coli Revealed Involvement of the waaWVL Operon in Biofilm
Formation
Benoit Chassaing,a,b* Estelle Garénaux,a,bJessica Carriere,a,bNathalie Rolhion,a,b* Yann Guérardel,c,dNicolas Barnich,a,b
Richard Bonnet,a,b,eArlette Darfeuille-Michauda,b,e
Clermont Université, UMR 1071 Inserm, Université Auvergne, Clermont-Ferrand, Francea; INRA USC 2018, Clermont-Ferrand, Franceb; Université de Lille 1, Unité de
Glycobiologie Structurale et Fonctionnelle, Villeneuve d’Ascq, Francec; CNRS, UMR 8576, Villeneuve d’Ascq, Franced; Centre Hospitalier Universitaire, Clermont-Ferrand,
Francee
ABSTRACT
Ileal lesions of patients with Crohn’s disease are colonized by adherent-invasive Escherichia coli (AIEC), which is able to adhere
to and to invade intestinal epithelial cells (IEC), to replicate within macrophages, and to form biofilms on the surface of the
in-testinal mucosa. Previous analyses indicated the involvement of the
Epathway in AIEC-IEC interaction, as well as in biofilm
formation, with
Epathway inhibition leading to an impaired ability of AIEC to colonize the intestinal mucosa and to form
bio-films. The aim of this study was to characterize the
Eregulon of AIEC strain LF82 in order to identify members involved in
AIEC phenotypes. Using comparative in silico analysis of the
Eregulon, we identified the waaWVL operon as a new member of
the
Eregulon in reference AIEC strain LF82. We determined that the waaWVL operon is involved in AIEC lipopolysaccharide
structure and composition, and the waaWVL operon was found to be essential for AIEC strains to produce biofilm and to
colo-nize the intestinal mucosa.
IMPORTANCE
An increased prevalence of adherent-invasive Escherichia coli (AIEC) bacteria was previously observed in the intestinal mucosa
of Crohn’s disease (CD) patients, and clinical observations revealed bacterial biofilms associated with the mucosa of CD
pa-tients. Here, analysis of the
Eregulon in AIEC and commensal E. coli identified 12 genes controlled by
Eonly in AIEC. Among
them, WaaWVL factors were found to play an essential role in biofilm formation and mucosal colonization by AIEC. In addition
to identifying molecular tools that revealed a pathogenic population of E. coli colonizing the mucosa of CD patients, these
re-sults indicate that targeting the waaWVL operon could be a potent therapeutic strategy to interfere with the ability of AIEC to
form biofilms and to colonize the gut mucosa.
C
rohn’s disease (CD) and ulcerative colitis (UC) are
multifac-torial diseases that occur in individuals with genetic
predispo-sitions and in whom an environmental or infectious trigger causes
an abnormal immune response (
1
,
2
). Clinical observations show
that bacterial biofilms are associated with the mucosa of
inflam-matory bowel disease (IBD) patients (
3
). The mean density of the
mucosal biofilm is 2-fold higher in IBD patients than in patients
with inflammatory bowel syndrome or controls, and the bacteria
are mostly adherent (
3
). Other lines of evidence suggest that
bac-teria play a role in the onset and perpetuation of IBD (
4
,
5
). Several
independent studies have reported the abnormal presence of
ad-herent-invasive E. coli (AIEC) bacteria in the ileal mucosa of CD
patients (
6–12
). In addition to their ability to adhere, these E. coli
bacteria are able to invade intestinal epithelial cells (IEC). The
adhesion and invasion process of the reference AIEC strain LF82
involves type 1 pili, flagella, outer membrane proteins, outer
membrane vesicles, and long polar fimbriae (
13–17
). In addition,
analysis of the genome sequence of AIEC strain LF82 revealed the
presence of several known virulence genes and four putative
pathogenic islands carrying virulence-related genes (
18
).
We previously reported that the
Epathway plays a crucial role
in AIEC strain LF82 but not in nonpathogenic E. coli K-12
MG1655 by regulating adhesion, invasion, and biofilm formation
processes (
19
). However, the molecular link between
Epathway
activation and AIEC phenotypes is still unclear. The
Efactor, also
called RpoE, is activated by stresses that interfere with the folding
of outer membrane proteins (OMPs) (
20–22
), such as the
osmo-larity encountered in the gastrointestinal tract (
15
). As expected
Received 4 December 2014 Accepted 3 February 2015 Accepted manuscript posted online 9 February 2015
Citation Chassaing B, Garénaux E, Carriere J, Rolhion N, Guérardel Y, Barnich N, Bonnet R, Darfeuille-Michaud A. 2015. Analysis of theEregulon in Crohn’s disease-associated Escherichia coli revealed involvement of the waaWVL operon in biofilm formation. J Bacteriol 197:1451–1465.doi:10.1128/JB.02499-14. Editor: V. J. DiRita
Address correspondence to Benoit Chassaing, chassaingbenoit@yahoo.fr, or Richard Bonnet, richard.bonnet@udamail.fr.
* Present address: Benoit Chassaing, Institute for Biomedical Sciences, Center for Inflammation, Immunity and Infection, Georgia State University, Atlanta, Georgia, USA; Nathalie Rolhion, Institut Pasteur, Unité des Interactions Bactéries-Cellules, Paris, France.
E.G. and J.C. contributed equally to this work.
This article is dedicated to our esteemed mentor and coauthor Arlette Darfeuille-Michaud, who sadly passed away on 28 June 2014 (68).
Supplemental material for this article may be found athttp://dx.doi.org/10.1128 /JB.02499-14.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
from its role in the stress response, the
Eregulon includes genes
that encode periplasmic foldases, proteases, and chaperones that
aid in OMP folding. In addition,
Etranscribes an array of
bio-synthetic enzymes involved in phospholipid, fatty acid,
lipopoly-saccharide (LPS), and membrane-derived oligolipopoly-saccharide
synthe-sis and transport, as well as a number of other cell envelope
proteins, including lipoproteins, inner membrane proteins, and
envelope proteins of unknown function (
23–25
).
The aim of the present study was to decipher the molecular
mechanism of
Epathway involvement in the pathogenesis of
AIEC and to identify AIEC virulence factors with
E-regulated
expression. We report here the involvement of the
E-mediated
pathway in the ability of AIEC strains to form biofilms and to
colonize the intestinal mucosa via transcription of the waaWVL
operon. This operon is transcribed in response to
Epathway
activation, is involved in AIEC lipopolysaccharide synthesis, and
is essential for AIEC strains to produce biofilm and to colonize the
intestinal mucosa.
MATERIALS AND METHODS
Ethics statement. Animal protocols were approved by the Committee for Ethical Issues, CEMEA Auvergne (permit number CE16-0927-2956), and all animals were used in accordance with the European Community guidelines for the care and use of animals (86/609/CEE).
Reference bacterial strains, plasmids, and cell lines. Bacterial strains and plasmids used in this study are listed in Table S1 in the supplemental material. Bacteria were grown routinely in LB broth (BD) overnight at 37°C without shaking. Antibiotics were added to medium at the following concentrations: ampicillin (50g ml⫺1), kanamycin (50g ml⫺1), and chloramphenicol (25g ml⫺1).
When experiments involved RseA/B and RpoE protein overexpres-sion, a concentration of 20 g liter⫺1of arabinose was used; we previously reported that this concentration leads to a strong and reliable decrease in E pathway activity (19). For experiments involving induction of
waaWVL expression (seeFig. 3Band7; see Fig. S1 and S4 in the supple-mental material), a concentration of 5 g liter⫺1of arabinose was used.
Intestine 407 cells (I-407; derived from human intestinal embryonic jejunum and ileum) were purchased from Flow Laboratories, Inc., McLean, VA. Cultured cells were maintained in an atmosphere contain-ing 5% CO2at 37°C in modified Eagle medium (Seromed; Biochrom KG, Berlin, Germany) supplemented with 10% (vol/vol) fetal bovine serum (Lonza), 1% nonessential amino acids (Lonza), 1%L-glutamine (Lonza), 200 U of penicillin, 50g of streptomycin, and 0.25 g of amphotericin B per liter, and with 1% minimal essential medium (MEM) vitamin mix X-100 (Lonza).
Construction of isogenic mutants and trans-complementation as-says. Isogenic mutants were generated with a PCR product using the method described by Datsenko et al. (26) and modified by Chaveroche et al. (27). Primers used are listed in Table S2 of the supplemental material. For trans-complementation assays, a PCR product containing the entire 3,389-bp waaWVL operon was cloned into the pBAD24 vector (28), and the rseAB operon was cloned into the pBAD33 vector, as previously de-scribed (19) (see Tables S1 and S2 in the supplemental material).
E-binding consensus sequence elaboration and determination of
E-regulated genes in AIEC reference strain LF82 and E. coli K-12 strain
MG1655. The HMMER software program (version 3.0;http://hmmer .janelia.org/) was used for the identification of theE-binding motifs within 28 sequences previously reported to be bound by theEprotein (23) (see Table S3 in the supplemental material). HMMER uses probabi-listic models called profile hidden Markov models to identify likely motifs within the input set of sequences. It produces a consensus sequence and a position-specific probability matrix, which provides probabilities associ-ated with each base at each position. The alignedEpromoter sequences were visualized using sequence logo (29) (http://weblogo.berkeley.edu/).
We then applied the HMMER program, using the motif matrix previously determined, to search for the motif in the whole genome of E. coli K-12 strain MG1655 and AIEC reference strain LF82 (18,30). The algorithm in HMMER calculates position z-scores for the motif at each possible posi-tion within a sequence. Only the motif hits with a HMMER score higher than 1 and located less than 1,100 bp from the translation start point were considered putativeE-binding sites (23).
Promoter expression assay. To generate the lacZ fusion promoter, promoters of genes topA, ORF1, ychH, and of operon waaWVL were am-plified by PCR (see Table S2 in the supplemental material). The resulting 360-bp fragments contained the promoter sequence of the corresponding gene/operon as well as the putative RpoE-binding site identified by in
silico analysis. These PCR fragments were then ligated into the plasmid
vector pRS550 (31) and designated pRS550-topA, pRS550-ORF1, pRS550-waaW, and pRS550-ychH, respectively.-Galactosidase activi-ties were analyzed with a-galactosidase assay kit (Qiagen) with strains harboring these pRS550 constructs as well as the pBAD30 empty vector or pBAD30-rpoE in LB culture medium.-Galactosidase activity of each sample was determined by measuring the optical density at 420 nm (OD420) at 24 h, and the number of bacteria in each sample was calculated based on OD620measurements, from which Miller units were deter-mined.
RNA manipulations, reverse transcription (RT), and RT-PCR. Cul-tures were grown at 37°C in LB, LB plus 20 g liter⫺1NaCl, cell culture medium (DMEM plus 10% fetal bovine serum) containing 2% sodium choleate, M9 minimal medium (Invitrogen) supplemented with glucose at 4 g liter⫺1, CaCl2at 0.1 mM, and MgSO4at 2 mM, or M63 minimal medium (U.S. Biological) supplemented with glucose at 8 g liter⫺1and MgSO4at 1 mM. At an OD620of 0.2 and when needed,L-arabinose at 20
g liter⫺1was added to induce the overexpression of RseAB, andL -arabi-nose at 5 g liter⫺1was added to induce the overexpression of WaaWVL. Total RNA was extracted at 4 h, 16 h, 24 h, or from overnight-cultured bacteria and treated with DNase (Roche Diagnostics) to remove contam-inating genomic DNA.
For RNA extraction of biofilm-associated bacteria, strains were grown overnight in Luria-Bertani broth with 5 g liter⫺1of glucose (Euromedex) at 35.5°C, after which 1/100 dilutions were made in M63 minimal me-dium (U.S. Biological) supplemented with 8 g liter⫺1glucose. Fifteen-milliliter aliquots were then placed in wells of non-cell-treated polysty-rene petri plates and incubated at 30°C without shaking. At different time points, plates were washed once, bacteria were harvested using a scraper, and RNAs were extracted as previously described. Bacterial growth in M63 minimal medium supplemented with 8 g liter⫺1(0.8%) glucose was used as a control.
The RNAs were reverse transcribed and amplified using primers spe-cific to rpoE, yjiW, lpfA, ORF1, ORF2, yliF, ygcU, waaWVL, ORF3, gnd,
uidC, yafT, and ychH mRNAs or 16S rRNA (see Table S2 in the
supple-mental material). Amplification of a single expected PCR product was confirmed by electrophoresis on a 2% agarose gel. RT-PCR was per-formed using an Eppendorf Realplex system, and the RNA levels were quantified using RNA master SYBR green I (Roche Diagnostic) with 0.25 g of total RNA.
Adhesion and invasion assay. The bacterial adhesion assay was per-formed as described previously (32). Briefly, intestine 407 cells were seeded in 24-well tissue culture plates with 4⫻ 105cells per well. Mono-layers were then infected at a multiplicity of infection of 10 bacteria per cell in 1 ml of cell culture medium without antibiotics and with heat-inactivated fetal calf serum (FCS; PAA Laboratories). After a 3-h incuba-tion period at 37°C, monolayers were washed three times in phosphate-buffered saline (PBS, pH 7.2). Epithelial cells were then lysed with 1% Triton X-100 (Euromedex) in deionized water. Samples were diluted and plated onto Mueller-Hinton agar plates to determine the number of CFU corresponding to the total number of cell-associated bacteria (adherent and intracellular bacteria). To determine the number of intracellular bac-teria, fresh cell culture medium containing 100g ml⫺1gentamicin was
added for 1 h to kill extracellular bacteria. Monolayers were then lysed with 1% Triton X-100, and bacteria were quantified as described above.
Biofilm formation assays. Biofilm formation assays were performed using a previously described method (33). Strains were grown overnight in Luria-Bertani broth with 5 g liter⫺1of glucose (Euromedex) at 35.5°C, after which 1/100 dilutions were made in M63 minimal medium (U.S. Biological) supplemented with 8 g liter⫺1(0.8%) glucose. Aliquots (130 l) were then placed in wells of non-cell-treated polystyrene microtiter plates and incubated overnight at 30°C without shaking. Afterwards, ODs were read at 630 nm in order to determine bacterial growth. Wells were washed once, adherent bacteria were stained with 1% crystal violet solu-bilized in ethanol, and ODs were read at 570 nm. Specific biofilm forma-tion (SBF) was calculated using the following formula: SBF⫽ (AB ⫺ CW)/G, in which AB is the OD570of the attached and stained bacteria, CW is the OD570of the stained control wells containing only bacterium-free medium (to eliminate unspecific or abiotic OD values), and G is the OD630as a measure of cell growth in broth (34,35). Assays were per-formed in triplicate.
Biofilm formation assays were also performed using paraformalde-hyde (PFA)-fixed intestinal epithelial I-407 cell monolayers. Briefly, con-fluent I-407 monolayers were fixed for 15 min in 4% PFA. After washing, bacterial strains expressing green fluorescent protein (GFP) (36) and di-luted in M63 medium supplemented with 8 g liter⫺1glucose were applied and incubated overnight at 30°C without shaking. For visualization, in-fected epithelial monolayers were fixed again for 15 min in 4% PFA, phal-loidin-tetramethyl rhodamine isothiocyanate (TRITC) was used to visu-alize actin, and Hoechst stain was used to visuvisu-alize nuclei. The slides were examined with a Zeiss LSM 510 Meta confocal microscope.
Image processing via COMSTAT. Images of biofilms at the surface of intestinal epithelial cell I-407 monolayers were analyzed with the com-puter program COMSTAT1 (37). A fixed threshold value was used for all image stacks, and values of roughness and thickness were determined.
Mouse ileal loop experiments. Six-week-old FVB wild-type male mice were starved for 24 h before surgery, with water available ad libitum. They were anesthetized, and their intestines were exteriorized through a midline incision (38). Two or three intestinal segments (about 1 cm) without Peyer’s patches were ligated and inoculated by mixed inocula comprising equivalent numbers (5⫻ 107CFU) of bacteria of two strains. Six hours postinfection, the number of each bacterial strain associated with the mucosa of ligated loops was determined to establish the compet-itive index (CI), which provides a senscompet-itive measurement of the relative degree of attenuation (39). Surgery was performed under ketamine-xyla-zine anesthesia, and all efforts were made to minimize suffering. Mice were killed by cervical dislocation according to animal care procedures.
LPS purification. AIEC strain LF82 (serotyped as O83:H1) and the LF82-⌬waaW::pBAD24-waaWVL mutant were grown overnight at 37°C in 150 ml of Luria-Bertani medium with or without 5 g liter⫺1arabinose. LPS was isolated according to the hot phenol-water procedure (40) with some modifications. Briefly, bacteria were collected by centrifugation, washed twice in PBS containing 0.15 mM CaCl2and 0.5 mM MgCl2, and then disrupted by sonication. To eliminate remaining nucleic acids and proteins, lysates were treated with 200g/ml proteinase K (1 h, 65°C, with gentle mixing) and then with 40g/ml DNase and 80 g/ml RNase (37°C, in the presence of 1l/ml 20% MgSO4and 4l/ml chloroform overnight with gentle mixing). Finally, an equal volume of hot (68°C) 90% phenol was added to the mixtures, followed by vigorous shaking at 68°C for 15 min. Suspensions were then cooled on ice and centrifuged at 8,500⫻ g for 15 min. Aqueous phases were pooled, and phenol phases were reextracted with 10 ml distilled water at 68°C. Pooled aqueous phases were extensively dialyzed against distilled water at 4°C, and purified LPS product was finally lyophilized.
SDS-PAGE analysis. LPS (25g) was separated by SDS-PAGE on a 4% stacking and 15% separating gel and subsequently revealed by either silver staining, periodic acid-Schiff (PAS) staining, or anti-O83 antigen serum immunoblotting. Rabbit antiserum against E. coli LPS O83 was
generously provided by Lothar Beutin (Department of Biological Safety, Robert Koch Institut, Berlin, Germany).
LPS composition analysis by nuclear magnetic resonance. Prior to nuclear magnetic resonance (NMR) spectroscopic analysis, samples were repeatedly exchanged in2H
2O (99.97% purity; Euriso-top, CEA Saclay, France) with intermediate freeze-drying and then dissolved in 500l of D2O (Euriso-top). Chemical shifts were expressed in parts per million downfield from the signal of the methyl groups of acetone. Samples were analyzed in 5-mm tubes, and one-dimensional proton1H experiments were recorded on a Bruker spectrometer at 9.4 T. Assignment of spectra was performed using the Topspin 3.0 program (Bruker Biospin) for spec-tra visualization and overlap.
Statistical analysis. Numerical values were expressed as means with standard errors of the means (SEM). Statistical comparisons were per-formed using a 2-tailed Student t test, unless the variables required a 2-tailed Fisher exact test. A P value less than 0.05 was considered statisti-cally significant.
RESULTS
Identification of the
Eregulon in AIEC strain LF82 and E. coli
K-12 strain MG1655. The
Epathway was previously reported to
be involved in the pathogenesis of AIEC strain LF82 by regulating
adhesion and invasion of intestinal epithelial cells, as well as the
biofilm formation process (
19
). This occurs through regulation of
expression of flagella, type 1 pili, and still-uncharacterized factors
involved in the interaction of AIEC bacteria with host cells (
15
,
19
). Our aim in the present study was to search in the genome of
strain LF82 for still-unknown virulence factors whose expression
is dependent on the
Epathway. To identify specific genes whose
transcription is regulated by
Ein strains LF82 and MG1655, the
HMMER software program (version 3.0;
http://hmmer.janelia
.org/
) was used to define
E-binding consensus motifs by using 28
DNA sequences previously reported to be bound by
E(greA,
yaeT, ygiM, rpoH, fkpA, rpoE, bacA, yggN, yfeY, clpX, yhjJ, yfeK,
ybfG, ddg, yfgM, plsB, mdoG, yhbG, yfjO, rseA, yeaY, htrA, sixA,
dsbC, sbmA, yieE, yraP, and yfgC) (
23
) (see Materials and Methods
and Table S3 in the supplemental material). The
E-binding
con-sensus sequence obtained is presented in
Fig. 1A
and was used to
perform genome-wide predictions of
E-binding consensus sites
within the genomes of AIEC strain LF82 and K-12 strain MG1655.
This method allowed the identification of 53 genes whose
tran-scription is putatively regulated by
Ein strain MG1655 (see Table
S4 in the supplemental material) and of 52 genes whose
transcrip-tion is putatively regulated by
Ein strain AIEC strain LF82 (see
Table S5 in the supplemental material). Importantly, the
compar-ison of these two
Eregulons identified 40 genes commonly
reg-ulated in the two strains, 13 genes specifically regreg-ulated in K-12
strain MG1655 (
Table 1
) and 12 genes specifically regulated in
AIEC strain LF82 (
Table 2
).
In order to confirm the results of this in silico analysis, we
intended to measure mRNA levels of the 12 AIEC-specific genes in
a mutant of AIEC strain LF82 deleted for the
E-encoding gene.
However, as previously reported, such a mutant is lethal in
Esch-erichia coli (
41
,
42
). In order to counteract this methodology
prob-lem, we used the LF82 strain transformed with the pBAD24-rseAB
plasmid, which allows the expression of the anti-sigma factors
RseA and RseB under the control of an arabinose-dependent
pro-moter, preventing
Einteraction with RNA polymerase (
19
,
43
,
44
). As a control, quantification of rpoE mRNA levels showed that
overexpression of RseAB in a growth medium containing 20 g
liter
⫺1arabinose led to decreased expression of the rpoE gene in
the LF82::pBAD24-rseAB construct (0.63-fold
⫾ 0.12-fold
de-crease; P
⫽ 0.045) (
Fig. 1B
). In addition, overexpression of RseAB
also led to decreased expression of the genes yaeT and rpoH, whose
transcription is under the control of a
E-regulated promoter,
thereby validating our strategy of utilizing RseAB overexpression
(
Fig. 1B
). Expression analysis of the 12 genes identified in silico as
having transcription putatively under the control of
Ein LF82
bacteria but not in MG1655 bacteria (yjiW, lpfABCDE, ORF1,
ORF2, yliF, ygcU, waaWVL, ORF3, gnd, uidC, yafT, and ychH)
showed that only three of them presented decreased mRNA
ex-pression after inhibition of the
Epathway: ORF1, specific to LF82
and encoding a hypothetical protein with unknown function;
ychH, encoding a hypothetical inner membrane protein; and gene
waaW from the waaWVL operon, which encodes three enzymes
predicted to be involved in LPS biosynthesis (expression levels
relative to the wild-type (WT) strain of 0.78
⫾ 0.12, 0.61 ⫾ 0.04,
and 0.59
⫾ 0.20, respectively). However, statistical analysis
indi-cated that only the transcription of waaW was significantly
de-creased (P
⫽ 0.039).
Similar experiments were performed in other culture media,
such as LB containing NaCl at 20 g liter
⫺1(previously reported to
lead to
Epathway activation [
15
]), cell culture medium
contain-ing 2% sodium choleate (previously reported to lead to increased
long polar fimbria expression [
45
]), or M63 medium containing 8
g liter
⫺1glucose (medium used for the biofilm formation assay).
Quantification of rpoE mRNA levels showed that overexpression
of RseAB led to decreased expression of the rpoE gene in all the
media used (
Fig. 1C
). Similarly, decreased expression levels of the
genes waaW and ychH were observed under all growth conditions.
Of note, with RseAB overexpression, ORF1 expression was
de-creased only in cell culture medium containing 2% sodium
cho-leate (
Fig. 1C
).
We next performed a
-galactosidase assay (
19
,
31
) in order to
confirm these findings. For this purpose, we cloned DNA
se-quences encompassing putative ORF1, ychH, and waaWVL
pro-moters upstream of a lacZ reporter gene in the pRS550 plasmid
and measured
-galactosidase activity. We found that promoters
of these genes led to low
-galactosidase synthesis, and the analysis
of decreased
-galactosidase synthesis in response to RseAB
over-expression was not viewed as an efficient way to identify
regula-tion of expression of these genes by
E. Instead, we decided to
measured
-galactosidase activity in the lacZ-negative E. coli
strain BW25113, deleted for the RpoE-encoding gene and
com-plemented with the pBAD30 empty vector or with the
pBAD30-rpoE vector (
46
). While this strain is likely to contain suppressor
mutations that counteract the lethality normally observed with
such a deletion, as previously reported (
47
), we utilized it as a way
of analyzing a potential increase of
-galactosidase activity in the
presence versus in the absence of the RpoE protein. Results,
pre-sented in
Fig. 1D
and expressed as the fold variation in strain
FIG 1 (A) Sequence logos ofEpromoter motifs. Motifs were identified upstream of the 28 mapped transcription starts in E. coli K-12 strain MG1655. Sequence
logos (http://weblogo.berkeley.edu/) (29) of the⫺35, ⫺10, and ⫹1 start site motifs and the A/T-rich UP sequences are indicated. (B) mRNA levels of genes belonging to the predictedE-specific regulon of AIEC strain LF82. Results are expressed as the fold variation in the LF82 strain overexpressing the inhibitory
complex RseA-RseB of theEpathway, determined using 20 g liter⫺1arabinose, relative to that of the wild-type strain. 16S rRNA levels were measured as a
reference. Data are means⫾ SEM from three separate experiments. (C) mRNA levels of rpoE, ORF1, waaW, and ychH genes in AIEC strain LF82 grown in LB medium, LB medium plus NaCl 20 g liter⫺1, cell culture medium plus 2% sodium choleate, or M63 medium plus 8 g liter⫺1glucose. Results are expressed as the fold variation in the LF82 strain overexpressing the inhibitory complex RseA-RseB of theEpathway, determined by using 20 g liter⫺1arabinose, relative to that
of the wild-type strain. 16S rRNA levels were measured as a reference. Data are means⫾ SEM from three separate experiments. (D) Activation of topA, ORF1,
waaWVL, and ychH promoters in the BW25113⌬rpoE strain transformed with the pBAD30 empty vector or pBAD30-rpoE vector in the presence of 20 g liter⫺1
arabinose. Data are presented as-galactosidase activity of the BW25113 ⌬rpoE::pBAD30-rpoE strain relative to activity of the BW25113 ⌬rpoE::pBAD30 strain. Data are means⫾ SEM from three separate experiments. *, P ⬍ 0.05; **, P ⬍ 0.01; ***, P ⬍ 0.001.
TABLE 1 Predicted RpoE regulon members of commensal E. coli K-12 strain MG1655 compared to adherent-invasive E. coli strain LF82 Transcription unit Genome location (bp) Presence in AIEC strain LF82 HMMER
score Protein function RpoE promoter sequencea
yfjO 2764666/2764732 ⫺ 5.46 Hypothetical protein from prophage CP4-57
CCTGAACTACGCACCATTGAAGGTGTCTTAAAAAGTAA
ybfG 715860/715926 ⫺ 4.73 Hypothetical protein AAGGAACTTAATATTTAAAAAATGTTCCATACAATTCC
bacA 3202193/3202259 ⫹ 4.44 Phosphatase GTTAAACCAAACGGTTATAACCTGGTCATACGCAGTAG
insH 3651409/3651474 ⫺ 3.96 Transposase GTATGAAAGATTGGTTATCCTGGCCTCTAAAAATTTAA
araE 2980372/2980437 ⫹ 3.69 Arabinose transporter TAATGAACTTTATGAATTTTATCTGCTGTAAAATTAGG
ygcJ 2879889/2879955 ⫺ 3.22 Hypothetical protein TGGTGAACGTTTTGACCAAAAAATCATCGATAAGACAT
yraH 3285300/3285365 ⫺ 2.42 Hypothetical adhesin ATCTTAAAGTTCAGTCTATTTAATGTTCAATGAAATAT
matA 310641/310706 ⫹ 1.34 Hypothetical regulatory protein
ATCGAAAATAATTAAACTTAATCTCGTTTAACCTTTAT
livK 3595759/3595824 ⫹ 1.30 Periplasmic protein AAAGGCACTTTTTTCTGTTTATCTATCAATAAATTCAG
rutR 1073295/1073360 ⫹ 1.14 Transcriptional regulator ATCTAAAATATCTGGTAAAAAGTGGACTAAACGGTCAA
puuA 1359058/1359123 ⫺ 1.12 Protein involved in synthesis of
gamma-Glu-putrescine
AAATGACCTTTATGTTCAATATTTTTTCAATCTAGCAG
yfeN 2522140/2522206 ⫹ 1.12 Outer membrane protein CTTAAACCTTCCGCCATATTGGTAATCGCAGAGACCGC
ydiY 1804272/1804336 ⫹ 1.04 Hypothetical membrane protein
TATTTAAAATTTTGCAGATAAATATATATAAATAAAAA aBoldface portions of sequences represent⫺35 and ⫺10 motifs of the putative RpoE-binding site.
BW25113-
⌬rpoE::pBAD30-rpoE compared to strain
BW25113-⌬rpoE::pBAD30 revealed that expression of RpoE in the presence
of 20 g liter
⫺1arabinose led to transcriptional activation of both
waaWVL and ychH promoters. However, the topA promoter (a
gene that does not harbor any RpoE-binding sequence and was
used as a control) and the ORF1 promoter were not activated
when RpoE was expressed (
Fig. 1D
). Altogether, these data
con-firmed that the waaWVL operon and ychH gene are new members
of the
Eregulon in the AIEC strain LF82.
Involvement of specific
Eregulon members in the ability of
LF82 to interact with host cells. We next addressed the
involve-ment of the genes waaWVL and ychH, which belong to the
Eregulon, in the adhesion and invasion processes of AIEC strain
LF82 by generating isogenic mutants. Of note, we failed to obtain
any mutant deleted for the waaW gene or for the entire waaWVL
operon, suggesting that deletion of waaWVL is lethal in AIEC
strain LF82. To overcome this lethality, strain LF82-
⌬waaW::
pBAD24-waaWVL was constructed, and the phenotype of the
re-sulting construct was analyzed in the absence of arabinose. This
model allowed very low basal expression of all three transcripts,
waaW, waaV, and waaL, as a consequence of leaky expression
from the pBAD promoter, which can counteract lethality (see Fig.
S1 in the supplemental material), and waaW, waaV, and waaL
transcript expression levels were fully restored in the presence of 5
g liter
⫺1arabinose (see Fig. S1). Complementation was performed
with all three genes, since waaWVL is an operon, and the deletion
of waaW was found to also significantly alter the expression of the
genes waaV and waaL (see Fig. S1).
Before studying the ability of
⌬waaW and ⌬ychH mutants to
adhere to and invade IEC, we confirmed that their growth and
viability in cell culture medium were not affected (see Fig. S2 in the
supplemental material). Neither of the two mutants generated
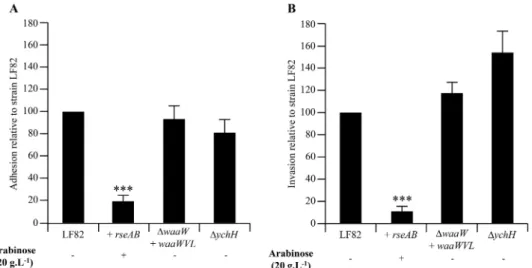
showed any decrease in their ability to interact with IEC (
Fig. 2A
and
B
) compared to AIEC LF82 bacteria overexpressing the
Einhibitory complex RseAB, which had a significantly decreased
abilities to adhere to and to invade IEC, with 19.6%
⫾ 5.9% and
11.4%
⫾ 5.0% residual adhesion and invasion, respectively.
Involvement of
Eregulon members in the ability of LF82 to
form biofilm. Martinez-Medina et al. reported that biofilm
for-mation is a novel pathogenic feature of the AIEC pathovar (
33
),
and we previously demonstrated that the
Epathway is involved
in biofilm formation of AIEC strain LF82 (
19
), but the bacterial
factor(s) involved remains unidentified. To elucidate whether
Eregulon members ychH and waaWVL are involved in AIEC
bio-film formation, we compared biobio-film formation on a plastic
sur-face by wild-type strain LF82, strain LF82 overexpressing RseA
and -B, LF82-
⌬waaW::pBAD24-waaWVL, and ⌬ychH isogenic
mutants. While inhibition of the
Epathway by using 20 g liter
⫺1arabinose led to a significant decrease in the ability of strain LF82
to form biofilms, the isogenic mutant deleted for ychH was not
modified in its ability to form biofilm (
Fig. 3A
). In contrast, the
LF82-⌬waaW::pBAD24-waaWVL strain had a strongly decreased
biofilm formation ability in the absence of arabinose, similar to
that observed with inhibition of the
Epathway (
Fig. 3A
). Of note,
type 1 pilus and flagellum expression levels were unchanged in the
LF82-⌬waaW::pBAD24-waaWVL strain (see Fig. S3 in the
sup-TABLE 2 Predicted RpoE regulon members of adherent-invasive E. coli strain LF82 compared to commensal E. coli K-12 strain MG1655Transcription unit Genome location (bp) Presence in K-12 strain MG1655 HMMER
score Protein function RpoE promoter sequencea
yjiW 4707553/4707620 ⫹ 5.05 Hypothetical endonuclease TGAAATTATGGATTATTTTATAACTCTAAAGAGTCA
lpfABCDE 3761291/3761358 ⫺ 3.55 Long Polar Fimbrae encoding
operon
TCAACTTATTGCAAAAATTAATATTCAGTAAAAATAA ORF 1 1494961/1495026 ⫺ 3.00 Hypothetical protein, function
unknown, with putative DNA-binding homeodomain
CGAAAACTTAAAAAAATAATGCGTCAGATCTGATAAA
ORF 2 2127613/2127679 ⫺ 2.40 Hypothetical protein with unknown function, with signal peptide and 5 transmembrane domains
TTAAAAAATTAGTCCCTTCGATTGTCTCTACAGGTGTT
yliF 826377/826442 ⫹ 1.74 Hypothetical diguanylate cyclase
GGAAATTGGCGAACTATTCCTGGTCTATCAACCGATTG
ygcU 2876616/2876679 ⫹ 1.54 Hypothetical dehydrogenase GGAAATGATTGAAAAACAGGGGGTCGAAGTTGAT
waaWVL 3845291/3845359 ⫹ 1.53 Involved in LPS biosynthesis TGAAATACTGGCCTATAATTTTAAAACAGTAAAAGTAT ORF 3 3121782/3121848 ⫺ 1.51 Hypothetical protein with
unknown function, with signal peptide and 2 transmembrane domains
Gnd 2123143/2123209 ⫹ 1.30 Gluconate-6-phosphate dehydrogenase
AGAAACATTATCAAAATTAAATTTACAAAAAATATAGG
uidC 1670175/1670241 ⫹ 1.24 Hypothetical protein with unknown function, with signal peptide and transmembrane domains
TCAACGATATCACTAGTTAATATTCAATAAAAATAAT
yafT 241096/241161 ⫹ 1.20 Hypothetical aminopeptidase TGCACGTTATGATTTTCATTTTTCTATTGATTTAATG
ychH 1269466/1269531 ⫹ 1.15 Hypothetical inner membrane protein
TGAAATAAGGGTTGTAATTGTGATCACACCCGCACATA aBoldface portions of sequences represent⫺35 and ⫺10 motifs of the putative RpoE-binding site.
plemental material). The defect in biofilm formation of strain
LF82-⌬waaW::pBAD24-waaWVL was fully restored in the
pres-ence of 5 g liter
⫺1arabinose, revealing that complementation of
the LF82-⌬waaW mutant fully restored a WT-like phenotype
(
Fig. 3B
). Overexpression of the waaWVL operon in
nonpatho-genic E. coli strain MG1655 was not sufficient to induce increased
biofilm formation (see Fig. S4 in the supplemental material). In
addition, we observed an increased level of waaWVL mRNA in
LF82 bacteria forming biofilms, compared to planktonic bacteria
grown in the same minimal medium (
Fig. 3C
). The induced
ex-pression of waaWVL during biofilm formation was parallel to that
of the
E-encoding gene (
Fig. 3C
), similar to what we observed
when LF82 bacteria were grown in various minimum media (see
Fig. S5 in the supplemental material), supporting the idea that
WaaWVL factors are the key missing elements involved in
Einhibition pathway-associated phenotypes.
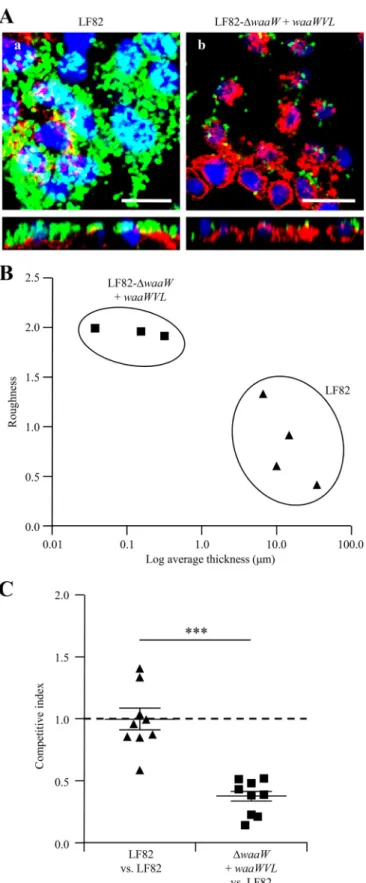
The biofilm formation abilities of strain LF82 and mutant
LF82-⌬waaW::pBAD24-waaWVL were also studied on the
sur-face of PFA-fixed intestinal epithelial cells in the absence of
arabi-nose. Compared to strain LF82, which was able to form a strong
biofilm on the surface of the intestinal epithelial cells, as observed
in the z-section, the mutant LF82-⌬waaW::pBAD24-waaWVL
was unable to produce such a biofilm, with only a few diffusely
adhering bacteria observed on the surfaces of intestinal
epithe-lial cells (
Fig. 4A
). Computational analysis of these biofilm
structures with the computer program COMSTAT1 (
37
)
re-vealed a dramatic 91.3-fold decrease in the biofilm thickness of
the LF82-⌬waaW::pBAD24-waaWVL strain compared to that
of the LF82 wild-type strain (P
⬍ 0.05) (
Fig. 4B
). A 2.3-fold
increase in the roughness coefficient was also observed for the
LF82-
⌬waaW::pBAD24-waaWVL strain compared to the LF82
strain, indicating increased biofilm heterogeneity with
microcolo-nies (
Fig. 4B
).
Finally, the role of WaaW in bacterium-intestinal mucosa
in-teractions was analyzed by using an intestinal ileal loop assay as an
in vivo model. Intestinal ileal loops were inoculated with a mixed
inoculum comprising equivalent numbers of wild-type LF82
bac-teria and LF82-⌬waaW::pBAD24-waaWVL bacbac-teria in the
ab-sence of arabinose, and the strains were compared by competitive
index (CI) analysis, which provided a sensitive measurement of
the relative degree of attenuation (
39
). The analyses of in vitro
cocultures of LF82 wild-type bacteria and LF82-⌬waaW::
pBAD24-waaWVL bacteria in the absence of arabinose revealed
that both strains remained stable over time (see Fig. S6 in the
supplemental material), and the intestinal ileal loop assays
re-vealed that LF82 with inhibition of waaWVL expression had a
mean CI of 0.38
⫾ 0.05, indicating that the depletion of WaaWVL
expression greatly impaired intestinal mucosa colonization (P
⬍
0.0001) (
Fig. 4C
).
waaWVL overexpression in a
Emutant restores a
wild-type-like phenotype. Several observations suggested that WaaWVL
ex-pression is the missing link between
Epathway activation and the
ability of AIEC to form biofilms and to colonize the intestinal
mu-cosa. First, similar phenotypes were observed between strains
im-paired in the
Epathway or in WaaWVL expression (
19
) (
Fig. 3A
).
Moreover, a perfect correlation occurred between
Epathway
ac-tivation and WaaWVL factor synthesis during the biofilm
forma-tion process (
Fig. 3C
). In order to test this hypothesis, we next
analyzed if waaWVL overexpression was able to reverse
pheno-types observed in a
Emutant to “WT-like” phenotypes. For this
purpose, pBAD24-waaWVL was transformed in AIEC strain LF82
overexpressing the
Epathway inhibitors RseA and -B (the rseAB
operon was subcloned into the pBAD33 vector in order to have
antibiotic and replication origin compatibility). waaWVL
overex-pression was found to be sufficient to fully restore a WT-like
phe-notype in AIEC strain LF82 overexpressing the RseAB inhibitory
complex, at both biofilm formation (
Fig. 5A
) and intestinal
mu-cosa colonization (
Fig. 5C
). As previously observed, rseAB
over-expression and/or waaWVL overover-expression had no effect on
bio-film formation or on intestinal mucosa colonization of strain
MG1655 (
Fig. 5B
and
D
).
WaaWVL factors are not involved in biofilm formation
abil-ities of all E. coli strains belonging to the B2 phylogroup. BLAST
analysis (
http://blast.ncbi.nlm.nih.gov
) showed that the waaWVL
operon is also present in two other sequenced AIEC strains (NRG
857C [
48
] and UM146 [
49
]), as well as in urinary pathogenic E.
FIG 2 Involvement of theE-mediated pathway in the ability of LF82 to interact with host cells. Adhesion (A) and invasion (B) abilities of LF82,LF82::pBAD24-rseAB (in the presence of 20 g liter⫺1arabinose), and LF82-⌬waaW::pBAD24-waaWVL and LF82-⌬ychH mutants (in the absence of arabinose) with intestinal epithelial cells (I-407). Each value is the mean⫾ SEM of at least four separate experiments. ***, P ⬍ 0.001.
coli strains CFT073 (
50
), UTI89 (
51
), and 536 (
52
), which are
responsible for urinary tract infections, and in avian pathogenic E.
coli (APEC) strain 01 (
53
) (see Fig. S7A in the supplemental
ma-terial). All these strains, including AIEC LF82, belong to the
phy-logenetic group B2, and sequence analysis revealed that the AIEC
strain LF82 genome is close to those of UPEC and APEC strains
(
18
). Comparison of nucleotide sequences showed that waaWVL
operons shared 99 to 100% homology between these strains (see
Fig. S7B in the supplemental material), suggesting that these
fac-tors may be involved in biofilm formation in UPEC and APEC
strains.
We therefore generated
⌬waaW::pBAD24-waaWVL
con-structs in UPEC strains (CFT073, UTI89, and 536), APEC
strain 01, and commensal E. coli strain HS. We observed that
the abilities of wild-type UPEC, APEC, and commensal strains
to form biofilms were significantly lower than those of AIEC
strains LF82 and LF134 (
Fig. 6
). Growth of these strains was not
affected after waaW depletion (see Fig. S8 in the supplemental
material), and unexpectedly, none of them was affected in
bio-film formation ability after WaaWVL depletion, compared to
AIEC strains LF82 and LF134, for which a significant decrease
in biofilm formation was observed after waaW depletion
(
Fig. 6
). These data suggest that the involvement of the
waaWVL operon in mediating biofilm formation is a
mecha-nism specific to AIEC strains.
WaaWVL depletion leads to modification of the LPS
struc-ture in AIEC strain LF82. Based on previous report describing the
involvement of WaaW, WaaV, and WaaL proteins in LPS
biosyn-thesis (
54–59
), LPS from LF82 and LF82-
⌬waaW::pBAD24-waaWVL strains grown in the absence or in the presence of
arabi-nose were extracted and purified using the hot phenol-water
procedure before silver staining, PAS staining, and anti-O83
im-munoblotting. PAS staining and the use of an antibody directed
toward the O-antigen of LPS revealed a ladder-like pattern of
LPS that ranged in size from 20 to 70 kDa for all three strains
used (
Fig. 7A
to
C
). In the absence of arabinose, the
LF82-⌬waaW::pBAD24-waaWVL mutant exhibited differences in the
LPS pattern compared to the WT AIEC strain LF82, as revealed by
silver and periodic acid-Schiff staining. In particular, we noticed
the accumulation of shorter semirough LPS species (around 20
and 27 kDa) after waaWVL depletion, indicating impaired LPS
synthesis. In agreement with their identification as biosynthetic
intermediates and consequently devoid of polymerized
O-poly-saccharides, these two bands were not revealed by the anti-O83
FIG 3 Involvement of the waaWVL operon in biofilm formation by AIEC strains. (A) SBF index of AIEC strains LF82, LF82::pBAD24-rseAB (in the presence of 20 g liter⫺1arabinose), and LF82-⌬waaW::pBAD24-waaWVL and LF82-⌬ychH mutants (in the absence of arabinose). Data are means ⫾ SEM from three separate experiments. (B) Biofilm formation abilities of AIEC strain LF82 and LF82-⌬waaW::pBAD24-waaWVL mutants in the absence or in the presence of 5 g liter⫺1arabinose. Data are means⫾ SEM from three separate experiments. LF82 wild-type strain was defined as 100%. (C) Activation of the Epathway andwaaW expression in AIEC strain LF82 during the biofilm formation process. The fold variation of rpoE and waaW mRNA levels in wild-type strain LF82 forming
biofilm (at 4 h, 16 h, and 24 h), relative to those of the wild-type strain grown in M63 broth. 16S rRNA levels were measured as a reference. Data are means⫾ SEM from three separate experiments. *, P⬍ 0.05; **, P ⬍ 0.01.
antibody. Importantly, a normal WT-like LPS pattern was
ob-served when the strain LF82-⌬waaW::pBAD24-waaWVL was
grown in the presence of 5 g liter
⫺1arabinose (
Fig. 7A
to
C
),
revealing that complementation of the LF82-⌬waaW mutant fully
recovered the impaired LPS synthesis. These data were
subse-quently confirmed in NMR experiments (
Fig. 7D
). Ring protons
from sugar classically resonate between 3 and 5.5 ppm, and the
superimposition of the three one-dimensional
1H-NMR spectra
of wild-type AIEC strain LF82 and the LF82-
⌬waaW::pBAD24-waaWVL strain (with or without arabinose) indicated differences
in the signal intensities corresponding to sugar molecules. Those
domains of the spectra correspond to beta-anomeric proton spin
systems, indicating that these are beta-monosaccharides (a chain
of
-Glc) mainly affected by the number of glucose repetitions.
NMR signals are quantitative, and we identified a loss of
approx-imately 20% of
-Glc chain length, which can be correlated with
impaired LPS biosynthesis associated with an overall shortening
of the LPS population and accumulation of LPS intermediates in
the LF82-
⌬waaW mutant (
Fig. 7
).
DISCUSSION
Among bacteria adherent to the ileal mucosa of CD patients,
ad-herent-invasive E. coli has been observed (
7
), and we have
identi-fied type 1 pili and flagella as important virulence factors
mediat-ing the interaction of bacteria with intestinal epithelial cells (
13
,
14
). The role of the
Epathway in the ability of E. coli to interact
with intestinal epithelial cells was previously reported for AIEC
strain LF82 (
19
), with the observation that inhibition of this
path-way greatly decreased adhesion and invasion processes (
19
). This
involvement of the
Epathway in adhesion/invasion phenotypes
of AIEC strains was found to be linked to expression of flagella and
type 1 pili. Moreover, inhibition of the
Epathway led to a
de-creased ability of AIEC strain LF82 to form biofilms, which was
previously reported to be another characteristics of AIEC strains
(
33
), and such decreased biofilm formation was not observed with
nonpathogenic E. coli K-12 strain MG1655 (
19
). These findings
suggested that any gene whose transcription is under the control
of
Ecould be involved in biofilm formation by AIEC strains and
that this factor(s) or its
E-dependent expression is absent in K-12
strain MG1655.
In this study, the identification of genes whose transcription is
under the control of the
Efactor and are putatively involved in
biofilm formation as well as in intestinal mucosal colonization was
performed, based on a previous study by Rhodius and
collabora-tors, which identified the
Eregulon in E. coli K-12 strain MG1655
by using a defined
Econsensus fixation domain and in silico
genome screening (
23
). In the present study, we precisely defined
a
Econsensus fixation domain based on the analysis of 28 E. coli
genes known to be
Eregulated, and we subsequently screened the
genomes of AIEC LF82 and E. coli K-12 MG1655 strains in order
FIG 4 Involvement of the waaWVL operon in the AIEC strain LF82 interac-tion with the intestinal mucosa. (A) Confocal analysis of LF82 and LF82-⌬waaW::pBAD24-waaWVL biofilm formation at the surface of a PFA-fixed monolayer of intestinal epithelial I-407 cells in the absence of arabinose. Bac-teria expressing GFP were used. Actin is labeled in red (with phalloidin-TRITC), and nuclei are labeled in blue with Hoescht. Bar, 50m. (B) Rough-ness versus average thickRough-ness of the biofilm structure of LF82 and
LF82-⌬waaW::pBAD24-waaWVL on the surface of PFA-fixed monolayers of intestinal epithelial I-407 cells at 24 h in the absence of arabinose. Images were quantified using the computer program COMSTAT, and each spot represents results from an independent experiment. (C) The CI of the LF82-⌬waaW:: pBAD24-waaWVL strain compared to the LF82 wild-type strain in the absence of arabinose. Intestinal ileal loops were inoculated by mixed inocula compris-ing equivalent numbers of wild-type LF82 and LF82-
⌬waaW::pBAD24-waaWVL bacteria, and the strains were compared by CI analysis. ***, P⬍
to identify the
Eregulon. This screening method led to the
iden-tification of 53 genes in strain MG1655 and 52 genes in strain LF82
whose transcription levels are putatively regulated by
E.
Com-parison of these two sets of genes allowed the identification of 12
genes specific to AIEC strain LF82. Of note, the analysis of the
LF82
Eregulon did not identify any
Econsensus fixation
do-main upstream of fim and fli operons, encoding, respectively, type
1 pili and flagella, nor upstream of intermediate factors involved
in transcriptional activation of these operons, revealing that
addi-tional experiments are needed to decipher the molecular
mecha-nism that lead to decreased expression of type 1 pili and flagella
during
Epathway inhibition. A recent study showed that the
Epathway promotes flagellum expression in Salmonella enterica
se-rovar Typhi under hyperosmotic stress (
60
), but the mechanism
involved has not yet been identified.
Analysis of phenotypes associated with deletion of the 12 AIEC
FIG 5 waaWVL overexpression complementEmutant phenotypes in AIEC strain LF82. (A) SBF index of LF82::pBAD33::pBAD24, LF82::pBAD33-rseAB::pBAD24, LF82::pBAD33::pBAD24-waaWVL, and LF82::pBAD33-rseAB::pBAD24-waaWVL strains in the presence of 20 g liter⫺1arabinose. Data are means⫾ SEM from three separate experiments. The result with the LF82 wild-type strain was defined as 100%. (B) SBF index of MG1655::pBAD33::pBAD24, MG1655:: pBAD33-rseAB::pBAD24, MG1655::pBAD33::pBAD24-waaWVL, and MG1655::pBAD33-rseAB::pBAD24-waaWVL strains in the presence of 20 g liter⫺1 arabinose. Data are means⫾ SEM from three separate experiments. The result with the LF82 wild-type strain was defined as 100%. (C) Competitive index of LF82::pBAD33::pBAD24, LF82::pBAD33-rseAB::pBAD24, LF82::pBAD33::pBAD24-waaWVL, and LF82::pBAD33-rseAB::pBAD24-waaWVL strains compared to the LF82 wild-type strain in the presence of 20 g liter⫺1arabinose. Intestinal ileal loops were inoculated with mixed inocula containing equivalent numbers of bacteria of the two strains, and the strains were compared based on CI analysis. (D) Competitive index of MG1655⫹pBAD33⫹pBAD24,
MG1655::pBAD33-rseAB::pBAD24, MG1655::pBAD33::pBAD24-waaWVL, and MG1655::pBAD33-rseAB::pBAD24-waaWVL strains compared to results with the MG1655
wild-type strain in the presence of 20 g liter⫺1arabinose. Intestinal ileal loops were inoculated by mixed inocula containing equivalent numbers of bacteria of the two strains, and the strains were compared based on CI analysis. **, P⬍ 0.01; ***, P ⬍ 0.001.
strain LF82-specific
Eregulon members showed that none of
them was involved in the adhesion and invasion processes, but
that one of them, the waaWVL operon, plays a crucial role in
biofilm formation. Indeed, the decrease in biofilm formation after
waaW depletion was similar to the decrease observed after
Epathway inhibition. In addition, we used an in vivo intestinal ileal
loop assay model to perform competitive index analysis, as
previ-ously used for the identification of virulence factors in Salmonella
spp. (
39
) and Listeria monocytogenes (
61
), providing a sensitive
measurement of the relative degree of attenuation of wild-type
strains and corresponding mutants. We identified that, after
de-pletion of the WaaW-encoding gene, the presence of AIEC LF82
bacteria on the surface of murine intestinal mucosa was
dramati-cally impaired. Of note, flagellum and type 1 pilus expression
lev-els were unchanged after depletion of WaaW, demonstrating that
these two factors were not involved in the decrease of biofilm
formation and intestinal mucosa colonization observed. The
anal-ysis of biofilm-associated AIEC LF82 bacteria revealed that both
Eand waaWVL expression levels are highly activated during the
biofilm formation process, suggesting that both play a key role in
this process. Importantly, we observed that waaWVL
overexpres-sion fully restores a wild-type phenotype in AIEC strain LF82 that
overexpresses the RseAB inhibitory complex, as shown based on
biofilm formation and intestinal mucosa colonization levels.
Fur-thermore, rseAB overexpression and/or waaWVL overexpression
did not modify the weak ability of K-12 E. coli strain MG1655 to
form biofilms or to colonize the intestinal mucosa.
BLAST and Interproscan analyses revealed that the three
en-zymes encoded by the waaWVL operon are WaaW, an LPS
␣-1,2-galactosyl transferase, WaaV, a
-1,3-glucosyltransferase, and
WaaL, a lipid A-core surface polymer ligase with an O-antigen
ligase-related domain. These three enzymes are predicted to be
involved in LPS biosynthesis and, more specifically, in core
oligo-saccharide and O-antigen biosynthesis (
54
,
55
). Multiple reports
have revealed that modification of LPS composition and/or length
can affect the virulence of pathogenic bacteria, as shown for
Shi-gella flexneri (
62
), and changes in LPS composition or length can
also affect biofilm formation, as reported for Pseudomonas
aerugi-nosa (
63
) and uropathogenic Escherichia coli (
64
).
LPS analysis in AIEC strain LF82 revealed that WaaWVL
de-pletion is associated with accumulation of shorter semirough LPS
species, indicating impaired LPS synthesis. These data correlate
with the previously reported functions of WaaW, WaaV, and
WaaL proteins in LPS biosynthesis (
54–59
), and they indicate that
these modifications of LPS length and structure might be the cause
of the phenotypes observed with the LF82-⌬waaW mutant, as
previously suggested for E. coli (
65
) and for Candida albicans (
66
).
Based on our original finding that waaWVL disruption leads to
lethality in various AIEC strains, but can be easily depleted
with-out any associated lethality in nonpathogenic E. coli strains as well
as in UPEC and APEC strains (in which WaaWVL enzymes were
found to not have any effect on biofilm formation [this report and
reference
58
]), we hypothesize that WaaWVL enzymes may also
play a role on AIEC biofilm formation in an AIEC-specific and
LPS-independent manner. Such AIEC-specific involvement of
WaaWVL enzymes in biofilm formation could indeed explain
why waaWVL depletion leads to an altered biofilm formation
abil-ity only in AIEC strains, for example, by regulating AIEC-specific
factor expression or membrane anchorage, resulting in an altered
biofilm formation ability.
BLAST analysis indicated that the waaWVL operon is present
in various E. coli strains belonging to the B2 phylogroup, including
the sequenced CD-associated AIEC strains NRG857C and
UM146, APEC strain 01, and UPEC strains 536, UTI89, and
CFT073. Compared to the wild-type UPEC strain UTI89, a
⌬rpoE
⌬rseAB mutant was reported to be less able to form biofilms (
67
).
In the present study, we demonstrated that WaaWVL factors were
not involved in APEC and UPEC strains biofilm formation, since
depletion of the waaWVL operon in APEC and UPEC strains did
not lead to any modification of biofilm production. We also
ob-served that depletion of the waaWVL operon did not modify the
biofilm formation ability of nonpathogenic E. coli strain HS, as
FIG 6 The waaWVL operon is not involved in biofilm formation of UPEC, APEC, and HS strains. SBF indexes of AIEC strains LF82 and LF134, UPEC strains CFT073, 536, and UTI89, APEC strain 01, and commensal E. coli strain HS and of the⌬waaW::pBAD24-waaWVL constructions in the absence of arabinose. Data are means⫾ SEM from four separate experiments. **, P ⬍ 0.01; ***, P ⬍ 0.001.previously described for waaL in commensal E. coli (
65
). One
explanation of such a finding is that the waaWVL operon might
not be efficiently expressed in these strains, which would explain
why they are less able to form biofilms than AIEC strain LF82.
However, when additional biofilm experiments were conducted
with overexpression of a cloned waaWVL operon in these strains,
we failed to identify any increased biofilm formation (data not
shown), suggesting that WaaWVL-mediated biofilm formation in
AIEC strain is a specific mechanism.
As we previously reported, specific activation of the
Epath-way occurs in AIEC strain LF82 during adhesion to intestinal
ep-ithelial cells as well as during biofilm formation processes. The
FIG 7 Analysis of purified LPS from AIEC strain LF82 and the LF82-⌬waaW ⫹ pBAD24-waaWVL mutant in the absence or presence of 5 g liter⫺1arabinose.(A to C) Electrophoretic profile of purified LPS separated on a 15% SDS-PAGE gel and revealed by silver staining (A), periodic acid-Schiff staining (B), or anti-O83 immunoblotting (C). (D) Proton NMR spectra of LPS oligosaccharide preparations from AIEC strain LF82 and the LF82-⌬waaW::pBAD24-waaWVL mutant in the absence or presence of 5 g liter⫺1arabinose. Arrows indicate signal differences between the various strains.
new data presented here demonstrate that the waaWVL operon,
whose transcription (which is
Edependent) is activated during
bacterial interaction with intestinal epithelial cells as well as
dur-ing biofilm formation, plays an important role in the ability of
bacteria to form biofilms and to colonize the intestinal mucosa
and might play a role in AIEC colonization of the intestinal
mu-cosa in CD patients.
ACKNOWLEDGMENTS
This study was supported by the Ministère de la Recherche et de la Tech-nologie, Institut National de la Santé et de la Recherche Médicale and the Université d’Auvergne (UMR Inserm 1071), the Institut National de la Recherche Agronomique (USC INRA 2018), and grants from the Associ-ation F. Aupetit (AFA).
We thank Dietrich H. Nies for providing the BW25113⌬rpoE isogenic mutant, Lothar Beutin for providing rabbit antiserum against E. coli LPS O83, Andrew T. Gewirtz (Georgia State University, Atlanta, GA) for help-ful discussions, Amanda R. Arnold and Hao Q. Tran (Georgia State Uni-versity, Atlanta, GA) for critically reading the manuscript, and the CICS platform for confocal microscopy.
We have no financial conflicts of interest.
REFERENCES
1. Kaser A, Zeissig S, Blumberg RS. 2010. Inflammatory bowel disease. Annu Rev Immunol 28:573– 621. http://dx.doi.org/10.1146/annurev -immunol-030409-101225.
2. Stappenbeck TS, Rioux JD, Mizoguchi A, Saitoh T, Huett A, Darfeuille-Michaud A, Wileman T, Mizushima N, Carding S, Akira S, Parkes M, Xavier RJ. 2011. Crohn disease: a current perspective on genetics, au-tophagy and immunity. Auau-tophagy 7:355–374.http://dx.doi.org/10.4161 /auto.7.4.13074.
3. Swidsinski A, Weber J, Loening-Baucke V, Hale LP, Lochs H. 2005. Spatial organization and composition of the mucosal flora in patients with inflammatory bowel disease. J Clin Microbiol 43:3380 –3389.http://dx .doi.org/10.1128/JCM.43.7.3380-3389.2005.
4. Sartor RB. 2008. Microbial influences in inflammatory bowel diseases. Gastroenterology 134:577–594. http://dx.doi.org/10.1053/j.gastro.2007 .11.059.
5. Chassaing B, Darfeuille-Michaud A. 2011. The commensal microbi-ota and enteropathogens in the pathogenesis of inflammatory bowel diseases. Gastroenterology 140:1720 –1728.http://dx.doi.org/10.1053 /j.gastro.2011.01.054.
6. Darfeuille-Michaud A, Neut C, Barnich N, Lederman E, Di Martino P, Desreumaux P, Gambiez L, Joly B, Cortot A, Colombel JF. 1998. Presence of adherent Escherichia coli strains in ileal mucosa of patients with Crohn’s disease. Gastroenterology 115:1405–1413.http://dx.doi.org /10.1016/S0016-5085(98)70019-8.
7. Darfeuille-Michaud A, Boudeau J, Bulois P, Neut C, Glasser AL, Bar-nich N, Bringer MA, Swidsinski A, Beaugerie L, Colombel JF. 2004. High prevalence of adherent-invasive Escherichia coli associated with ileal mucosa in Crohn’s disease. Gastroenterology 127:412– 421.http://dx.doi .org/10.1053/j.gastro.2004.04.061.
8. Martinez-Medina M, Aldeguer X, Lopez-Siles M, Gonzalez-Huix F, Lopez-Oliu C, Dahbi G, Blanco JE, Blanco J, Garcia-Gil LJ, Darfeuille-Michaud A. 2009. Molecular diversity of Escherichia coli in the human gut: new ecological evidence supporting the role of adherent-invasive E. coli (AIEC) in Crohn’s disease. Inflamm Bowel Dis 15:872– 882.http://dx.doi .org/10.1002/ibd.20860.
9. Martin HM, Campbell BJ, Hart CA, Mpofu C, Nayar M, Singh R, Englyst H, Williams HF, Rhodes JM. 2004. Enhanced Escherichia coli adherence and invasion in Crohn’s disease and colon cancer. Gastroenter-ology 127:80 –93.http://dx.doi.org/10.1053/j.gastro.2004.03.054. 10. Baumgart M, Dogan B, Rishniw M, Weitzman G, Bosworth B,
Yantiss R, Orsi RH, Wiedmann M, McDonough P, Kim SG, Berg D, Schukken Y, Scherl E, Simpson KW. 2007. Culture independent analysis of ileal mucosa reveals a selective increase in invasive
Esche-richia coli of novel phylogeny relative to depletion of Clostridiales in
Crohn’s disease involving the ileum. ISME J 1:403– 418.http://dx.doi.org /10.1038/ismej.2007.52.
11. Sasaki M, Sitaraman SV, Babbin BA, Gerner-Smidt P, Ribot EM, Garrett N, Alpern JA, Akyildiz A, Theiss AL, Nusrat A, Klapproth JM. 2007. Invasive Escherichia coli are a feature of Crohn’s disease. Lab Invest 87:1042–1054.http://dx.doi.org/10.1038/labinvest.3700661.
12. Eaves-Pyles T, Allen CA, Taormina J, Swidsinski A, Tutt CB, Eric Jezek G, Islas-Islas M, Torres AG. 2007. Escherichia coli isolated from a Crohn’s disease patient adheres, invades, and induces inflammatory responses in polarized intestinal epithelial cells. Int J Med Microbiol 298:397– 409.
http://dx.doi.org/10.1016/j.ijmm.2007.05.011.
13. Barnich N, Boudeau J, Claret L, Darfeuille-Michaud A. 2003. Regula-tory and functional co-operation of flagella and type 1 pili in adhesive and invasive abilities of AIEC strain LF82 isolated from a patient with Crohn’s disease. Mol Microbiol 48:781–794. http://dx.doi.org/10.1046/j.1365 -2958.2003.03468.x.
14. Boudeau J, Barnich N, Darfeuille-Michaud A. 2001. Type 1 pili-mediated adherence of Escherichia coli strain LF82 isolated from Crohn’s disease is involved in bacterial invasion of intestinal epithelial cells. Mol Microbiol 39:1272–1284. http://dx.doi.org/10.1111/j.1365-2958.2001 .02315.x.
15. Rolhion N, Carvalho FA, Darfeuille-Michaud A. 2007. OmpC and theE
regulatory pathway are involved in adhesion and invasion of the Crohn’s disease-associated Escherichia coli strain LF82. Mol Microbiol 63:1684 – 1700.http://dx.doi.org/10.1111/j.1365-2958.2007.05638.x.
16. Rolhion N, Barnich N, Bringer MA, Glasser AL, Ranc J, Hebuterne X, Hofman P, Darfeuille-Michaud A. 2010. Abnormally expressed ER stress response chaperone Gp96 in CD favours adherent-invasive Escherichia coli invasion. Gut 59:1355–1362.http://dx.doi.org/10.1136/gut.2010.207456. 17. Chassaing B, Rolhion N, Vallee A, Salim SY, Prorok-Hamon M, Neut C, Campbell BJ, Soderholm JD, Hugot JP, Colombel JF, Darfeuille-Michaud A. 2011. Crohn disease-associated adherent-invasive E. coli bac-teria target mouse and human Peyer’s patches via long polar fimbriae. J Clin Invest 121:966 –975.http://dx.doi.org/10.1172/JCI44632.
18. Miquel S, Peyretaillade E, Claret L, de Vallee A, Dossat C, Vacherie B, Zineb el, Segurens HB, Barbe V, Sauvanet P, Neut C, Colombel JF, Medigue C, Mojica FJ, Peyret P, Bonnet R, Darfeuille-Michaud A. 2010. Complete genome sequence of Crohn’s disease-associated adherent-invasive E. coli strain LF82. PLoS One 5:e12714.http://dx.doi.org/10.1371 /journal.pone.0012714.
19. Chassaing B, Darfeuille-Michaud A. 2013. TheEpathway is involved in
biofilm formation by Crohn’s disease-associated adherent-invasive
Esch-erichia coli. J Bacteriol 195:76 – 84.http://dx.doi.org/10.1128/JB.01079-12. 20. Mecsas J, Welch R, Erickson JW, Gross CA. 1995. Identification and characterization of an outer membrane protein, OmpX, in Escherichia coli that is homologous to a family of outer membrane proteins including Ail of Yersinia enterocolitica. J Bacteriol 177:799 – 804.
21. Missiakas D, Betton JM, Raina S. 1996. New components of protein folding in extracytoplasmic compartments of Escherichia coli SurA, FkpA and Skp/OmpH. Mol Microbiol 21:871– 884.http://dx.doi.org/10.1046/j .1365-2958.1996.561412.x.
22. Rouviere PE, De Las Penas A, Mecsas J, Lu CZ, Rudd KE, Gross CA. 1995. rpoE, the gene encoding the second heat-shock sigma factor, sigma E, in Escherichia coli. EMBO J 14:1032–1042.
23. Rhodius VA, Suh WC, Nonaka G, West J, Gross CA. 2006. Conserved and variable functions of theEstress response in related genomes. PLoS
Biol 4:e2.http://dx.doi.org/10.1371/journal.pbio.0040002.
24. Dartigalongue C, Missiakas D, Raina S. 2001. Characterization of the
Escherichia coli sigma E regulon. J Biol Chem 276:20866 –20875.http://dx .doi.org/10.1074/jbc.M100464200.
25. Rezuchova B, Miticka H, Homerova D, Roberts M, Kormanec J. 2003. New members of the Escherichia coliEregulon identified by a
two-plasmid system. FEMS Microbiol Lett 225:1–7.http://dx.doi.org/10.1016 /S0378-1097(03)00480-4.
26. Datsenko KA, Wanner BL. 2000. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97:6640 – 6645.http://dx.doi.org/10.1073/pnas.120163297. 27. Chaveroche MK, Ghigo JM, d’Enfert C. 2000. A rapid method for
effi-cient gene replacement in the filamentous fungus Aspergillus nidulans. Nucleic Acids Res 28:E97.http://dx.doi.org/10.1093/nar/28.22.e97. 28. Guzman LM, Belin D, Carson MJ, Beckwith J. 1995. Tight regulation,
modulation, and high-level expression by vectors containing the arabi-nose PBADpromoter. J Bacteriol 177:4121– 4130.