THE JOURNAL OF INFECTIOUS DISEASES • VOL. 134, SUPPLEMENT • AUGUST 1976
© 1976 by the University of Chicago. All rights reserved.
Tobramycin Adenylyltransferase: A New
Aminoglycoside-Inactivating Enzyme from
Staphylococcus epidermidisPartha Santanam and Fritz H. Kayser From the Institute of Medical Microbiology, University of Zurich, Zurich, Switzerland
Certain strains of Staphylococcus epidermidis resistant to the aminoglycoside anti-biotics were shown to contain an enzyme that inactivates the kanamycins, neo-mycins, butirosins, paromomycin, gentamicin A, amikacin, and tobramycin by adenylylation. Tobramycin adenylyltransferase, as this enzyme is called, was found to be optimally active at pH 5.5. With paromomycin or neomycin Band C as substrates, however, two pH values (5.5 and 9.0) for optimal activity were observed. The enzyme requires Mg+ + for activity and is stabilized significantly by dithiothreitol. It is probable that the 4'-hydroxyl group of ring I of the anti-biotics is adenylylated. Those aminoglycosides that are not substrates for the enzyme lack a hydroxyl group in the corresponding position.
An epidemiological study of the frequency of resistance of staphylococci to aminoglycosides
showed that 2% of isolates of Staphylococcus
epidermidis were inhibited by ~ 12.5 ug of to-bramycirr/rnl. Such tobramycin-resistant (TmR
) strains were also resistant to kanamycin A, paro-momycin, butirosin, and gentamycin A and inter-mediately resistant to neomycin and amikacin. In a genetic study of TmR in such strains, it was found that resistance was conferred by a small
plasmid (mol wt, 2.7 X 106 daltons) present in
multiple copies [1]. Loss of this plasmid resulted in tobramycin-susceptible variants, which were also susceptible to the other aminoglycosides ex-cept streptomycin.
We now report that the mechanism of resis-tance to aminoglycosides in these staphylococci can be explained by the production of an adenyl-ylating enzyme. We propose to call this enzyme by the trivial name tobramycin adenylyltransfer-ase, because it was detected in staphylococci unusually resistant to tobramycin, and tobramycin was found to be one of the best substrates for it.
Materials and Methods
Antibiotics and chemicals. Tobramycin was
supplied by Eli Lilly and Company
(India-Please address requests for reprints to Dr. F. H. Kayser, Institute of Medical Microbiology, 8028 Zurich, Switzerland.
S33
napolis, Ind.) ; the kanamycins A, B, and C,
amikacin (BB-K8), and butirosin by Bristol
Laboratories (Syracuse, N.Y.); neomycins Band C and spectinomycin by the Upjohn Co. (Kala-mazoo, Mich.); paromomycin by Parke, Davis and Co. (Detroit, Mich.); and the gentamicins Cll C:!, C1ll , and A and sisomicin by Schering Corp. (Bloomfield, N.J.). The other antimicro-bial agents were provided by the pharmacy of the University Hospital (Zurich, Switzerland).
Radiolabeled chemicals were bought from the
Radiochemical Centre (Amersham, England) .
Dithiothreitol and adenosine triphosphate (ATP) were obtained from Calbiochem (Los Angeles, Calif. ). All other inorganic salts of analytic grade were obtained from Merck (Darmstadt, Federal Republic of Germany).
Organism. The representative strain used for our studies was S. epidermidis strain 109, which was isolated from clinical material obtained in Zurich. In addition to tobramycin, this strain is resistant to penicillin (Pen"), chloramphenicol
(CmR
) , tetracycline (TcR) , streptomycin (SmR) , and sulfamethoxazole (SMZH) and is moderately resistant to methicillin (Meth"). The organism was maintained on brain-heart infusion agar (Difco, Detroit, Mich.) containing 20 ug of neomy-cirr/rnl. For production of enzyme, the strain was subcultured on brain-heart infusion agar
contain-ing 10 ug of neomycin Byrnl. The culture was
inoculated into 2 ml of a medium composed of
(Difco), 50 g; NaCl, 10 g; glucose, 4 g; and an amount of water sufficient to make 1 liter. The pH was adjusted to 7.2. For growth of the in-oculum, this medium contained 10ugof neomy-cin B/ml. After overnight incubation, 1 ml of the inoculum was inoculated into 100 ml of fresh medium, which was incubated in a shaking water bath at 37 C. At the beginning of the exponential phase of growth, 1 ml of neomycin B (400/lg/
ml) was added to the culture. When the culture reached an OD of about 1 at 550 nm, cells were harvested by centrifugation at 11,000gfor 10 min at 4 C and were washed twice with 40 ml of a cold buffer containing 0.01 M Tris and 0.03 M NaCI (pH 7.8).
Production of enzyme. The osmotic shock method [2], slightly modified, was used. The washed cells were suspended in 10 ml of 0.033 M Tris-HCl buffer (pH 7.3) containing 0.003 M EDTA and 20% sucrose. After it stood at room temperature (about 20 C) for 30 min, the sus-pension was stirred with a magnetic stirrer for 5 min and centrifuged for 15 min at 12,000gand 4 C. The supernatant was discarded, and the re-maining sucrose was removed carefully from the wall of the centrifuge tubes with cotton swabs. The pellet of cells was suspended in 5 ml of an ice-cold solution of 0.0005 M MgCI2 , and the
suspension was aspirated forcefully about 20 times with a 5-ml pipette. After stirring for 5 min and centrifugation for 15 min (22,000g) at 4 C, the supernatant was carefully decanted to a flask containing 150 III of a 0.04 M solution of dithiothreitol.
For preparation of extracts by sonification, the washed cells were "Suspended in an ice-cold solu-tion of 0.0005 M MgCI2, frozen at - 70 C,
thawed rapidly, and sonified for 3 min at 55 W (Branson sonifier B-12; Branson, Danbury, Conn. ). The procedure was repeated once, after which the suspension was centrifuged for 15 min at 22,000 gat 4 C. The supernatant was decanted and stabilized with dithiothreitol.
For preparation of lysates, cells were lysed by lysostaphin according to procedures described by Shaw and Brodsky [3].
Enzymatic assay. The methods used were similar to those described by Benveniste et al. [4] and to those by Davies et al. [5]. The assay for adenylylation consisted of 20 III of cell extract
(0.5-1.0 mg of protcin/rnl), 1.5 umol of buffer, 0.5-0.7 umol of MgCb, 55 nmol of dithiothreitol, 20 nmol of [2-3H]ATP (specific activity, 60
IlCi/llmol), and 0.6-0.8 nmol of drug base (total volume, 65 Ill). After incubation at 35 C for various periods, 50-Ill samples were pipetted onto a 2-cm2phosphocellulose paper (Whatman P-81; Balsbon, Maidstone, England), which was washed twice with double-distilled water and dried at 55 C. Radioactivity was counted in a Packard Tri-Carb scintillation spectrometer (Packard Instruments, Downers Grove, Ill.) in 10 ml of· a toluene-based scintillation fluid. Reaction mixtures devoid of either the enzyme or the drug were used as con-trols for nonspecific binding of [2-3H]ATP. When
adenylylation had to be measured before com-pletion of the reaction, the tubes containing the reaction mixtures were chilled in ice-water for termination of the reaction. The amount of drug used in all experiments was within the linear range of a quantitative assay.
Acetylating and phosphorylating activities in the cell extract were checked according to methods described by Haas and Davies [6] and Benveniste and Davies [7].
Assays for inactivation of drugs. After com-pletion of the reaction, 50 III of the reaction mixture containing the ingredients mentioned above were placed in wells punched in antibiotic agar no. 3 (Difco) containing 104 spores of Bacillus subtilis (ATCC 6633)/ml as the test organism, and inactivation of drugs was checked. After incubation at 30 C for 24 hr, the degree of inactivation was measured by comparison of the zones of inhibition with appropriate controls.
MIC of drugs. The MIC of antibiotics was determined in Mueller-Hinton broth (Baltimore Biological Laboratories, Baltimore, Md.) [8].
Protein assay. The protein content of cell extracts was measured by the method of Lowry et al. [9].
Results
Enzymatic inactivation of aminoglycosides. Natural resistance to aminoglycosides in gram-positive and gram-negative bacteria is usually the result of a chemical modification and the con-sequent inactivation of the antibiotics by certain enzymes [10]. Since the known
aminoglycoside-Tobramycin Adenylyltransjerase S35
Figure 1. pH dependence of adenylylation reaction with tobramycin adenylyltransferase. Reaction mix-tures were prepared at the indicated pH values with use of appropriate buffers. (0 - - 0 )
=
citrate-phosphate buffer, pH 3.8-6.0; ( e - - e )=
Tris-maleate buffer, pH 5.5-7.5; ( 6 - - 6 )=
barbital buffer, pH 7.5-9.0; ( . . )=
glycine-NaOH buffer, pH 9.0-10.0. Incubation at 35 C lasted for 2.5 min with tobramycin and for 9 min with paromo-mycin. These periods of incubation were in the linear range of assay at the indicated pH values.18 12 PAROMOMYCIN 2 3 I, 5 6 7 8 9 10 2 3 4 5 6 7 8 9 10 pH TOBRAMYCIN 20 16 21, ~o ~ 12 x ~ a. ~ J=8 inactivating enzymes are periplasmic enzymes and
are released from R-factor-containing
gram-negative bacteria by osmotic shock [2], the same procedure was used for extraction of the enzyme from S. epidermidis strain 109.
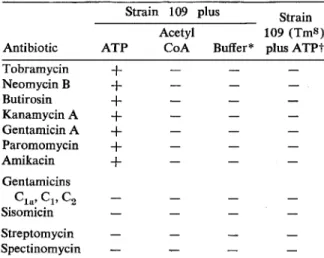
Those drugs that are substrates for the enzyme were inactivated when the reaction mixture con-tained ATP. No inactivation was observed when
the reaction mixture contained either acetyl
coenzyme A or cell extracts from tobramycin-susceptible variants. All of the drugs except gentamicins C1a, C2, and C3, sisomycin,
strepto-mycin, and spectinomycin were inactivated (table
1) .
Adenylylation of aminoglycosides. Since ATP was required for inactivation of the enzyme, we assumed that the antibiotics were modified either by O-adenylylation or O-phosphorylation. No transfer of radioactivity from [y_32P]ATP to the aminoglycosides was detectable in the enzymatic assay. Radioactivity was, however, transferred to
the drugs from [2-3H]ATP. The optimal pH for
enzymatic reaction with tobramycin, kanamycin A, B, and C, the butirosins, and gentamicin A
was 5.5; with amikacin it was 6.0, although
adenylylation occurred over. a broad range of pH (4.5-9.5). Adenylylation of paromomycin, neo-mycin B, and neoneo-mycin C had two pfl optima, 5.5 and 9.0 (figure 1).
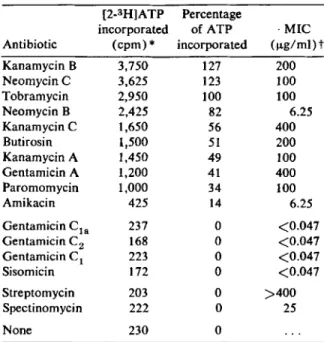
The efficiencies of adenylylation of the different aminoglycosides by tobramycin adenylytransferase
at pH 5.5 are listed in table 2. It can be seen
that this enzyme is different from both
strepto-mycin-spectinomycin adenylyltransferase and
gentamicin adenylyltransferase, since
strepto-mycin and spectinostrepto-mycin, the gentamicinsCta, Ct,
and C2, and sisomicin are not adenylylated [10].
At pH 5.5, tobramycin is one of the best
sub-strates, whereas amikacin is a rather poor one. In general, the efficiency data reflect the phenotypic resistance of strain 109, except in the case of neo-mycin B, which is readily adenylylated but weakly inhibitory to the strain. Adenylylation therefore may inactivate the drug incompletely, an effect resulting in a low MIC value.
When the kinetics of adenylylation were
examined, it was found that some antibiotics were Strain
109 (TmS )
Buffer
*
plus ATPt AcetylCoA ATP
Table 1. Effects of adenosine triphosphate (ATP) and acetyl coenzyme A (acetyl CoA) on inactivation of aminoglycoside antibiotics by lysates of osmotic-ally shocked Staphylococcus epidermidis strain 109.
Strain 109 plus Antibiotic
NOTE. Inactivation was determined by the agar diffu-sion test as described in Materials and Methods. A plus sign indicates that the antibiotics were inactivated; a minus sign indicates no inactivation in comparison with results in appropriate controls. Unlabeled ATP (20 nmol) or unlabeled acetyl CoA (10 nmol) was used in each assay.
*
The buffer used was Tris-maleate (pH 5.5).t Strain 109 (TmS ) is a spontaneous, tobramycin-sus-ceptible variant of strain 109.
Tobramycin
+
NeomycinB+
Butirosin+
Kanamycin A+
Gentamicin A+
Paromomycin+
Amikacin+
Gentamicins Cta'c,
C2 Sisomicin Streptomycin SpectinomycinGentamicin CIa 237 0 <0.047 Gentamicin C2 168 0 <0.047 Gentamicin C I 223 0 <0.047 Sisomicin 172 0 <0.047 Streptomycin 203 0 >400 Spectinomycin 222 0 25 None 230 0
'" ATP
=
adenosine triphosphate. The assays wereper-formed as described in Materials and Methods. Samples were incubated for 6 min with all substrates to ensure an assay in the linear range. Efficiency of adenylylation was calculated with reference to radioactivity of the adenyly-lated tobramycin. A count of 2,950 cpm/min was equated with 100% efficiency. Controls devoid of either enzyme or drug had an average value of 230 cpm.
t MICs of drugs were determined in Mueller-Hinton
broth [8].
Table 2. Adenylylation of aminoglycoside otics by celI-free extracts of Staphylococcus midis strain 109. [2-3H]ATP Percentage incorporated of ATP ( cpm) '" incorporated Discussion T08RAMVC!N
In recent years it has become apparent that, under
some circumstances, S. epidermidis can be
con-sidered to be as pathogenic as Staphylococcus
aureus. Certain infections, such as endocarditis,
urinary tract infections, and colonization of
ventriculoatrial shunts are typically caused by S. epidermidis[11-14]. In addition, this organism can sometimes be isolated in pure culture from pyogenic material. The isolated bacteria often prove to be quite resistant to antibiotics [15-17].
Resistance to all ~-lactam antibiotics, to the
tetracyclines, to erythromycin, streptomycin, and the sulfonamides, and often also to kanamycin, neomycin, and chloramphenicol has been demon-strated [18]. Thus the new aminoglycoside anti-biotics may become significant aids in the treat-ment of staphylococcal infections.
However, we have found that some S.
epider-midis strains are resistant to multiple drugs, including many of the new aminoglycoside anti-biotics. A previous genetic study revealed that the determinants of resistance to aminoglycosides in representative strains of these staphylococci are plasmid-borne and can be transduced between
antibi- epider-200 100 100 6.25 400 200 100 400 100 6.25 ·MIC (ug/rnljt 127 123 100 82 56 51 49 41 34 14 3,750 3,625 2,950 2,425 1,650 1,500 1,450 1,200 1,000 425 Antibiotic Kanamycin B NeomycinC Tobramycin Neomycin B Kanamycin C Butirosin Kanamycin A Gentamicin A Paromomycin Amikacin 10 20 30 40 50 60 0 10 20 30 40 50 60 TIME (minutes)
Figure 2. Kinetics of adenylylation of various aminoglycoside antibiotics. Samples were incubated at 35 C for various periods, after which 50-~1 portions were pipetted onto a phosphocellulose paper which was washed and counted for radioactivity.
adenylylated at approximately the same rate but not to the same extent (figure 2). A possible explanation for this behavior is that the end products differ slightly in ability to inhibit the
enzymatic reaction. However, such inhibition
does not seem to be a major factor in determin-ing levels of resistance in intact bacteria (table 2).
Properties of tobramycin adenylyltransferase.
Of the three methods tested, osmotic shock yields a preparation with the highest enzymatic activity. Like other aminoglycoside-inactivating enzymes, the enzyme has an absolute requirement for Mg+ + ions. Dithiothreitol stabilizes the enzyme significantly. The transferase is stable for at least
60 days when stored at -40 C in the presence of
dithiothreitol. Nearly 90% of activity was lost after storage for 60 days at 4 C and for one week at 22 C (table 3). (Y) I a x ::E o, u I I (Y) KANAMVCIN B
100, 100, 100 100, 100, 87 100, 100, 61 100, 100, 11 100, 87, 0 100, 80, 0 100, 9, 0 Tobramycin Adenylyltransjerase
Table 3. Properties of tobramycin adenylyltrans-ferase.
Adenylylation
Property (% )*
Preparation of enzymet
Osmotic lysate (0.5 mg of protein/ml) 100
Sonified extract (1.0 mg of protein/ml) 54
Lysed extract (2.9 mg of proteiri/rnl ) 11
Mg++requirement (per assay)t
No~ 4 0.05J,tmol 33 0.1 umol 64 0.3 umol 88 0.5 urnol 97 0.7 urnol 100 1.0 urnol 95 2.0 umol 91
Dithiothreitol requirement (per assay)§
None 44 55 nmol 100 Stability (days) t
o
1 6 7 20 30 60*Adenylylation values listed in relation to the property
of stability were obtained at -40 C, 4 C, and 22 C, re-spectively.
t Activity was measured after incubation for 30 min at 35 C, with neomycin B as substrate, in Tris-HCI buffer (pH 8.0). A count of 3,658/min was equated with 100% activity.
t Activity was measured after incubation for 15 min at
35 C, with tobramycin as substrate, in Tris-maleate buffer (pH 5.5). A count of 5,276/min was equated with 100% activity.
§Activity was measured after incubation for 6 min at
35 C, with neomycin B as substrate, in Tris-rnaleate buffer (pH 5.5). A count of 3,884/min was equated with 100% activity.
strains with rather high frequency [1]. The size and other characteristics of this extrachromo-somal element are not unusual for staphylococci [1]. However, the plasmid is unusual, insofar as it carries the determinants for production of an aminoglycoside-inactivating enzyme with a broad range of activity. To our knowledge, an enzyme with the properties of tobramycin adenylyltrans-ferase has not been found in either gram-positive or gram-negative bacteria.
S37
Although the end products of enzymatic re-action are yet to be isolated and identified, the substrate profile strongly suggests that adenylyla-tion occurs at the 4' -hydroxyl group of ring I of these antibiotics. The kanamycins, the neomycins, the butirosins, paromomycin, amikacin, genta-micin A, and tobramycin all contain such a group in this position, whereas sisomicin and gentamicins Ct a, Ct, and C2 do not (figure 3). Except for
butirosin, all of the substrates have yet another hydroxyl group that can be adenylylated (i.e., the 4"-hydroxyl group of ring III of tobramycin, gentamicin A, the kanamycins, and amikacin, and the 4'"-hydroxyl group of ring IV of the neo-mycins and paromomycin. Whether or not this second hydroxyl group is also adenylylated re-mains to be determined after analysis of the adenylylated antibiotics.
The demonstration of a tobramycin adenylyl-transferase that inactivates the classical amino-glycosides as well as some of the newer antibiotics of this group implies that no guarantees of free-dom from resistance to antibiotics of the future
can be given. Thus, although resistance to
gentamicin has not been found in staphylococci isolated in our area despite use of the drug for a decade, it is perhaps only a matter of time before
resistant strains appear. If such resistance is
plasmid-borne, the widespread intraspecies and perhaps also interspecies distribution [19] of this plasmid under appropriate selective conditions in vivo might result. Therefore, an important aspect of antistaphylococcal chemotherapy in the future should be a restriction of the use of the new, highly potent antibiotics to cases in which these drugs are absolutely necessary.
References
1. Rosendorf, L. L., Kayser, F. H. Transduction and
plasmid deoxyribonucleic acid analysis in a
multiply antibiotic resistant strain of
Staphylo-coccus epidermidis. J. Bacteriol. 120: 679-686, 1974.
2. Neu, H. C., Heppel, L. A. The release of enzymes
from Escherichia coli by osmotic shock and
during the formation of spheroplasts. J. BioI.
Chern. 240:3685-3692, 1965.
3. Shaw, W. V., Brodsky, R. F. Characterization of
chlorarnphenicol-acetyltransferase from
chloram-phenicol-resistant Staphylococcus aureus. J.
A
c
.z.,.s:
KANAIotYCINS A NHZ OH B NHZ NHZ C OH NHZHO~~
HO~HZ'N~NI0 H N AI II RZ CHBZO~0 OH NHZ R 0 ' "HO~"'"
I' OH H~Z'N~O R4 RI RZ R3 R4 NEOIlAYCINB NHZ H H H CHZNHZ NEOIlAYCIN C NHZ H H CHZNHZ H PARQllAOIlAYCIN OH H H { CH ZHNH2 CH;NHZ}B
D
RI RZ R3 R4 R5 R6 GENTAIot'C1NS A H OH OH OH H OH Cia H NHZ H H OH CH3 Cz CH3 NHZ H H OH CH3 CI CH3 NHCH3 H H OH CH3Figure 3. A, structure of kanamycins. Positions in ring I are numbered 1'-6'; in ring II, 1-6; and in ring III, 1"-6". Positionyis the 2' position. Tobramycin is 3'-deoxykanamycin B. Amikacin is 1-hydroxyamino-butyryl-kanamycin A. B, structure of gentamicins, Sisomicin is 4', 5'-dehydrogentamicin Cia' C, structure of
neo-mycins. D, structure of butirosins.
4. Benveniste, R, Yamada, T., Davies, r, Enzymatic
adenylylation of streptomycin and spectinomycin
by R-factor resistant Escherichia coli. Infec.
Immun. 1: 109-119, 1970.
5. Davies, J., Brzezinska, M., Benveniste,R R-factors:
biochemical mechanism of resistance to amino-glycoside antibiotics. Ann. N.Y. Acad. Sci. 182: 226-233, 1971.
6. Haas, M. 1., Davies, J. Enzymatic acetylation as a means of determining serum aminoglycoside concentrations. Antimicrob. Agents Chemother. 4:497-499, 1973.
7. Benveniste, R, Davies, J.Aminoglycoside
antibiotic-inactivating enzymes in actinomycetes similar to those present in clinical isolates of antibiotic-resistant bacteria. Proc. Natl. Acad. Sci. U.S.A. 70:2276-2280, 1973.
8. Ericsson, H. M., Sherris, J. C. Antibiotic sensitivity
testing: report of an international collaborative
study. Acta Pathol. Microbiol. Scand. [B] 79
(Suppl. 217):1-90,1971.
9. Lowry, O. H., Rosebrough, N. J., Farr, A. L.,
Randall, R J. Protein measurement with the
Folin phenol reagent. J. Biol. Chern. 193 :265-275, 1951.
10. Benveniste, R., Davies, J. Mechanisms of antibiotic resistance in bacteria. Annu. Rev. Biochem. 42: 471-505, 1973.
11. Holt, R. J. The colonisation of ventriculo-atrial
shunts by coagulase-negative staphylococci In
M. Finland, W. Marget, and K. Bartmann [ed.}.
Bayer Symposium III. Bacterial infections.
Springer-Verlag, Berlin, 1971, p. 81-90.
12. Holt, R J. Coagulase-negative staphylococci as op-portunist pathogens. Arzneim. Forsch. 21:325-328,
1971.
13. Holt, R J. The pathogenic role of
coagulase-negative staphylococci. Br. J. Dermatol. 86
(Suppl. 8) :42-47, 1972.
14. Keys, T. F., Hewitt, W. J. Endocarditis due to micrococci and Staphylococcus epidermidis. Arch. Intern. Med. 132:216-220, 1973.
Tobramycin Adenylyltransferase
15. Andriole, V. T., Lyons, R. W. Coagulase-negative
Staphylococcus. Ann. N.Y. Acad. Sci.
174:533-544, 1970.
16. Bentley, D. W., Hahn, J. J., Lepper, M. H.
Trans-mission of chloramphenicol-resistant
Staphylo-coccus epidermidis: epidemiologic and laboratory
studies. J. Infect. Dis. 122:365-375, 1970.
17. Marsik, F. J., Parisi, J. T. Significance of
Staphylo-coccus epidermidis in the clinical laboratory. Appl.
S39
Microbiol. 25: 11-14, 1973.
18. Pulverer, G., Damen, G. Neugebauer, M. Antibiotic resistance of Staphylococcus albus. Med. Micro-bioI. Immunol. 158:32-43, 1972.
19. Yu, L., Baldwin, J. N. Intraspecific transduction in
Staphylococcus epidermidis and interspecific
trans-duction between Staphylococcus aureus and
Staphylococcus epidermidis. Can. J. Microbiol. 17:767-773, 1971.