A Sequence in
M13
Phage
Detects
Hypervariable
Minisatellites
in Human
and Animal DNA
GILBERT VASSART, MICHEL GEORGES, RITA MONSIEUR,
HUGUETrE BROCAS,
ANNE
SOPHIE
LEQUARRE, DANIEL CHRISTOPHE
The term "DNA fingerprint" has been used to describe the extensive restriction
fragment length polymorphism associated with hypervariable minisatellitespresent in
thehuman genome.Untilnow, it was necessary tohybridizeSouthern blots tospecific
probes cloned from human genomic DNA in order to obtain individual-specific restriction patterns.The presentstudydescribes the surprising finding that the
insert-free,wild-type M13 bacteriophage detects hypervariableminisatellitesin human andin animalDNA, provided no competitor DNA is used during hybridization.Theeffective sequence in M13 was traced to two dustersof15-base pair repeats within the protein III gene of the bacteriophage. This unexpected use of M13 renders the DNA
fingerprinting technologymorereadilyavailable to molecular biology laboratories. E XTENSIVE RESTRICTION FRAGMENT
length polymorphism (RFLP)
asso-ciated with moderately repeated se-quences has recently been described in the
human genome (1-4). The hypervariable
nature of these "minisatellites" has been exploited as a meansof developinga
power-fiul
fingerprinting technique withapplica-tions inpaternitytesting,forensicmedicine, andmapping ofthe human genome (5, 6). The probes used in such determinations were originally derived from segments of
DNA discovered by chance within anony-mous (3)orgene-associated piecesof
geno-mic DNA (1, 4, 7). We report here that a
sequence present intheprotein IIIgene of the widely used M13 bacteriophage vector
(8) allows detection of a distinct set of
hypervariable minisatellites in human and animal DNA.
In the course ofa systematic search of RFLPs associated with the thyroglobulin
gene, we observed that a set of probes correspondingtoDNA segmentssubcloned in the 'bacteriophage M13 gave different resultswhen used in the classical hybridiza-tion mixture[Denhardt's medium
plus
her-ring sperm DNA (9)] or in a milk-based cocktail[Blottomedium(10)].Acomplicat-ed andhighlypolymorphic patternwas
sys-B C
10 11 1 2 3
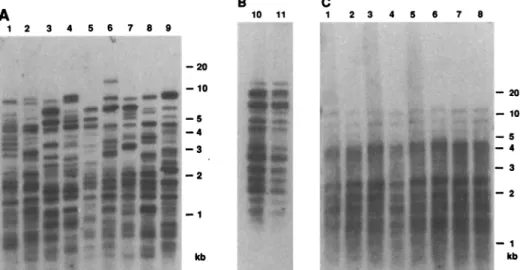
tematically observed with the enzyme Hae III in the lattercondition. It soon became evident thatthe pattern was unrelated to the nature of theinsert and thatwild-type M13 bacteriophages gave the same results. Nine unrelatedindividuals displayeddifferent pat-tems while monozygotic twins were indis-tinguishable (Fig. 1, A and B). Similar blots hybridized with an M13 probe in the pres-ence of herring sperm DNA showed no
polymorphism (Fig. 1C).
The logical explanation for this finding
wasthat a segment in M13hybridized to a
hypervariable minisatellite that could be competed forby fish DNA.If this were true,
G. Vassart, R. Monsieur, H. Brocas, D. Christophe,
InstitutdcRechercheInterdisciplinaire, Universite Libre
de Bruxelles, Campus Erasme, RoutedeLennik808, 1070 Bruxelles.
M.Georges andA. S. Lequarre, Departementde
G&nEti-que, Faculte de Medecine Vt&rinaire, Universite de
Lige,ruedesVeterinaires45, 1070Bruxelles.
A 1 2 3 0: 0000;; CEdDD; 4 5 6 7 8 5 6 B 4 5 6 4940 00 f00: 0 4334 3975 2895 1I*: 2289 1809 952 -20 -10 -5 -4 -3 -2 -1 kb
Fig. 1.Hypevariable polymorphic patternsdetected in human DNAwithwild-type M13 DN probe. Individuals 1to9areunrelatedCaucasians; 10 and 11aremonozygotic twins. Hae III-dige DNA(5,ug) from white blood cellswasseparated ina1% agarosegelin40 mMtris-acetate, 1 EDTA,pH 7.7,ataconstantvoltage of 9 V/cm for about 4 hours. The gelsweresoaked in 1.5MN
0.5M NaOH(2 x 30minutes) followed by equilibrationin IMNH4acetate,0.03MNaOH(2 minutes)and transferovernightontonitrocellulose filters(Schleicher &Schuell)in thesamesolui Single-stranded M13 (200 ng) was labeled with "2P-dATP to a specific activity of 0.5x
10-1.0x 10-9 cpm per microgram of DNA (13). After a 2-hour prehybridization, the filters X hybridized overnightat42°C in the following solutions: (A and B) 40%formamide, 6x SSC (stan4
salinecitrate), EDTA 5 mM, 0.25%dried skimmedmilk; (C)samemediumexceptthat thepowd
milkwasreplaced by 5x Denhardt'sanddenaturedherringspermDNA(100,ug/ml). The filters X allwashed under identical conditions: twice for 15 minutes in 2xSSC,0.1%SDS,roomtemperat four times for 15minutesin 2x SSC,0.1%SDS,65°C;and twice for 30minutesin1x SSC, 6
FilterswereexposedtoKodak XAR-5filmsat-70°C withanintensifyingscreen.
- 20
Fig. 2. IdentificationofM13sequences
hybridiz--10 ingtoherringspermDNA. M13RF DNAwas
5 digested with Xmn I (lanes 1 and 4), BstNI(lanes
4 2and5), and ClaI(lanes3 and6).Two identical
blots were prepared and hybridized to a total 3 M13probe (A)ortoM13sequencesselectedby
2 hybridization toherring spermDNA(B). M13
RFDNA(200 ng)wasdigestedwith restriction endonucleases and identical blotswere prepared asdescribed(Fig. 1). (A) The filterwas
hybrid--1 izedto atotal P-labeled M13
probe.
(B)
The kb filter washybridized
to labeled M13 sequencesthat had beenselectedby hybridizationtoherring
sperm DNA. A 100-cm2 nitrocellulose filterwas
Aas saturated withherringspermDNAby soakingit
ested for 30 minutesinto NH4Ac lM-NaOH 0.03M mM containing 2.5 mg/mil denatured herringsperm
ZaCl, DNAand bakedat80°C for 2 hours. The filter
x30 wasprehybridizedfor2hours in 40%formamide,
tion. 6x SSC,5mMEDTA, 0.25%powdered milkat
9to 42°C, and hybridized overnight toa total
32P-were labeled M13 DNA probe. After washing, the
dard affinity-purified M13 sequences were eluted by lered boiling the filterfor 10 minutes in 10 ml of 10 were mMtris HCIpH 8, 1mMEDTA. About 0.1% of
ture; the inputradioactivitywasrecovered. Hybridiza-5°C. tion and washing offilterswas asdescribed inFig.
1A. 6 FEBRUARYI987 A 1 2 3 4 REPORTS 683 on May 3, 2009 www.sciencemag.org Downloaded from
A 6862 61 B 1833 1 I GAGGGTGGTGGCTCT 6433x/ \ 2 \GAGGGTGGCGGTTCT 1013 3 GAGGGTGGCGGTTCT 4 GAGGGTGGCGGTaCT 5940
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~1894
2283 5 GGcGGCGGCTCT 6 GGTGGTGGTTCT 7 GGTGGCGGCT'CT 8 GAGGGTGGTGGCTCT 1965 9 GAGGGTGGCGGTTCT 10 GAGGGTGGCGGCTCT 11 GAGGGaGGCGGTTCc 2246 12 GGTGGTGGCTCT 13 GGTtcCGGT 2-401Cla I 265 Consensus AGGGTGGXGGXTCT
HaeIII 0 sequence I
(GluGlyGlyGlySer )n
Xmn v
Fig. 3. (A) M13 restriction map. The thick lines show the restriction fragments that werepreferentially
recognized bytheaffinity-purifiedM13sequences. The open boxesgive the localization of the tandem repeats.(B)Alignmentof tandem repeats present in the protein III gene ofM13. The coordinates are as described in (14).
14 15 16 17 18 19 B 1 2 3 4 5 6 7 8
t -
St.-~~w ....
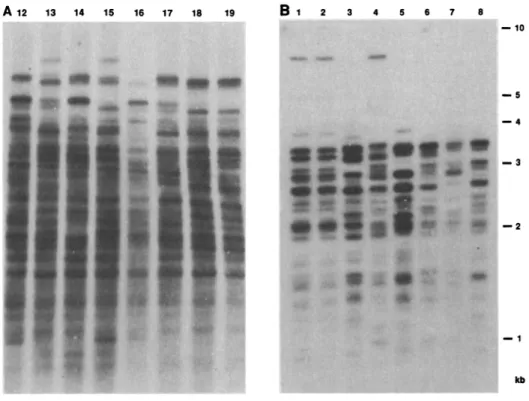
Fig. 4. Hypervariable polymorphic patterns detected in human (A) and bovine (B) DNA by hybridizationto atandemrepeatpresentinM13 (15). (A) Individuals 12to 17are members ofa Caucasianpedigree,18and 19aremonozygotic twins; (B) 1and2aremonozygotictwinsobtainedby embryo transferofasurgicallybisected bovinemorula;bovines 3to8areunrelated animals from the
"BelgianWhiteandBlue"breed. The280-bpM13 HaeIII-Cla Irestrictionfragmentwasobtained
afteragarosegel electrophoresisandelectroelution.Thefragment(100 ng)waslabeled with32ptoa specific activity of 0.5 x 10- to1 x 109cpm permicrogramof DNA(13).Thisprobewasused in Southernblottingexperimentsasdescribed in the legendtoFig. 1 with themilk-based cocktail for
hybridization.
it should be possible to affinity-purify the bacteriophagesequenceinvolvedbyaround
ofhybridization-selection toherringsperm DNA. A radioactively labeled M13 probe
waspurifiedinthiswayandhybridizedtoa
Southern blot containing restriction
frag-mentsofthereplicative formofM13 (Fig.
2B). Total, unpurified M13 was used as a control on a replicate blot (Fig. 2A). The
results indicated that the affinity-purified 684
probe was enriched in sequences
corre-sponding to a region between coordinates
1013and2528ontheM13map(Fig. 3A).
Inspection ofthis region ofthe phage
se-quence revealed the presence of a 15-bp
motifrepeated in tandem at two places in theprotein III gene and corresponding to (Glu, Gly, Gly, Gly, Ser)n (Fig. 3B).A 280-bpfragment essentiallymade ofoneof the
repeat clusters was purified and used as a
probe on a Southern blot of humanDNA cleaved with Hae III. A typical
hyperpoly-morphic pattern was obtained (Fig. 4A). The pattern was clearly different from that obtained with Jeffreys' probe (5, 6) under
similarconditions. Highlypolymorphic
pat-terns were also obtained with DNAof bo-vine (Fig. 4B), equine, murine, and canine origin (11), whichextendstheusefulness of
the M13 to pedigree analyses in animal breeding.
Our results leadtothesurprising conclu-sion that one of the most commonly used
DNA vectors is able, by itself, to detect hypervariable minisatellites in a variety of mammals. The observation that fishDNA, which is frequently added in excess to
hy-bridization mixtures, competes for the
probe explains why this phenomenon had
remained unnoticed. It also suggests that
omission of carrier DNAduring
hybridiza-tion should help in finding additional
probes of this type. Finally, the wide conser-vation of sequences showing
cross-hybrid-ization withtheM13repeat posesthe ques-tionoftheir putativeroleormechanismof
maintenanceduringevolution. An interest-ing clue may befoundinthe recent discov-ery
that
anotherhighly conservedminisatel-lite
isassociatedwithtransposableelements(12).
REFERENCESAND NOTES
1. A. J. Jeffreys, V. Wilson, S. L. Thein, Nature
(London) 314,67(1985).
2. D. R.Higgs,S. E. Y.Goodbourn,J. S. Wainscoat,
J. B.Clegg, D.J.Weatherall, NuceicAcids Res. 9,
4213(1981).
3. A. R. Wyman and R.White,Proc.Natl. Acad.Sci.
U.SA.77, 6754(1980).
4. S. E.Y.Goodbourn,D. R.Higgs, J.B.Clegg,D.J.
Weatherall,ibid.80, 5022 (1983).
5. A.J. Jeffreys,J. F. Y. Brookfield, R. Semeonoff,
Nature(London)317,818(1985).
6. P.Gill,A.J.Jeffreys,D.J.Werrett,ibid.318,577
(1985).
7. G.I.Bell,M.J.Selby,W.J. Rutter,ibid.295,31
(1982).
8. J.Messing, B.Gronenborn,B.Muller-Hill, P. H.
Hofschneider,Proc.Natl.Acad. Sci.U.SA.74,3642
(1977).
9. T. Maniatis,E. F.Fritsch, J. Sambrook,Molecular
Clonitg:ALaboratoryManual(ColdSpnrngHarbor Laboratory, Cold Spring Harbor, NY,1982).
10. D. A.Johnson,J. W. Gautsch, J.R.Sportsman, J.
H.Elder, GencAnal.Technol.1,3(1984).
11. M.Georgesetal.,inpreparation.
12. D. Liebermann, B. Hoffman-Liebermann, A. B.
Troutt,L. Kedes,S.N.Cohen, Mol. Cell. Biol.6,
218(1986).
13. A. P.Feinberg and B. Vogelstein, Anal.Biocaem.
132,6(1983).
14. P. M, vanWezembeck, T.J.M.Hulsebos, J. G.
Schoenmakers,Gene11,129(1980).
15.Thesequenceandproceduredescribed inthe
pre-sentstudyarethesubjectofapendingpatent. 16.Thecontnuoussupportand cnticalinterestofJ.E.
Dumontand R.Hansetaredeeplyacknowledged.
We thank A. Deplano and M. Mairie for expert
technical assistance and D. Leemansfor the
prepara-tionof themanuscript.WearegratefultoA.Massip forproviding thebovine monozgtictwins. Sup-portedbyActionsConcert6es(Minist&edela
Poli-tique Scientifique) and AssociationRecherche
Bio-medicaleetDiagnostica.s.b.l.
29September1986;accepted12December1986
SCIENCE,VOL. 235 A12 13
on May 3, 2009
www.sciencemag.org