact genetically with cwlT (Dewitt, 2013). fRefers to experimental evidence of gene function.
ICEBsJ is induced when its host is subject to DNA damage or when surrounded by a high density of potential recipient cells (Fig. 3). ImmR is a phage-like repressor of ICEBs] genes that
is degraded by ImmA, a protease conserved in some phage and other mobile genetic elements
(Auchtung et al., 2005; Bose et al., 2008; Burrus et al., 2002). ImmA activation occurs in
response to two signals: 1) DNA damage and the recA-dependent SOS response; and 2)
activation of the cell-signaling protein RapI (Bose and Grossman, 2011; Bose et al., 2008).
ICEBsJ encodes cell-cell sensory genes rapI and phrI (Fig. 3). PhrI is a secreted peptide, and PhrI import leads to RapI inhibition and ICEBsJ repression (Auchtung et al., 2005). This
system prevents ICEBsJ transfer to neighboring cells that already contain ICEBsJ. RapI is also
inhibited during exponential growth by AbrB, a regulator of transition-state genes. Nutrient
limitation and high cell density lead to AbrB inactivation and Rap-dependent induction of
DNA damage
I
RecA High density of - PhrlICEBs1+ cells peptide
Signal e Rapi
- ImmA
----*Im
High cell density -i AbrB Protease
/starvation Repressor
Fig. 3. ICEBsI gene regulation. Adapted from (Johnson and Grossman,
positive regulation, and red lines show negative regulation.
ImmR -- ICEBs1 gene Repressor expression
2015). Black arrows denote
111
ImmR represses Pxis, a promoter that drives expression of the
recombination-directionality-factor xis (Auchtung et al., 2007; Lee et al., 2007). Following ImmR degradation,
xis and the recornbinase int (integrase) direct ICEBsI excision (Bose et al., 2008; Lee et al.,
2007). The Pxis operon also encodes the genes required for DNA processing and conjugation of ICEBs1 DNA (Fig. 2A) (Auchtung et al., 2005, 2007). Conjugation requires the DNA transfer and replication genes relaxase nicK and processivity factor helP (Lee and Grossman, 2007; Thomas et al., 2013) (discussed below), as well as the type IV secretion system encoded by the
con genes and cwIT (Fig. 2A) (DeWitt and Grossman, 2014; Lee et al., 2012; Leonetti et al.,
2015). The host translocase pcrA is required for unwinding of the ICEBsI DNA (Thomas et al., 2013), and ICEBs1 cannot transfer out of ApcrA donors (Fig. 1, transfer of ssDNA via the TIVSS) (Lee et al., 2010). Other host genes may be required for ICEBsJ conjugation.
Tn916 biology and regulation
Tn96 is an ~18 kb element originally isolated from pathogenic E. faecalis (see map, Fig. 2B, and Table 2, gene functions) (Franke and Clewell, 1981). Tn96 and Tn916-like
elements have a broad host range among Gram-positive species, including B. subtilis,
Clostridium difficile, and Streptococcus pneumoniae (reviewed in (Clewell et al.; Roberts and Mullany, 2011)). Tn96 integrates in AT-rich regions (Mullany et al., 2012; Scott, 1993), and
can be found in multiple copies within one host cell (Norgren and Scott, 1991). The promiscuity
of Tn96 integration site selection is due to fact that the Tn96 site-specific recombinase Int can
catalyze recombination between non-homologous sequences (reviewed (Rajeev et al., 2009)).
Tn96 encodes tet(M), a gene conferring resistance to tetracycline (Burdett, 1991), and exposure of host cells to tetracycline induces expression of tet(M) and Tn96 excision and conjugation
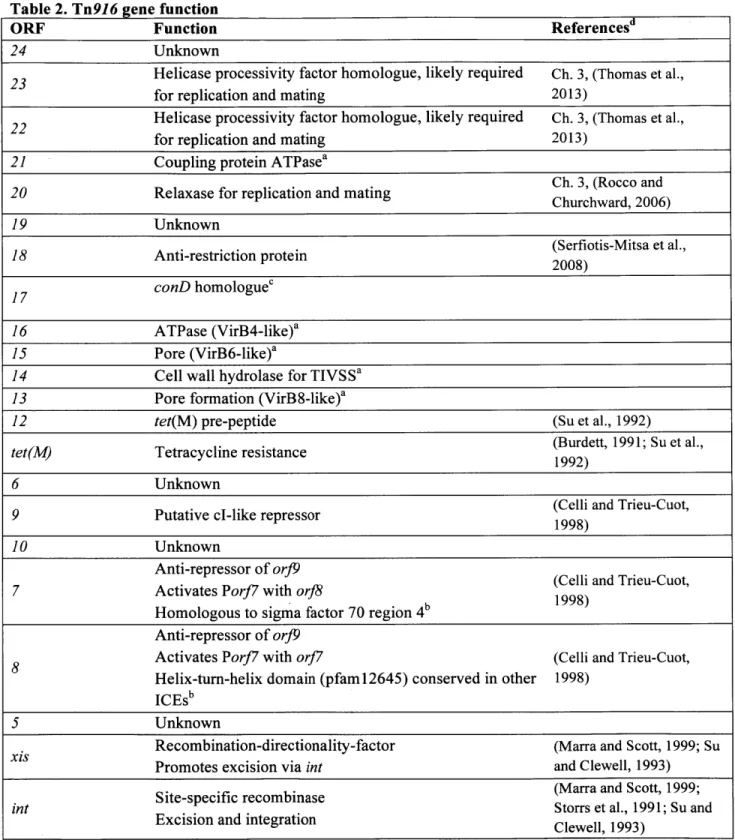
Table 2. Tn916 gene function
ORF Function Referencesd
24 Unknown
23 Helicase processivity factor homologue, likely required Ch. 3, (Thomas et al., for replication and mating 2013)
22 Helicase processivity factor homologue, likely required Ch. 3, (Thomas et al., for replication and mating 2013)
21 Coupling protein ATPasea
Ch. 3, (Rocco and 20 Relaxase for replication and mating Churchward, 2006)
19 Unknown
18 Anti-restriction protein (Serfiotis-Mitsa et al., 2008)
17 conD homologuec
16 ATPase (VirB4-like)a 15 Pore (VirB6-like)a
14 Cell wall hydrolase for TIVSSa 13 Pore formation (VirB8-like)a
12 tet(M) pre-peptide (Su et al., 1992)
tet(M) Tetracycline resistance (Burdett, 1991; Su et al.,
1992)
6 Unknown
9 Putative cl-like repressor (Celli and Trieu-Cuot, 1998)
10 Unknown
Anti-repressor of orJ9 (Celli and Trieu-Cuot, 7 Activates Porf7 with orf8
1998) Homologous to sigma factor 70 region 4
Anti-repressor of orJ9
8 Activates Porf7 with orJ7 (Celli and Trieu-Cuot, Helix-turn-helix domain (pfam12645) conserved in other 1998)
ICEs'
5 Unknown
Recombination-directionality-factor (Marra and Scott, 1999; Su Promotes excision via int and Clewell, 1993)
Site-specific recombinase (Marra and Scott, 1999;
int Storrs et al., 1991; Su and
Excision and integration Clewell, 1993) aTIVSS protein as predicted in (Alvarez-Martinez and Christie, 2009b).Homologues or analogues from the Ti plasmid of Agrobacterium tumefaciens are indicated in parentheses. dPrediction based on a high confidence association with a domain model in the NCBI Conserved Domain Database
(http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). 'Homologous to conD from ICEBsJ, which is predicted to be a VirB3-like protein and, analogous to the ICEBs1 machinery, may be required for localization and function of ATPase orf16 (Berkmen et al., 2011; Leonetti et al., 2015). dRefers to experimental evidence of gene function.
Tetracycline is one of several antibiotics that disrupts ribosome function (Kohanski et al.,
2010). The drug binds to the 30S subunits of the ribosome and inhibits binding of charged
tRNAs, leading to translation inhibition. Tet(M) is a so-called ribosome protection protein
(Roberts and Roberts, 1996), and increasing Tet(M) concentrations leads to tetracycline
displacement from ribosomes in vitro (Burdett, 1996). Tet(M) exhibits GTPase activity (Burdett,
1991), but the exact mechanism of Tet(M)-dependent tetracycline release from ribosomes is
unknown (Burdett, 1996).
Su et al. proposed that tetracycline promotes expression of tet(M) via a unique
attenuation mechanism that is not well understood (Su et al., 1992). orf12 encodes a tet(M)
leader transcript (Fig. 2B), and tetracycline increases RNA polymerase read-through of orf12
and production of a longer transcript encompassing tet(M) (Su et al., 1992). orf12 encodes rare
amino acids and contains several inverted repeats that may form secondary structures, including
a predicted factor-independent terminator. Su et al. proposed that transcription and translation of
orfl2 initiates normally in the absence of drug, but ribosome stalling at rare codons facilitates formation of the orf12 terminator hairpin, terminating transcription. In the presence of
tetracycline, charged-tRNAs, including tRNAs corresponding to the rare codons in orf12,
accumulate, and efficient translation of orf12 prevents formation of the factor-independent
terminator, allowing transcription to proceed through tet(M) (Su et al., 1992). The model predicts
that the effects of tetracycline on Tn916 gene expression are non-specific, and that any antibiotic
that binds ribosomes should increase tet(M) production and promote Tn916 induction.
Although the mechanism underlying tet(M) transcription is ambiguous,
tetracycline-dependent induction of Tn916 is well-documented (Celli and Trieu-Cuot, 1998;
Tn916 induction mechanism is depicted in Fig. 4. orJ9 encodes a putative cl-like repressor, and
or19 transcription decreases in the presence of tetracycline (Celli and Trieu-Cuot, 1998). Notably, one of the tet(M) transcripts encompasses or9, producing an antisense transcript that
could down-regulate or19 expression. The tet(M) transcript also extends through orJ7 and or8,
whose products activate their own promoter, PorJ7, and further repress Por9.
Tetracycline-dependent enhancement of tet(M) transcription thus initiates a positive feedback loop from PorJ7
that drives Tn916 induction.
Por17 drives transcription of int and xis, leading to element excision (Celli and
Trieu-Cuot, 1998). int+ Tn916 generates a large transcript initiating from PorJ7 or Ptet(M) that
encompasses the conjugation genes orJ24-orfl3 (Fig. 2B). Thus excision and circularization
generate a new contiguous operon that drives expression of the conjugation genes (Fig. 4B), and,
unlike ICEBsJ (Lee and Grossman, 2007), excision is required for expression of the conjugation
A2
A
attL
Ptt
et(M)
Pof]x
is tconjugation 12 Porf9
and replication leader T transcript
(-)Tet
+ Tet
B
Fig. 4. Tn9J6 gene regulation.
Ptet(M)
P orf7
Fig. 4 Continued. Promoters are bent arrows. Black arrows represent transcription. ORFs in the gene regulation module are dark gray and ORFs in the conjugation and replication module are light gray. Black rectangles are att sites as described in the legend of Fig. 2.
A) Model for tetracycline regulation of gene expression adapted from (Celli and Trieu-Cuot, 1998). The thickness of the black arrows (transcripts) represents relative abundance in the absence and presence of tetracycline. Not all transcripts are shown. Black arrows align with the relevant promoters, and, in the case of orJ9 and orf12 leader transcript, the termination sites. Without tetracycline (-Tet), transcription initiates from Ptet(M) and terminates at the downstream terminator ("T" shape). PorJ9 drives expression of or9, encoding a putative repressor. In the presence of tetracycline (+ Tet), Ptet(M) drives transcription of longer mRNAs, including orf7 and or8, and mRNA antisense to orJ9. orJ7 and orJ8 together positively regulate PorJ7 (green lines and arrow) and repress PorJ9 (red lines and cross bars). B) Circularization of Tn916 is necessary for expression of conjugation and replication genes (Celli and Trieu-Cuot, 1998). The black arrow represents contiguous transcription from Ptet(M) and PorJ7 through the conjugation genes. Multiple transcripts are generated from excision. The thickness of the arrow does not represent transcript abundance. Adapted from (Johnson and Grossman, 2015).
Autonomous replication of integrative and conjugative elements in the donor
An ICE integrates into its host genome, enabling stable vertical inheritance. However, the
mechanisms that ensure maintenance of the excised element in the donor and recipient were
previously unknown. My thesis work and the work of others show that several ICEs are capable
of autonomous replication (Burrus and Waldor, 2003; Carraro et al., 2015; Lee et al., 2010;
Ramsay et al., 2006; Wang et al., 2001).
ICEBsI replicates autonomously via a rolling circle mechanism (Lee et al., 2010), as depicted in Fig. 5. Rolling circle replication of plasmids and phages initiates when an
element-encoded relaxase nicks one strand of the DNA at a specific site and covalently binds the 5'
phosphate group of the nicked strand (Khan, 2005). Consistent with this model, replication of
ICEBsJ requires the relaxase NicK, which initiates unidirectional replication from the nic site in
oriT (Lee and Grossman, 2007; Lee et al., 2010). Host translocase PcrA acts as a helicase to unwind nicked ICEBsJ (Lee et al., 2010; Thomas et al., 2013). Processive unwinding by PcrA
requires the helicase processivity factor HelP encoded by ICEBs1 (Thomas et al., 2013).
Replication of ICEBs1also requires the host replicative polymerase, which, analogous to other
rolling circle models, initiates replication of the un-nicked strand from the free 3' end at the nic
site. Following unwinding and replication of the un-nicked template, NicK re-circularizes the
nicked strand and generates a ssDNA circle. ICEBsJ encodes at least two single strand origins of
replication that are important for duplication of the ssDNA circle and ICEBsJ replication (Ch. 2).
Host DNA poll, DNA polIII, and DNA ligase complete replication and formation of the second
NicK oriT
000
C EssICEBs1
Nicking at oriT free 3' end 3' Unwinding; DNA synthesis copy #1 Poll sso RNA primersso recruits RNAP
to prime
copy #2
Fig. 5 Rolling circle replication of ICEBsI. NicK, oriT HelP and sso are ICEBs1-encoded factors important for replication (Lee et al., 2010; Thomas et al., 2013; Wright et al., 2015) (see Ch. 2). NicK nicks at oriT and covalently attaches to the DNA. PolIll-driven DNA synthesis initiates from the free 3'. Processive unwinding by PcrA and HeIP separate the DNA strands, facilitating DNA synthesis (Thomas et al., 2013). NicK-driven strand transfer reactions release the first dsDNA copy of ICEBsI and a ssDNA circle. sso is active in its single-stranded form and recruits RNAP polymerase to synthesize an RNA primer (Khan, 1997). Poll initiates DNA replication from the RNA primer. PollIl and ligase complete synthesis of the second copy of ICEBsI. Newly-synthesized DNA is shown as a dotted line.
27
new DNA
HePP
Autonomous replication is required for excised ICEBs1 to be stably maintained in
dividing donor cells (Lee et al., 2010). Replication of ICEBsI may also be important in recipient
cells (Thomas et al., 2013). Notably, replication is not required for ICEBsJ conjugation (Lee et
al., 2010).
Cararro et. al (Carraro et al., 2015) recently showed that ICE R391 replicates
autonomously in Gram-negative E. coli, and that replication is important for element stability.
The relaxase and origin of transfer are important for R391 replication, suggesting that the
element may also use rolling circle replication. The related ICE SXT also appears to be
replicative in E. coli (Burrus and Waldor, 2003). Other ICEs and ICE-like elements may also be
capable of autonomous, plasmid-like replication (Carraro et al., 2016; Ramsay et al., 2006; Wang
et al., 2001).
DNA transfer and replication in the recipient
Relaxase proteins mediate the cleavage (nicking) and ligation of one strand of DNA via
an ATP-independent, topoisomerase I-type mechanism (Chandler et al., 2013; Dempsey et al.,
1992a; Watson et al., 2014) (Fig. 6A). Relaxase proteins contain a catalytic tyrosine residue, and
the hydroxyl (OH) group of tyrosine undergoes nucleophilic attach of the phosphodiester bond in
the DNA backbone, generating a phosphotyrosine bond between the enzyme and the 5' end of
the cleaved DNA (Chandler et al., 2013; Watson et al., 2014). The reaction is reversed when the
free OH group on the 3' end of cleaved DNA or another DNA strand undergoes a nucleophilic
attack on the phosphotyrosine bond, thus regenerating a phosphodiester bond between DNA
strands. Relaxases cleave between specific bases within a relaxase-specific nic site (Chandler et
biochemical and structural data indicate that the hairpin structure and basepair composition
surrounding the nic site are important for recognition of the DNA substrate by the relaxase
(Chandler et al., 2013). Relaxases are highly specific to their cognate nic site; related relaxases
with similar nic sites will not cross-react (e.g., (Dempsey et al., 1992b; Guja and Schildbach,
2015)).
DNA strand exchange, as occurs during rolling circle replication and in the recipient cell
following conjugative DNA transfer, requires the activity of two catalytic tyrosine residues that
act sequentially (Chandler et al., 2013) (Fig. 6B). Conjugative and rolling circle relaxases encode
two catalytic tyrosines (e.g., TrwC of conjugative plasmid R388) or operate as dimers (e.g.,
RepC of rolling circle plasmid pT 181) in order to carry out the DNA strand exchange reactions
required for replication and DNA transfer (Grandoso et al., 2000; Novick, 1989; Rasooly and
Novick, 1993). Recircularization (ligation) of the nicked strand of rolling circle plasmids and
conjugative DNA requires nicking by a second catalytic tyrosine at a second nic site generated
during replication. The resulting free 3' OH group can undergo nucleophilic attack and
A
B
Relaxase c-v H 5' 3' DNA |( V 0 0 0 O- 0-0-+Nicking
0 0 5' p O ...- Osp..-- _ 3' 0 \ | 0- 0-0 5'1 ' O P... O' B...OH 3'0-+
Re-Ligation 0 0 0 I I I 0- 0- 0-nic site Copy 24Nicking
Cl
Re-LigationCl
cOpy1 3'B
Fig. 6 Relaxase-mediated DNA nicking and ligation. The relaxase and relevant residues are shown in blue. Red arrows represent nucleophilic attacks. (A) Mechanism of cleavage (nicking) and re-ligation. The mechanism of catalysis is described in the text. The OH group and benzyl ring of the catalytic tyrosine residue are shown. The long black line represents the sugar-phosphate backbone of a strand of DNA with the 5' and 3' ends shown. A "zoomed in" view of the chemistry is shown below the DNA backbone. Phosphate groups and hydroxyl (OH) groups are shown with short black or blue lines
representing covalent bonds between atoms. The squared "B" represents the deoxyribose and associated base linked together by phosphodiester bonds in the DNA strand. (B) Strand exchange catalyzed by relaxase during rolling circle replication and conjugation requires sequential nicking by two catalytic tyrosines. The relaxase can act as a monomer with two catalytic tyrosines as shown, or can undergo strand exchange using two relaxase molecules. The initial nicking, unwinding, and synthesis of the leading strand as depicted in Fig. 5 is not shown. Generation of a second nic site following leading strand replication facilitates the strand transfer reactions necessary for release of a free ssDNA circle during rolling circle replication (Fig. 5) or after conjugative transfer. The boxed "Y"s represent the catalytic tyrosines. DNA strands are shown as black or gray lines, and the nic site is depicted as two rectangles within the DNA strands, with the bases to the "left" and "right" of the nick shown in different shades. Gray strands were part of the original monomeric plasmid, and the black DNA strand is the leading strand of the second copy of plasmid. The direction of leading strand DNA synthesis is shown by small black arrows.
The relaxase and covalently attached DNA strand transfer to the recipient cell during conjugation (Draper et al., 2005; Garcillan-Barcia et al., 2007; Luo and Isberg, 2004) (Fig. 1; Fig 7). Recircularization (ligation) of the ssDNA can theoretically be catalyzed by relaxase in the donor or recipient (Chandler et al., 2013). Analogous to the replication cycle of rolling circle phage M13, multiple copies of single-stranded ICE DNA attached to one relaxase molecule may be generated and transferred (Fig. 7).
A
Nicking in donorNy
Nicking0'
3' ReLi OnyNoB
Nicking in recipient Nicking Recipient Donor 0 Re-Li ton 3, Recipient Donora
3
F'~ ~
5 53 32 5'Fig. 7 Termination of DNA strand transfer by relaxase in the donor or recipient. Transfer of multimers of covalently-attached ICEs is shown. DNA strands are shown as black or gray lines, and each segment of gray or black represents a full-length copy of single- or double-stranded ICE. The nic site is depicted as two rectangles within the DNA strands, with the bases to the "left" and "right" of the nick shown in different shades. Relevant 5' and 3' ends of the ICE DNA are labeled. The direction of leading strand DNA synthesis in the donor is shown by small black arrows. Second strand synthesis from the sso is shown as a dotted line with the small black arrows showing the direction of DNA synthesis. Following second strand synthesis, the monomers may be resolved by homologous recombination. Relaxase is in blue, and the boxed "Y"s represent the catalytic tyrosines. (A) Termination of strand transfer by relaxase nicking in the donor. (B) Termination of strand transfer by relaxase nicking in the recipient. Note that both scenarios assume that the transfer machinery translocates the free 3' end generated after nicking into the recipient cell.
Rolling circle replicating plasmids and phages that are stable encode a single strand origin of replication (sso) or primase that facilitates priming of the ssDNA circular intermediate (Fig. 5; Fig. 8) (reviewed in (Khan, 1997)). An sso is strand-specific and only active in its single-stranded form . The sso sequence typically includes regions of dyad symmetry that form hairpins when single-stranded. ssos of M13 phage and plasmid pT 181 adopt folded structures that mimic promoters and recruit RNA polymerase (Fig. 8) (Bikard et al., 2010; Birch and Khan,
1992). These origins form single large hairpins, and are characterized as ssoA-type origins based on their structural features. The structural bases of the ssos from pBAA1 (ssoT-type) and
pUB 110 (sso U-type), which consist of multiple hairpins, are also known (Kramer et al., 1999; Seery and Devine, 1993). Mutational analyses show that the nucleotide identity of most of the
sso is not important for function. Instead, the secondary structure appears to direct sso function.
Furthermore, ssos are sometimes functional in certain species and not in others, and host range is in part dependent on recognition by host RNA polymerase (Kramer et al., 1998, 1999; Lorenzo-Diaz and Espinosa, 2009).
RNAP
3
sso
RNAP
primer
S
Fig. 8. The single strand origin of M 13 recruits host RNA polymerase. The model of M 13
sso-mediated recruitment of RNA polymerase (RNAP) is shown. The folded sso mimics a promoter, with the -35 and -10 sites shown. RNAP binding to the sso leads to primer synthesis (green). It is assumed that RNAP is displaced from the DNA when RNAP encounters the loop structure at the end of the sso hairpin. Single strand origins can form other secondary structures that recruit host priming factors. Priming facilitates replication of the ssDNA circle and synthesis of the second copy of a rolling circle plasmid as shown in Fig. 5.
Conjugative and mobilizable plasmids transfer as a ssDNA intermediate that re-circularizes in the recipient cell (Chandler et al., 2013; Fernandez-Lopez et al., 2014), and
efficient plasniid conjugation also requires duplication of the transferred ssDNA (Chatfield et al., 1982; Henderson and Meyer, 1996; Lanka and Barth, 1981; Lorenzo-Diaz and Espinosa, 2009; Wilkins et al., 1985). Conjugative and mobilizable plasmids utilize canonical RNAP-dependent ssos, as well as sites that mimic primosome assembly sites or recruit host primase directly (Bruand et al., 1995; Masai and K, 1996; Nomura et al., 1982; Wilkins and Lanka, 1993). In lieu of sites that recruit the host enzymes, sone conjugative plasmids encode a primase that directs priming of transferred DNA replication (Guiney et al., 1989; Wilkins et al., 1985).
Like plasmid conjugation, ICE transfer results in transfer and circularization of a ssDNA
intermediate (Fig. 1). In the recipient cell, the ICE recombinase catalyzes ICE integration into
the new host genome (Johnson and Grossman, 2015). ICEs were thought to encode an sso or
primase activity to drive replication of the transferred ssDNA because double-stranded DNA is
the substrate for most ICE recombinases (Clewell et al.; Wozniak and Waldor, 2010).
In Ch. 2, I describe the identification and characterization of the first single strand origin in an
ICE, and show that Sso activity is important for stable acquisition of transferred ICEBsJ DNA. Ch. 3 shows that Tn916 replicates autonomously in B. subtilis using a rolling circle mechanism,
References
Ahmed, A.M., Shinoda, S., and Shimamoto, T. (2005). A variant type of Vibrio cholerae SXT element in a multidrug-resistant strain of Vibrio fluvialis. FEMS Microbiol. Lett. 242, 241-247.
Alvarez-Martinez, C., and Christie, P. (2009a). Biological diversity of prokaryotic type IV secretion systems. Microbiol. Mol. Biol. Rev. 73, 775-808.
Alvarez-Martinez, C.E., and Christie, P.J. (2009b). Biological Diversity of Prokaryotic Type IV Secretion Systems. Microbiol. Mol. Biol. Rev. 73, 775-808.
Amita, Chowdhury, S.R., Thungapathra, M., Ramamurthy, T., Nair, G.B., and Ghosh, A. (2003). Class I integrons and SXT elements in El Tor strains isolated before and after 1992 Vibrio cholerae 0139 outbreak, Calcutta, India. Emerg. Infect. Dis. 9, 500-502.
Auchtung, J.M., Lee, C.A., Monson, R.E., Lehman, A.P., and Grossman, A.D. (2005). Regulation of a Bacillus subtilis mobile genetic element by intercellular signaling and the global DNA damage
response. Proc. Natl. Acad. Sci. U. S. A. 102, 12554-12559.
Auchtung, J.M., Lee, C.A., Garrison, K.L., and Grossman, A.D. (2007). Identification and
characterization of the immunity repressor (ImmR) that controls the mobile genetic element
ICEBs1 of Bacillus subtilis. Mol. Microbiol. 64, 1515-1528.
Beaber, J.W., Hochhut, B., and Waldor, M.K. (2004). SOS response promotes horizontal dissemination of antibiotic resistance genes. Nature 427, 72-74.
Bedzyk, L.A., Shoemaker, N.B., Young, K.E., and Salyers, A.A. (1992). Insertion and excision of Bacteroides conjugative chromosomal elements. J. Bacteriol. 174, 166-172.
Bellanger, X., Morel, C., Decaris, B., and Guddon, G. (2007). Derepression of excision of integrative and potentially conjugative elements from Streptococcus thermophilus by DNA damage response:
Implication of a cl-related repressor. J. Bacteriol. 189, 1478-148 1.
Bellanger, X., Morel, C., Gonot, F., Puymege, A., Decaris, B., and Guddon, G. (2011). Site-specific accretion of an integrative conjugative element together with a related genomic island leads to cis mobilization and gene capture. Mol. Microbiol. 81, 912-925.
Berkmen, M.B., Lee, C. a., Loveday, E.K., and Grossman, A.D. (2010). Polar positioning of a
conjugation protein from the integrative and conjugative element ICEBs 1 of Bacillus subtilis. J. Bacteriol. 192, 38-45.
Berkmen, M.B., Laurer, S.J., Giarusso, B.K., and Romero, R. (2011). The Integrative and Conjugative Element ICEBs1 of Bacillus subtilis. In: Bacterial Integrative Mobile Genetic Elements. (Landes Bioscience).
Bikard, D., Loot, C., Baharoglu, Z., and Mazel, D. (2010). Folded DNA in action: hairpin formation and biological functions in prokaryotes. Microbiol. Mol. Biol. Rev. 74, 570-588.
Birch, P., and Khan, S. a (1992). Replication of single-stranded plasmid pT181 DNA in vitro. Proc. Natl.
Acad. Sci. U. S. A. 89, 290-294.
Bose, B., and Grossman, A.D. (2011). Regulation of horizontal gene transfer in Bacillus subtilis by activation of a conserved site-specific protease. J. Bacteriol. 193, 22-29.
Bose, B., Auchtung, J.M., Lee, C.A., and Grossman, A.D. (2008). A conserved anti-repressor controls horizontal gene transfer by proteolysis. Mol. Microbiol. 70, 570-582.
Boyd, E.F., Almagro-Moreno, S., and Parent, M.A. (2009). Genomic islands are dynamic, ancient integrative elements in bacterial evolution. Trends Microbiol. 17, 47-53.
Bruand, C., Ehrlich, S.D., and Janniere, L. (1995). Primosome assembly site in Bacillus subtilis. EMBO J.
14, 2642-2650.
Burdett, V. (1991). Purification and characterization of Tet(M), a protein that renders ribosomes resistant to tetracycline. J. Biol. Chem. 266, 2872-2877.
Burdett, V. (1996). Tet(M)-promoted release of tetracycline from ribosomes is GTP dependent. J. Bacteriol. 178, 3246-325 1.
Burrus, V., and Waldor, M.K. (2003). Control of SXT integration and excision. J. Bacteriol. 185,
Burrus, V., and Waldor, M.K. (2004). Shaping bacterial genomes with integrative and conjugative elements. Res. Microbiol. 155, 376-386.
Burrus, V., Roussel, Y., Decaris, B., and Ge, L. De (2000). Characterization of a Novel Integrative Element , ICE Sti , in the Lactic Acid Bacterium Streptococcus thermophilus. Appl. Environ. Microbiol. 66, 1749-1753.
Burrus, V., Pavlovic, G., Decaris, B., and Guddon, G. (2002). Conjugative transposons: The tip of the iceberg. Mol. Microbiol. 46, 601-610.
Butala, M., Zgur-Bertok, D., and Busby, S.J.W. (2009). The bacterial LexA transcriptional repressor.
Cell. Mol. Life Sci. 66, 82-93.
Carraro, N., Libante, V., Morel, C., Decaris, B., Charron-Bourgoin, F., Leblond, P., and Gu6don, G. (2011). Differential regulation of two closely related integrative and conjugative elements from Streptococcus thermophilus. BMC Microbiol. 11, 238.
Carraro, N., Poulin, D., and Burrus, V. (2015). Replication and Active Partition of Integrative and Conjugative Elements (ICEs) of the SXT/R391 Family: The Line between ICEs and Conjugative Plasmids Is Getting Thinner. PLoS Genet. 11.
Carraro, N., Libante, V., Morel, C., Charron-Bourgoin, F., Leblond, P., and Gu??don, G. (2016). Plasmid-like replication of a minimal streptococcal integrative and conjugative element. Microbiol. (United Kingdom) 162, 622-632.
Carter, M.Q., Chen, J., and Lory, S. (2010). The Pseudomonas aeruginosa pathogenicity island PAPI-1 is transferred via a novel type IV pilus. J. Bacteriol. 192, 3249-3258.
Celli, J., and Trieu-Cuot, P. (1998). Circularization of Tn916 is required for expression of the transposon-encoded transfer functions: Characterization of long tetracycline-inducible transcripts reading through the attachment site. Mol. Microbiol. 28, 103-117.
Chandler, M., de la Cruz, F., Dyda, F., Hickman, A.B., Moncalian, G., and Ton-Hoang, B. (2013). Breaking and joining single-stranded DNA: the HUH endonuclease superfamily. Nat. Rev. Microbiol. 11, 525-538.
Chatfield, L.K., Orr, E., Boulnois, G.J., and Wilkins, B.M. (1982). DNA primase of plasmid ColIb is involved in conjugal DNA synthesis in donor and recipient bacteria. J. Bacteriol. 152, 1188-1195.
Clewell, D.B., Flannagan, S.E., and Jaworski, D.D. Unconstrained bacterial promiscuity:
Coburn, B., Guntram, A.G., and Finlay, B. (2007). Salmonella, the host and disease: a brief review. Immunol. Cell Biol. 85, 112-118.
Constins, O.S.X.T., Hochhut, B., Lotfi, Y., Mazel, D., Shah, M., Woodgate, R., Waldor, M.K., and Faruque, S.M. (200 1). Molecular Analysis of Antibiotic Resistance Gene Clusters in Vibrio cholerae 0139 and Molecular Analysis of Antibiotic Resistance Gene Clusters in Vibrio cholerae 0139 and 01 SXT Constins. Antimicrob. Agents Chemother. 45, 2991-3000.
Craig, N.L. (1996). Transposition. (Washington, D.C.: American Society for Microbiology).
Delavat, F., Mitri, S., Pelet, S., and van der Meer, J.R. (2016). Highly variable individual donor cell fates characterize robust horizontal gene transfer of an integrative and conjugative element. Proc. Natl. Acad. Sci. 201604479.
Dempsey, L. a, Birch, P., and Khan, S. a (1992a). Uncoupling of the DNA topoisomerase and replication activities of an initiator protein. Proc. Natl. Acad. Sci. U. S. A. 89, 3083-3087.
Dempsey, L.A., Birch, P., and Khan, S.A. (1992b). Six amino acids determine the sequence-specific DNA binding and replication specificity of the initiator proteins of the pT 181 family. J. Biol. Chem. 267, 24538-24543.
Dewitt, T.A. (2013). Characterization of a Bifunctional Cell Wall Hydrolase in the Mobile Genetic Element ICEBs 1. Massachusetts Institute of Technology.
DeWitt, T., and Grossman, A.D. (2014). The bifunctional cell wall hydrolase cwlT is needed for conjugation of the integrative and conjugative element ICEBs 1 in Bacillus subtilis and B. anthracis. J. Bacteriol. 196, 1588-1596.
Recombination with the Chromosomal tRNA Leu Gene by the Large Conjugative Haemophilus Resistance Plasmid Site-Specific Recombination with the Chromosomal tRNA Leu Gene by the Large Conjugative Haemophilus Resistance Plasmid. 46, 1602-1604.
Dorn, E., Hellwig, M., Reineke, W., and Knackmuss, H. (1974). Isolation and characterization of a 3-chlorobenzoate degrading Pseudomonad. Arch. Microbiol. 1, 61-70.
Doucet-Populaire, F., Trieu-Cuot, P., Dosbaa, I., Andremont, A., and Courvalin, P. (1991). Inducible transfer of conjugative transposon Tn 1545 from Enterococcus faecalis to Listeria monocytogenes in the digestive tracts of gnotobiotic mice. Antimicrob. Agents Chemother. 35, 185-187.
Draper, 0., Cesar, C.E., Mach6n, C., de la Cruz, F., and Llosa, M. (2005). Site-specific recombinase and integrase activities of a conjugative relaxase in recipient cells. Proc. Natl. Acad. Sci. U. S. A. 102,
16385-16390.
Fernandez-Lopez, C., Bravo, A., Ruiz-Cruz, S., Solano-Collado, V., Garsin, D.A., Lorenzo-Diaz, F., and Espinosa, M. (2014). Mobilizable Rolling-Circle Replicating Plasmids from Gram-Positive Bacteria: A Low-Cost Conjugative Transfer. Microbiol Spectr 2, 8.
Franke, A.E., and Clewell, D.B. (1981). Evidence for conjugal transfer of a Streptococcus faecalis transposon (Tn916) from a chromosomal site in the absence of plasmid DNA. Cold Spring Harb.
Symp. Quant. Biol. 45 Pt 1, 77-80.
Frost, L.S., Leplae, R., Summers, A.O., and Toussaint, A. (2005). Mobile genetic elements: the agents of open source evolution. Nat.Rev.Microbiol. 3, 722-732.
Fukushima, T., Kitajima, T., Yamaguchi, H., Ouyang, Q., Furuhata, K., Yamamoto, H., Shida, T., and Sekiguchi, J. (2008). Identification and characterization of novel cell wall hydrolase CwlT: A two-domain autolysin exhibiting N-acetylmuramidase and DL-endopeptidase activities. J. Biol.
Chem. 283, 11117-11125.
Garcillan-Barcia, M.P., Jurado, P., Gonzalez-Perez, B., Moncalian, G., Fernandez, L.A., and De La Cruz, F. (2007). Conjugative transfer can be inhibited by blocking relaxase activity within recipient cells with intrabodies. Mol. Microbiol. 63, 404-416.
Gelvin, S.B. (2003). Agrobacterium -Mediated Plant Transformation: the Biology behind the Agrobacterium -Mediated Plant Transformation: the Biology behind the " Gene-Jockeying" Tool. Microbiol. Mol. Biol. Rev. 67, 16-37.
Gohlke, J., and Deeken, R. (2014). Plant responses to Agrobacterium tumefaciens and crown gall development. Front. Plant Sci. 5, 155.
Grandoso, G., Avila, P., Cay6n, a, Hernando, M. a, Llosa, M., and de la Cruz, F. (2000). Two active-site tyrosyl residues of protein TrwC act sequentially at the origin of transfer during plasmid R388 conjugation. J. Mol. Biol. 295, 1163-1172.
Groisman, E. a, Hollands, K., Kriner, M. a, Lee, E., Park, S., and Pontes, M.H. (2013). Bacterial Mg 2+ Homeostasis, Transport, and Virulence. Annu. Rev. Genet. 47, 625-646.
Guiney, D.G., Deiss, C., Simnad, V., Yee, L., Pansegrau, W., and Lanka, E. (1989). Mutagenesis of the Tral core region of RK2 by using Tn5: Identification of plasmid-specific transfer genes. J.
Bacteriol. 171, 4100-4103.
Guja, K.E., and Schildbach, J.F. (2015). Completing the specificity swap: Single-stranded DNA recognition by F and R100 Tral relaxase domains. Plasmid 80, 1-7.
Hagege, J., Pernodet, J.L., Friedmann, A., and Guerineau, M. (1993). Mode and origin of replication of pSAM2, a conjugative integrating element of Streptomyces ambofaciens. Mol. Microbiol. 10, 799-812.
He, J., Baldini, R.L., D6ziel, E., Saucier, M., Zhang, Q., Liberati, N.T., Lee, D., Urbach, J., Goodman, H.M., and Rahme, L.G. (2004). The broad host range pathogen Pseudomonas aeruginosa strain PA14 carries two pathogenicity islands harboring plant and animal virulence genes. Proc. Natl.
Acad. Sci. U. S. A. 101, 2530-2535.
Henderson, D., and Meyer, R.J. (1996). The primase of broad-host-range plasmid R 1162 is active in conjugal transfer. J. Bacteriol. 178, 6888-6894.