HAL Id: hal-01898766
https://hal.archives-ouvertes.fr/hal-01898766
Submitted on 18 Oct 2018HAL is a multi-disciplinary open access archive for the deposit and dissemination of sci-entific research documents, whether they are pub-lished or not. The documents may come from teaching and research institutions in France or abroad, or from public or private research centers.
L’archive ouverte pluridisciplinaire HAL, est destinée au dépôt et à la diffusion de documents scientifiques de niveau recherche, publiés ou non, émanant des établissements d’enseignement et de recherche français ou étrangers, des laboratoires publics ou privés.
FIS-Dependent trans Activation of Stable RNA Operons
of Escherichia coli under Various Growth Conditions
Lars Nilsson, Hans Verbeek, Erik Vijgenboom, Cornelis van Drunen, Anne
Vanet, Leendert Bosch
To cite this version:
Lars Nilsson, Hans Verbeek, Erik Vijgenboom, Cornelis van Drunen, Anne Vanet, et al.. FIS-Dependent trans Activation of Stable RNA Operons of Escherichia coli under Various Growth Con-ditions. Journal of Bacteriology, American Society for Microbiology, 1992, 174 (3), pp.921-929. �10.1128/jb.174.3.921-929.1992�. �hal-01898766�
0021-9193/92/030921-09$02.00/0
Copyright© 1992, American Society for Microbiology
FIS-Dependent
trans
Activation of Stable
RNA
Operons
of
Escherichia coli under Various
Growth Conditions
LARS NILSSON,t HANS VERBEEK, ERIKVIJGENBOOM, CORNELISVANDRUNEN,
ANNE VANET,t AND LEENDERT BOSCH*
Departmentof Biochemistry, Leiden University, Gorlaeus Laboratories, P.O. Box9502,
2300 RA Leiden, TheNetherlands Received 14 June1991/Accepted 14 November 1991
InEscherichia colitranscription of the tRNA operonthrU(tuJB)andthe rRNA operon rrnB istrans-activated by the protein FIS. This protein, which stimulates the inversion of various viral DNA segments, binds
specifically to a cis-acting sequence (designated UAS) upstreamof the promoter of thrU (tuJB) and the P1 promoter of the rrnB operon. There are indications that this type of regulation is representative for the regulation of more stable RNA operons. In the present investigation we have studied UAS-dependent transcription activationofthe thrU(tuiB)operonin the presenceand absence ofFISduringanormal bacterial growthcycle andafter anutritional shift-up. Inearly log phase theexpressionof the operon risessteeply in
wild-typecells, whereafter it declines. Concomitantly, a peakof the cellular FIS concentration is observed.
Cellsin thestationary phasearedepleted of FIS. The rather abrupt increase of transcription activation depends
onthenutritionalqualityof the medium. It isnot seenin minimalmedium. Afterashift from minimaltorich medium, a peak of transcription activation andof FIS concentration is measured. Thispeakgetshigherasthe medium gets more stronglyenriched.We conclude thatacorrelation between changes of theUAS-dependent activation of the thrU (tuJB) operon and changes of the cellular FIS concentration under a variety of
experimental conditions exists. This correlation strongly suggests that the production of FIS responds to
environmentalsignals, thereby trans-activating the operon. Cells unabletoproduce FIS(fiscells) also showan
increaseofoperontranscriptionin theearly log phase andafteranutritionalshift-up,albeit lesspronounced
than that of wild-type cells. Presumably it is controlled by the ribosome feedback regulatory system. cis activationofthe operon by the upstreamactivatorsequenceis apparent in the absenceofFIS. This activation is constant throughout the entire growth cycle and is independent of nutritional factors. The well-known growth rate-dependent control, displayed by exponentially growing cells studied under various nutritional
conditions, is governed by two regulatory mechanisms: repression, presumably by ribosome feedback inhibition, andstimulation by transactivation. FISallows veryfastbacterial growth.
Thesynthesisof rRNA of Escherichia coli is finely tuned
to the cell's environmental conditions. Cells growing in a constantenvironmentdonotshowasignificantturnoveror a significant buildup of free rRNA or vacant ribosomes,
exceptatverylowgrowthrates(for reviews,seereferences 20, 21, and 26). Consequently, ribosomes are utilized at
maximal ornear-maximal capacity. Upon alteration of the
nutritionalcapacity of the medium, leadingto adifferent but
constantenvironment, cellspromptly readjust the synthesis of their rRNA and tRNAto meetthe demands ofanaltered
growth rate. In exponentially growing cells the
concentra-tionof ribosomes (and of rRNA) thusappears tobe
propor-tionaltogrowthrate(6, 8, 9). The mechanismunderlying this so-calledgrowthrate-dependentcontrol has been described
as a feedback inhibition of rRNA synthesis by ribosomes
(ribosomefeedback control) (5, 12, 13, 29)and/orinhibition
by ppGpp (stringent control) (4), the concentration of this
nucleotidebeingafunction ofthe growth medium (28). Gaal andGoursereported thatE. colimutantsunable to produce
ppGppare still undergrowth rate-dependent control. They concluded thatppGpp isnottheonly factor involved in this
type of regulation (7).
*Correspondingauthor.
tPresent address: Department of Cell Biology, University of Stockholm, S-106 91 Stockholm, Sweden.
t Present address: Institut deBiologie Physico-Chimique, 75005
Paris, France.
Interestingly, it has been reported that the synthesis of tRNA issubjectto the same regulatorymechanisms as the synthesis of rRNA (14, 32). While studying the regulation of thetRNA operonthrU
(tuJB),
werecently showed that it is also regulated by a positively acting control system.Up-streamof thisoperonacis-actingsequenceisfound, deletion of which results in an80to 90%drop oftranscription (31). Thissequence,called UAS forupstreamactivatorsequence,
appearedto be the targetofatrans-activating protein (34), whichwe subsequentlyidentifiedasthe protein FIS (3, 25).
Up till then this heat-stable protein was only known tobe involved in site-specific recombination (2, 17, 30). It stimu-lates theinversion of various viral DNAsegmentsbybinding
to arecombinational enhancer (16, 19).
It may be envisaged that more stable RNA operons are
activated intransby thissystem. Accordingly, the UASs of the tyrT, metY, thrU(tufB), and rrnB operonsall bindone
and thesameprotein (25, 33). Sequencesupstreamof theP1
promoters of all rRNA operons and the promoters of 13
tRNA operons (but not all tRNAoperons) match the con-sensus sequence for FIS-binding sites (18, 33). Recently
Rossetal.(18, 27)independently demonstratedthat FISacts asthetransactivator ofthe rrnB operon. Inthiscontextthe
question of under which conditions the trans activation controlsystembecomes operativearises.Preliminary stud-ies fromthislaboratory (3,25) revealedlargefluctuations in
the trans activation of the thrU (tufB) operon during a
normal bacterialgrowth cycle. Here wehave studied these
922 NILSSON ET AL.
TABLE 1. E.coli strains used inthisinvestigation
Strain Genotype or phenotype Reference orsource
MC1000 lacX74araDl39(araleu)7697 galU galKStrr 15
MC1000-fis767 lacX74 araD139(araleu)7697 galUgalKStrrfis Kanr 15
JM101 supEthi(lac-proAB) 22
JM101-fis767 supE thi (lac-proAB) fis Kanr Ourlaboratory
fluctuations in moredetail. We alsoaskedwhether changes
inthelevel oftransactivationoccurinresponseto environ-mental signals and whether they are accompanied by changes in thecellular FIS concentration. Of further interest was whether growth rate-dependent control is solely gov-erned by ribosome feedback (see above) orwhether trans
activation also plays a role in this process. Finally we describe the benefits for thecellof having FIS.
MATERIALS ANDMETHODS
Strains, plasmids, and growth media. TheE. coli strains used inthisstudyarelisted in Table1.Theplasmid pDS10 is described in reference31. Itharborstheoperonfusion thrU
(tufB)':galK, and the UAS extends from position-176tothe
promoter. pDS10AUAS is identicalto pDS10exceptthat it carriesadeletion extending from -500to-57.
M9 minimalmedium was supplemented with thiamine (1
jig/ml), essential amino acids (20 ,ug/ml), and succinate (1.0%) (30);inthecaseof minimal medium plus amino acids
supplementation was with 0.5% casein hydrolysate. LB
medium(per liter: 10gof Bacto Tryptone,5gof BactoYeast
Extract, 10gofNaCl, pH 7.5)wasprepared by the method
of Miller (23). In the nutritional shift-up experiment 0.1 volume ofa 1Ox concentrated LB medium without NaCl was added. Brain heart infusion medium was prepared
accordingtothe manufacturer'sdescription (Difco Labora-tories, Detroit, Mich.). In most of the experiments with a rich medium the mediumwasreplaced by LBplus glucose
(1%), since this improved the reproducibility oftheresults.
Determinationofgalactokinase activity.The bacteria
trans-formedwithpDS10 orpDSAUAS weregrownovernightin
the medium indicated. The cultures werediluted and
incu-bated further at 37°C. At the times indicated the optical densityat600nm
(ODwo)
was measured and samples werewithdrawn. Galactokinase activities and plasmidcopy
num-bersweredeterminedbythemethod of Adams and Hatfield
(1) with modifications byvanDelftetal. (31).
Bacterial growth.Growthwasmeasuredby reading
OD6w
(inmostcases)orby determinationof thedry weightof the
biomass(see Fig. 8).Inthe lattercasecellsweresedimented
andwashedwith0.83%NaCl,whereafter the sedimentwas
weighed afterlyophiization.
Determination ofthecellular FISconcentration. For each
determination cellextractswereprepared fromtwo
indepen-dentbacterial cultures (40hin minimalmediumpriortothe
shift-upandovernightpriortoagrowth cycleinLBmedium
plus 1% glucose). After sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) and Western
immunoblotting with FIS antibodies pretreated with an extract ofthefis strain MC1000-fis767, FIS was detected
with the ECL fluorescence kit of Amersham. Fluorescence signals were quantified by determining peak areas with a
laserscanner.Theseareas werecorrelated with theareaofa cross-reactingprotein representativefortheprotein
concen-tration inthe samples. Therelative FIS concentration
plot-ted in thefigures represents the indirect FIS concentration/ total soluble protein. It is corrected for blot and detection
efficiencybyusing purified FIS as aninternalstandard. The standard deviation was calculated for each point with the data of three independentWestern blots and two scans of each blot. Thelargest deviation found was 15%.
RESULTS
Experimental strategy. In order to study theexpression of thethrU(tufB)operon invivo under various cellular growth conditions we used the plasmid-borne operon fusion as previously described (31). This fusion, thrU
(tuJB)':galK,
which puts theexpression of galK under control of the thrU(tuJB)promoter, was introduced intogalK-defective E. coli strains. Expression of the operon was studied by measuring
galactokinaseactivity/femtomoleofplasmid. Inherent to this procedure is that a difference in life span of the galactokinase protein and the transcript may lead to anoverestimationof thenumber oftranscripts present at the times indicated and thatgalactokinaseactivitieslagsomewhatbehind changes in FISconcentration andtranscription. Whenrelevant, this is
pointedoutin thetext(e.g.,seeFig. 3A). Large fluctuations in galactokinase activity are observed during a bacterial growthcycle and after a nutritionalshift-up (cf.Fig. 2 to 4). These are duetosynthesisanddegradationof the
galactoki-nase protein, since they cannot be ascribed to comparable
fluctuations inplasmidcopy numbers.Changes in therateof
galactokinase mRNA or protein degradation, occurring
un-der the variousgrowth conditions, may thus bias the results. This drawback can beovercome asindicated below.
Since transcription activation of the operon depends on
the presence of the UAS, galactokinase activities were
measured in cells carrying a plasmid with an intact UAS
(designatedUAS+cells)orwith theUAS deleted
(designat-ed UAS- cells). Other regulatory mechanisms, known to
control stable RNA synthesis such as ribosome feedback inhibition andstringentresponse, have DNAdeterminantsin theregionfromposition -50to +1(11).DNAdeterminants oftranscriptionactivation, however,arefound in the
region
extendingfrom -131to -48(25, 31, 34). Thisenabledus to
distinguishbetweentheeffectofUAS-dependentactivation and thatofrepression by anotherregulatory mechanism(s).
Galactokinase activities of UAS+ cells thus reflect the effects of all regulatory mechanisms including activation,
whereas theactivities of UAS- cells reflect theeffectsof all mechanisms except activation. The ratio of galactokinase
activities of UAS+ and UAS- cells isa measure of activa-tion. It isindependent ofchanges in the rateof mRNA or
protein degradation.
We have alsostudiedtheeffect of FISonactivation andon
other regulatory systems by performing the same
experi-mentsinastrainlackingafunctionalfisgene (fis cells).The
complete lackof FIS infis cellswasconfirmed byWestern blotting withFIS-specific antibodies(datanotshown).
Expression of the thrU
(tall)
operon during a bacterialE E 400 * 200 :R A cm time(min.) 20 0 10~~~~~~~~~ 02 0.1 0 0.05 0 100 200 time(min.)
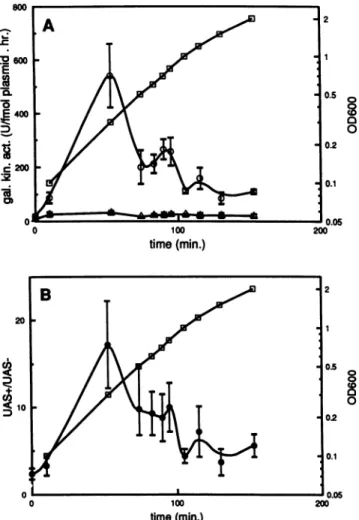
FIG. 1. Expression of the plasmid-borne thrU (tufB) operon
during abacterial growth cycle. E. coli MC1000 was transformed
with pDS10 or the plasmid deletion derivative lacking the UAS, pDS10AUAS (see Materials and Methods). The cells were grown
overnight in brain heart infusionmedium,dilutedtoanOD600of0.1, and incubated at 37°C. (A) Galactokinase activities expressed in
unitsperfemtomole of plasmidperhour.0,UAS+ cells;A,
UAS-cells; O,
OD6w.
(B) Ratios of galactokinaseactivities inUAS+andUAS- cells (0) as calculated from the data in panel A. Standard
errors(SE) of the ratiosweredeterminedby thefollowingformula:
[(SEof UAS+ activity/UAS+ activity)2 + (SE of UAS- activity/ UAS-activity)2]05.The errorbarsrepresenttheSE, ineachcase
based on three independent experiments. O,
OD6..
Backgroundlevels ofgalactokinase activities, representing nonspecific binding of radioactive materialtothe membraneandenzymaticactivityat0°C,
were7% ± 2% for UAS+ cells and 6% ±4% forUAS- cells.
growth cycle. With the strategy outlined above the
expres-sion of the thrU (tufB) operon was investigated during a
normal bacterial growth cycle in various media. Overnight
cultureswere diluted to an
OD6.
of0.1, and galactokinaseactivitiesweremeasured atvariousstagesthereafter. Figure
1A shows the results obtained with cells grown in a rich
medium (brainheart infusion). Large fluctuations in galac-tokinase levelsareseenduringthe growth cycle, which is in
agreement with our previous results (25). Almost
immedi-ately after dilution ofa stationary phase culture in fresh
medium, alargeincrease ofgalactokinaseactivityoccursin UAS+cells. Thatthisincrease ismainly duetoactivation of transcription can be concluded from the behavior of
UAS-cells.The latter cells also showanincrease ingalactokinase
activity, but it is much less pronounced. Apparently, an
additional regulatory mechanism differing from activation
becomes operative afterreinitiation ofgrowthbut the
stim-ulation of transcription by this mechanism is greatly
out-weighed by the activation of the operon. The increase in
activation is clearly reflected by the ratio ofgalactokinase
activities in UAS+ and UAS- cells (Fig. 1B).
Theinitial rise ingalactokinase activities is followedbya
dropinboth UAS+ and UAS- cells. The decline in
activa-tion is reflected bya decrease ofthe ratio ofgalactokinase
activities in UAS+ and UAS- cellsfrom 17toapproximately
10(Fig. 1B).Afterthis rather steepdrop,activation declines
further,
albeit more slowly, asindicated by ratios of 4 to 5when cells approach the stationaryphase.
Sensingthe nutritionalquality ofthemedium.Theincrease inactivation inearlylog phaseisaffectedbythecomposition
of the medium. This became apparent by studying the
UAS+-to-UAS- ratios during outgrowth of overnight
cul-tures in various media. The ratio varies from less than 5 in
minimal mediumto approximately 16in LB medium and 17
in brain heart infusion medium (maximal ratios are not
always
observed at the same cell concentration). Eventhough we cannot fully rule out that we have missed the
exact activation peak during the growth cycle, the large
differences observed indicate that the cells sense the
nutri-tional quality of the medium and respond with an altered
activation level.
Transcription regulationinthe presence andabsence ofFIS.
Transcription activation of the thrU (tufB) operonis depen-dent on the protein FIS (3, 25), as is clearly shown by
comparingthe expression ofthe operon inwild-type andfis
cells (Fig. 2). Overnight cultures ofboth types of cells are
devoid of FIS (see below). Accordingly, deletionof theUAS
has the sameeffectonthegalactokinaseactivities of station-arywild-type andfis cells, i.e., an approximately threefold reduction of the galactokinase activities (compare zero time
activities inFig. 2A andB). Apparently,we aredealinghere
witha cis effect ofthe UAS in the absence of FIS. Thiscis
effect should beconstantthroughoutthe entire growthcycle
infis
cells.This is indeed what theexperiment shows, ascanbe concluded from the UAS+-to-UAS- ratios (Fig. 2B and
Table 2). A FIS-independent effect of the UAS on the
expression of therrnB operonhas beenshown, both in vivo
and in vitro, by Gourse and coworkers (27). In contrast to
what is seen in fis cells, the UAS+-to-UAS- ratio rapidly increases in wild-type cells (Fig. 2A and Table 2). The
activation that becomes operative in these cells thusis dueto
twoeffects: one is dependentonFIS occurringimmediately after reinitiation ofgrowth, and the other most likely is a
FIS-independent cis effect induced by the nucleotide se-quence, since it acts continuously and at a constant level.
Figure 2A and B further show expression of the thrU
(tufi) operonafter deletion ofthe UAS in wild-type andfis cells. As mentioned above, an additional control
mecha-nism, different from UAS-dependent activation, becomes
apparent here. We suggest that it reflects derepression of ribosome feedback inhibition control.
Interestingly, the UAS-independent transcription persists
foralonger period of time infiscells than in wild-typecells,
leading to asignificantly highergalactokinase level infis cells than in wild-type cells. A similar phenomenon is observed
with therrnBoperon (27). Although further experiments are needed, these data suggest that UAS-dependent activation and theadditional regulatory system affect each other.
Ascanbe seen in Table 2 and Fig. 2A and B, expression of the thrU (tufB) operon with an intactUAS isalmostequal
924 NILSSON ET AL. X200 E
a
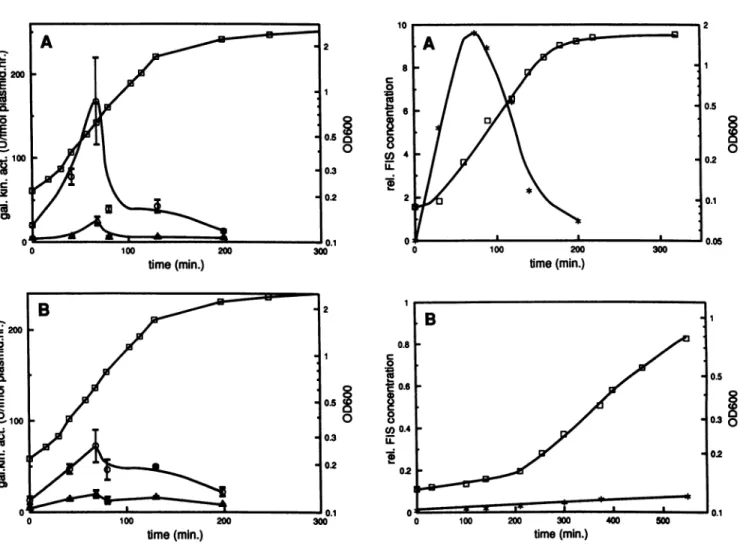
cL i100 C co time(min.) tme(min.)FIG. 2. Effectsof FIS and UAS on thrU(tuiB)expression during
growthcycle. Wild-typecells (MC1000) (A) andfiscells
(MC1000-fis767) (B) were transformed with the plasmids described in the legend to Fig. 1.Overnightcultures werediluted to anOD6 of 0.2 in LB medium and grown at 37°C. Galactokinase activities are
expressedasdescribedin the legend toFig. 1. Forsymbol
defini-tions,see thelegend toFig. 1.
inwild-typeandfis cells foracertainperiodof time (between
OD6.
of0.8and 1.6). Sincefis cells cannot use thetran-scription activationsystem, theyapparentlycompensatefor it by derepressing the ribosome feedback control system.
Cellularlevels of FIS. Thelarge fluctuations in
transcrip-TABLE 2. Galactokinaseactivity ratios of various cell types
(MC1000andMC1000-fis767) determinedduring
thegrowthcyclea
Galactokinaseactivityratio of:
OD6'WTUAS+/ WTUAS+/ fisUAS+/
WTUAS- fis UAS+ fis
UAS-0.2 4.1 ± 0.87 1.5 ±0.52 2.9± 1.0 0.4 12.0± 2.9 1.6 ±0.26 3.3± 0.45 0.6 6.7 ± 2.4 2.3 ±0.91 3.6± 1.2 0.8 6.5 ± 1.2 0.84±0.22 3.6± 1.3 1.6 6.8 ± 1.3 0.86± 0.16 3.0± 0.2 2.2 2.5± 0.32 0.59± 0.15 2.6 ±0.55 2.4 3.5 ±0.67 0.56±0.12 3.8 ± 1.0
a SeelegendstoFig.2Aforwild-type (WT)cellsandFig.2Bforfiscells.
c 0 co C3
1
1
Co 4 0.2 2 0.1 0 0.05 0 100 200 300 time(min.)B
0.8 ii~~~~~~~~~~~~~~~~~~~~~0.5
time(min.)FIG. 3. Cellular FIS concentration during growth cycle. Over-night cultures of the wild-type strain MC1000 were diluted to an
ODC of0.15 and grown at37°Cin LB medium plus 1% glucose (A) or in minimal medium (1% succinate) (B). At the times indicated the cellular FIS concentration was determined asdescrbed in Materials and Methods. *, relative FIS concentration (see Materials and
Methods);O.
OD6w.
tion activation of the thrU(tufB) operon duringthegrowth cycleraise thequestionof whether the cellularlevel of FIS also varies. Thompson et al., using a FIS-DNA binding
assay, reported that the FIS level dropped 70-foldas cells
wentfrom late logtostationary phase (30). Wedetermined the cellular FISconcentrationduring the entire growth cycle
by usingWesternblotting. Cells cultured inarich medium (LBplus1%glucose)showarapidincrease of their FIS level in the early log phase (Fig. 3A). A peak value of FIS is reachedapproximately 75 minafterinitiation of thegrowth cycle, whereafter a rather steep decline sets in. In the stationary phase FIS has dropped toa level atwhich it is undetectable andsohastranscriptionactivation. The
maxi-mum level of FIS does not exactly coincide with that of
galactokinase activity (cf. Fig. 2A and 3A). This is to be expected, since the galactokinaseactivities giveanindirect estimate of the number of transcripts and will lag behind changes in transcription. Conceivably, the fluctuations in FIS level largely govern the fluctuations in transcription
activationduringthegrowth cycleinthis medium.
Cells growing in minimal medium(1% succinate) do not
show an abrupt rise in FIS content. The slight gradual
increaseof FIS observed in
Fig.
3B isatthe lower limits of detection and may be ofquestionable significance.
We conclude that the fluctuations in the cellular concentration of FIS are sensitivetoenvironmentalsignals.
Since transcrip-tion activatranscrip-tion also varies withchanges
in thecompositionof themedium,
a correlation betweenchanges
in theUAS-dependent transcription
activation of the thrU(tufil)
operon andchanges
of the cellular FIS concentration becomes apparent. This correlationstrongly
suggests that environ-mental conditionssignal
trans activation of the operon byFIS. Furtherconfirmation of this correlation was obtained
by studying
the effectofa nutritionalshift-up.
Effects of a nutritional
shift-up.
Figure
4 shows there-sponseof
exponentially growing wild-type
andfis
cellsto ashift from minimal mediumtoLB medium. The
growth
rateof
wild-type
cellspromptly
increases from 0.4 to 2.2dou-blings
per h(not
shown),
whilegalactokinase activity
in-creases
approximately
10-foldover aperiod
of2hafter the shift(Fig.
4A).
This enhancedexpression
of the thrU(tuJB)
operon isgreatly
due to a rise intranscription activation,
since the
UAS+-to-UAS-
ratio increasesapproximately
fourfold
during
thisperiod.
Thecells also
respond
withachange
in their FIScontent(Fig.
4B).
As illustrated above(Fig. 3B),
cellsgrowing
inminimal medium do not
display
sudden fluctuations in FISconcentration
during
thegrowth cycle,
and thisconcentra-tion remains very low.
Immediately
afterchanging
thecomposition
of themedium,
however,
a steep rise of FISoccurswithina1-h
period,
whereafter it also declines rathersteeply
and levelsoffatarelatively
elevated level.Cells unable to
produce
FIS(fis
cells[Fig.
4C]) likewisesense this rather drastic
change
in thecomposition
of the medium. Thegalactokinase activity rises, although
not tothesame level as in
wild-type
cells,
but the UAS+-to-UAS-ratio remains constant.Apparently,
aregulatory
mecha-nism,
different fromtransactivation,
isresponsible
for this enhancement of the thrU(tuiB)
expression.
A limited nutritional shift up from 1% succinate to 1%
CasaminoAcids
plus glucose
also enhances thegrowth
rateof
wild-type
cells(from
0.4 to 1.1doublings
per h[not
shown])
and theexpression
ofthethrU(turB)
operon,as canbeconcluded from
Fig.
5.Galactokinaseactivity
risesover aperiod
of about 1h(Fig.
5A).
Thattransactivationpartici-pates in the enhanced
transcription
canbe concluded fromtheincreasesinthe
UAS+-to-UAS-
ratio and in cellular FIS level(Fig.
5A andB,
respectively).
Finally, fis
cells alsorespond
withanincreaseingalactokinase activity (Fig.
5C). Sincetransactivationdoesnottakeplace
in these cells(the
UAS+-to-UAS-
ratioremainsapproximately
constant),
the elevatedexpression
of theoperon isgoverned
by
adifferent controlmechanism,
mostlikely
ribosomefeedbackcontrol.Apparently
the responses ofwild-type
andfis
cells to alimited nutritional
shift-up
arequantitatively
rather similar butqualitatively
different in theregulatory
mechanismre-sponsible
for thechange
intranscription.
Pulse-chase experiments.
Although FIS-dependent
transactivation sofar has
only
been demonstrated in vivoandin vitro for the thrU(turB)
and rrnB operons (3, 18, 25, 27),various data suggest that the
regulation
of these operons isrepresentative
for that of more stable RNA operons (seeabove).
Inthiscontextonemay ask whether thestimulation ofstable RNAsynthesis by
a nutritionalshift,
as studiedmore
directly
by pulse-chase experiments,
differs inwild-type
andfis
cells. Adirectcomparison
of the results ofFig.4with those ofa
pulse-chase experiment
inwhich minimal medium is shiftedtoLBmedium istechnically
notfeasible,
C 0 (. c 8 CO) cn LD time(min.)
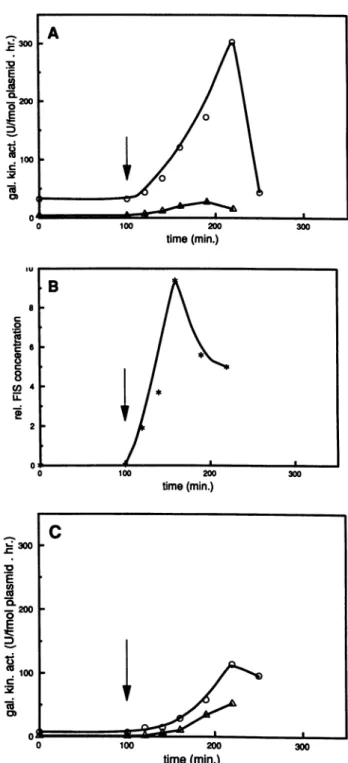
FIG. 4. Response to an extensive nutritional shift-up. Cells grown overnight in minimal medium were diluted 10-fold with minimal medium and grownat37°Cfor 2h,whereafter(arrow) the mediumwasshifted toLB mediumplus 1%glucose. At thetimes indicatedgalactokinase activityand FIS concentration were deter-mined as described in Materials and Methods. (A) Galactokinase activities ofwild-type MC1000 cells transformed with pDS10 (0)
(UAS+ cells) orwithpDS10AUAS (A) (UAS- cells);(B) relative cellular FIS concentration of MC1000 cells (*); (C) galactokinase
activitiesofMC1000-fis767(fis cells), transformed withpDS10 (0)
926 NILSSON ET AL. 100
time
(min.) 0.6 U)0.4 Er: time(min.) E isi
L200
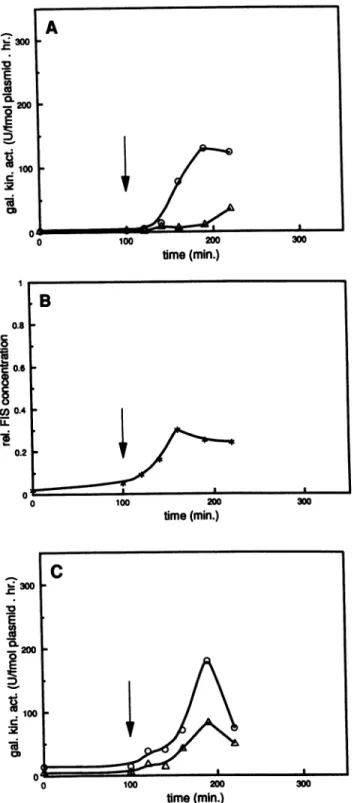
M 00 100 0 100 200 time(min.)FIG. 5. Responsetoalimitednutritional shift-up. Experimental
conditionswereidenticaltothosedescribedinthelegendtoFig. 4,
except that the medium was shifted from minimal to Casamino Acids(1%) plus glucose (1%).
sincethelabelisdilutedoutinLBmedium. Shiftingfrom1%
succinate to 1% Casamino Acids plus 1% glucose was
therefore carried outinstead. Cells receiveda2-min
[3H]u-ridine pulse which was followed by an 8-min chase. The resultsdiffered somewhatdependingonthe pretreatmentof
thecellspriortothe shift, butessentiallytheoutcomewas as
illustratedinFig.6. StableRNAsynthesisinbothwild-type
FIG. 6. Stable RNAsynthesisafter alimited nutritional shift-up, studiedby pulse-chase experiments. MC1000-fis767 (fis)cells were
cultured in minimal medium and shifted fromminimal medium to
Casamino Acids (1%) plus glucose
(1%).
One-milliliter culturesamples were pulse-labeledwith 37 KBqof
[3H]uridine
for 2min,
which wasfollowedby achasewith0.125mMnonlabeled uridine, whereafter0.2 ml of anonlabeled overnight cultureas acarrierand 3volumesof7%trichloroaceticacid wereadded.Precipitateswere
collected onGF/C filters, washed, dried, andcounted in a
liquid
scintillationcounter
U,
wild-type MC1000cells; O,fiscells.andfis cells increases immediately after the change of the
medium. We conclude that the response of stable RNA
synthesisto alimitednutritional shiftisrathersimilartothat of thrU
(tufB)
expression. Theresults of Fig. 6 are also in line with those of Fig. 5 in that they do not reveal largequantitativedifferences in the responses of wild-typeandfis
cells. Qualitatively the responses of both cell types differ;
however, since wild-type cells use trans activationto stim-ulate theexpression of the thrU
(tuJB)
operon(cf. Fig. 5B),whereasfiscells do not (seeDiscussion).
Growth rate-dependent trans activation. The fact that rrn
operons lacking the UAS are submitted to growth
rate-dependent control(11) indicates that cellsincapableoftrans
activation utilize the mechanismofnegative regulation by ribosome feedback. However, since the present
investiga-tion shows that the composition of the medium has a
pronouncedeffect on theUAS-dependenttransactivation,it doesnotseem verylikely thatgrowthrate-dependent regu-lation is solely
governed
by repression of stable RNAsynthesis. In order to investigate this question we have grownUAS+andUAS-cells indifferent media,permitting
a variation of the growth rates of between 0.9 and 2.0
doublingsperh.
Cells
wereharvestedat anOD6.
of0.4, and theirgalactokinase activities were determined. In this waythepresentresults arefullycomparabletothoseobtainedby
Gourseandcoworkers(11), whostudiedthe DNA
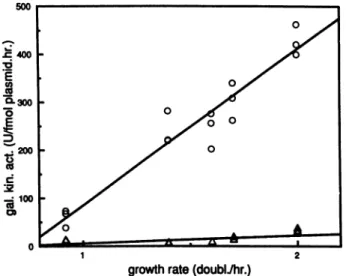
determi-nantsofgrowthrate-dependent controlof therrnBoperon. As isapparent fromFig.7, bothUAS+and UAS- cells show
alinearrelationshipbetweenthegalactokinase activitiesand thegrowthrates, inaccordancewith thefindingsfor therrnB
operon (11). In this range ofgrowth rates the increase in
galactokinase
activities is 2.6times for UAS-cells
but4.6times forUAS+
cells.
Increasedgrowthratethusleadsto anincreased UAS+-to-UAS- ratio. We conclude that two
controlmechanisms underliegrowthrate-dependent
regula-tion: repression, presumably by ribosome feedback, and
stimulationbytransactivation.
400 E °
1F
CL .5~~~~~~0
100 _ 0 Cu cm 0 1 2growth rate(doublJhr.)
FIG. 7. Expression of the thrU (tufB) operon in exponentially growing cells at various growth rates. Transformants described in thelegendtoFig.1weregrownovernightin LBmedium and diluted toanOD600of 0.1 in differentmedia. Growth rates were determined
duringtheearly log phase,andgalactokinase activities were mea-sured at an
ODwo
of 0.4.Background galactokinaselevels were6%± 1.5%forUAS+cellsand20%o±4%for UAS- cells(cf.legendto
Fig. 1). The media used were minimal medium, minimal medium
supplemented with Casamino Acids, LB medium, LB medium
supplemented withglucose, andbrain heart infusion medium (0, UAS+cells;A,UAS-cells).
FISallows very fast cellular growth. We have seen thatthe
expressionof thefis gene is verypronouncedwhen station-ary cells reinitiategrowthinarich medium (Fig. 1 to 3). A
rapid increase of FIS is accompanied by a rise of
FIS-dependent trans activation of the thrU(tufB) operon. One maytherefore expectwild-type cells to grow faster thanfis
cells under these conditions. Thisexpectation isborne out
bythefollowing experiment. Wild-type(MC1000) cells and
fis (MC1000-fis767) cellsweregrown in LBmedium supple-mented with 1% glucose to an
OD6.
of 0.3. The FIS concentration ofwild-type cells has then reached itsmaxi-mum (cf. Fig. 3A). At this stage both types of cells were diluted 40-fold, whereafter the
OD6.
was carefully moni-tored. Atregular intervals samples were collected to deter-minethedry weight ofthe accumulated cell mass. As can be concludedfromFig.8,growthratesof 2.2 and 1.4doublingsper hwerefound forwild-type andfiscells, respectively, on the basis ofreadings of
ODwo,
and growth rates of 1.8 and 1.3doublingsper h onthe basisof biomassassays. Theeffect ofthefisdeletion isnotstrainspecific, sinceweobtained thesameresultwith the strain JM101.
Recently, Gille et al. (10) reported that FIS binds and
bendsthe origin ofchromosomal DNAreplication, oriC,of
E. coli and that FIS is requiredforminichromosome
repli-cation. The difference in growth rate ofwild-type andfis cells, as measured here by reading OD and by assaying biomass, cannot be ascribed to a reduced replication ability. Weconclude that exponentially growingcellsable to trans-activate stableRNA synthesishave anadvantage over cells unabletotrans-activate and that one of the functions of FIS is to allow fast cellular growth (see Discussion).
DISCUSSION
Acceleration of cellular growth is accompanied by an elevated transcription of the thrU (tufB) operon. Such an
U - ~1.000 , 0,2 500/ 0 A. 0.1 200 0,05 100 0,02 50 2 4 time (hrs.)
FIG. 8. Growth of wild-type cells (MC1000) and fis cells
(MC1000-fis767) in rich medium. Cellsgrown toan
OD6w
of 0.3inLBmedium plus 1%glucosewerediluted 40-fold andgrownat37°C
in the samemedium.Atthetimesindicated,
OD6w
(opensymbols)was read and the dry weightof the accumulated cell mass(closed
symbols) was determined (see Materials andMethods). O and *, wild-type cells; A andA,fiscells.
acceleration clearly occurs after dilution of an overnight
culture in fresh medium(Fig. 1and2)and afterashift from minimal to rich medium (Fig. 4). In both cases the rise in
transcription is dependent on the UAS, the target of the protein FIS. Deletion of the UAS abolishes the increase of thrU (tufB) expression in the early log phase almost
com-pletely(Fig. 1A). Theratio ofgalactokinaseactivities of cells
carryingtheplasmid-borne operonfusion thrU(tufB)':galK
with an intact or a deletedUAS (the UAS+-to-UAS- ratio)
rises substantially (Fig. 1B). As pointed out above (see "Experimental strategy" above), this ratio is a measure of
transcription activation which is independent ofchangesin the rates of galactokinase mRNA andgalactokinaseprotein
degradation during the course of the experiment. In cells harboring the fis gene, this activation increasesimmediately
aftergrowthacceleration and ingeneraldeclines thereafter.
Incells unable toproduce FIS (fis cells) the
UAS+-to-UAS-ratio remains constant throughout the entire growthcycle.
ThisFIS-independent cisactivationis notaffectedby envi-ronmental conditions. We assume that it is induced by the
nucleotidesequenceof the UAS. Increases of the UAS+-to-UAS- ratioarealsoseeninwild-typecells after anutritional shift-up (Fig. 4A and 5A). In contrast this ratio remains constantinfiscells(Fig. 4C). Concomitant with the changes in transcription activation, the cellular FIS concentration rises in wild-type cells, both in early log phase and aftera
shift-up (Fig. 3A, 4B, and SB). Apparently, a correlation
between changes of the UAS-dependent activation of the thrU(tufB) operon andchanges of the cellular FIS concen-tration exists.
Various lines of evidence indicate that environmental conditions affect the FIS production and the activation of
transcription. First, the UAS+-to-UAS- ratios during bac-terialoutgrowth in freshmedium vary with thecomposition
of the medium. The large rise and fall of FIS, observed during the growth cycle inarich medium (Fig. 3A),are not
observedduringacyclein minimal medium(Fig. 3B). After
a shift from minimal to rich medium, however, both the
cellular FIS content and operon transcription activation increase promptly (Fig. 4B and A, respectively). The
in-928 NILSSON ET AL.
creases of FIS and oftranscription activation are largeras
themedium is more strongly enriched (cf. Fig. 4B and SB).
We conclude that environmental conditions affect
tran-scription activation and the FIS concentration in the same
way. This correlation, apparently occurring underavariety
ofconditions(seeabove), strongly suggests that it is FIS that trans-activatestranscriptionofthe thrU
(tuJB)
operon inE.coli. Thealternative model, that FIS concentrationschange
in response to changes instable RNAsynthesis,isnot avery logical one and has some rather bizarre implications. FIS-dependent trans activation apparentlyacts as a sensorof the nutritionalquality of the medium. These data raise
interest-ing questions concerning the regulation of de novo FIS
synthesis and the transduction of the environmental signals
involved.
As pointed out in the introduction, anumberof observa-tions indicate that the trans activation of the thrU
(tuJB)
operonisrepresentativefor sucharegulationofmorestable RNA operons.One and the sameproteinbinds in vitrotothe UASs ofthe tyrT, metY, andthrU
(tuJB)
operonsandtothe UAS upstream of the P1promoterof the rrnB operon (25).FIS-dependent transactivationof the rrnBoperon has been demonstratedbyRoss etal. (18, 27). Sequence comparisons
of upstream regions (18, 33) are also in line with the
assumption that more stable RNA operons (albeit not
nec-essarily all) are submitted to this regulatory system and respond to FIS. If so, this would indicate that one of the basic elements of growth control is the regulation of the de novo synthesis of FIS.
Cells unabletoproduceFIS(fis cells)alsoshow enhanced
transcriptional activityof the thrU(tufB)operon inearlylog phase of thegrowth cycle (Fig. 2B) andafter enrichment of themedium(Fig. 4C andSC),albeit in rich mediato alesser
extent than wild-type cells. Since no
FIS-dependent
transactivation of the operon takes place in these
cells,
theUAS+-to-UAS- ratio remains virtually constant,
irrespec-tive of the environmental conditions. The fact that
galactoki-nase activity of UAS+ cells exceeds that of UAS- cells
reflects cis activation of the operon by the UAS in the
absence of FIS.
Significant transcription
of the operon is thus observed infiscellsduringthegrowth cycle
(Fig. 2B).
In fact, wild-type andfis cells only show a
pronounced
difference inUAS-dependentexpression
of theoperondur-ingtheinitialtranscriptional jumpin
early log phase
(cf.
Fig.
2A and B). Thereafter, virtually no difference is seen foracertainperiod oftime (between
OD600
of 0.8 and1.6).
Apossible explanation of these data is thefollowing.
In wild-type cells this leveling off at a lower level is due toribosome feedback and adrop in FIS concentration.
Ribo-some feedback in these cells may result from a temporary
excess ofribosomes, present after the steep rise in stable RNAsynthesisinearly log phase.
Infis
cells this rise is lesspronouncedandsomay be ribosome feedback inhibition. On the other hand thesecells donot
trans-activate,
sotranscrip-tion in bothcelltypes reachesapproximatelythesamevalue.
Inaccordance with adifferentextentof ribosome feedback in wild-type and fis cells are the results obtained upon
deletingthe UAS of the reporter operon. In
wild-type
cellsexpressionof theoperon dropstovery low
levels,
whereasinfiscells theexpressionremainsatthe samelevel forlonger periodsof time (Fig. 2B).
When stable RNA synthesis is studied in
exponentially
growing cellsatvariousgrowthratesby
altering
the compo-sition of the medium, a linear relationship between theexpression of stable RNA operons and the
growth
rate is found(11, 24). Bothribosome feedback andtransactivationgovernthis
growth
rate-dependent regulation,
as isevident from Fig. 6, in which theexpression
of the thrU(tufB)
operonwas
investigated.
Inaccordancewithprevious
stud-iesof therrnBoperon(24),
growth-rate
dependent
controlis observed when the thrU(tufB)
operon is studied after deletion of the UAS. From thisfinding
one should notconclude, however, that ribosome feedback control is the
only
mechanismunderlying growth
rate-dependent
regula-tion. TheUAS+-to-UAS-
ratios measuredatvariousgrowth
ratesclearlyindicate that both the
positive
and thenegative
control mechanismsareoperative
under these conditions.Finally
thequestion
of whether the bacterial cell benefits fromharboring
afis
gene maybe asked.First,
therearetwocounteracting
regulatory
systems; apositively
operating
trans activation and a
negatively operating
ribosome feed-back may allow finetuning
oftRNAand rRNAsynthesis.
Aprerequisite
of such abalanced control is that transactiva-tion,
like ribosomefeedback, responds
to environmentalsignals.
The results of the presentstudy
demonstrate that this condition is fulfilled.Second,
FIS enables the cell togrow very fast. This may be
quite
anadvantage
when cells grow underthe conditionsprevailing
in nature.Until
recently
theE. coliprotein
FIS was knownonly
toplay
a role as a host factor in thereplication
of certainphages.
As such itdid notseemto contributetothewelfare of the bacterial cell. It has become evidentnowthat cellsdo benefit from this heat-stableprotein
andareabletoswitchonits
synthesis
when environmental conditions make this de-sirable.ACKNOWLEDGMENTS
Theplasmids containingthe UASof the thrU
(turB)
operonand deletion derivatives thereofwere generously donated by J. van Delft. Thestrains MC1000 and MC1000-fis767werekindlyprovidedby R. C. Johnson. The technical help of Anneke Kuipers is
gratefully acknowledged.WeareendebtedtoN.Goosen and P.van de Puttefor valuablehelpand suggestions.
TheinvestigationwassupportedinpartbytheCommission of the
EuropeanCommunities, BiotechnologyAction Programme (BAP),
Directorate-GeneralScience,Research andDevelopment,Brussels. L.N.wastherecipientofalong-termEMBOfellowship,and A.V.
wassupported bytheERASMUS program.
REFERENCES
1. Adams,C. W., andG. W.Hatfield. 1984. Effects ofpromoter
strengthsandgrowthconditions oncopynumber of
transcrip-tion-fusionvectors. J. Biol.Chem. 259:7399-7403.2. Betermier, M., V. Lefrere, C. Koch, R. Alazard, and M.
Chandler. 1989. The Escherichia coli
protein,
Fis:specific
bindingtotheends ofphageMuDNAandmodulationofphagegrowth. Mol. Microbiol. 3:459-468.
3. Bosch, L., L. Nilsson, E.
Vlgenboom,
and H. Verbeek. 1990.FIS-dependenttrans-activationof tRNA and rRNA operons of Escherichia coli.Biochim.Biophys. Acta1050:293-301.
4. Cashel, M., and K. E. Rudd. 1987.The stringentresponse,p.
1410-1438.In F.C. Neidhardt,J. L.Ingraham, B.
Magasanik,
K. B. Low, and M. Schaechter (ed.), Escherichia coli and Salmonellatyphimurium:cellularandmolecularbiology.Amer-icanSocietyforMicrobiology,Washington, D.C.
5. Cole, J. R., C. L. Olsson, J. W. B. Hershey, M.
Grunberg-Manago, and M. Nomura. 1987. Feedback
regulation
of rRNAsynthesisinEscherichia coli.J. Mol. Biol. 198:383-392. 6. Dennis, P. P., and H. Bremer. 1974. Differential rate of
ribo-somalprotein synthesis in Escherichia coli B/r. J. Mol. Biol. 84:407-422.
7. Gaal, T., and R. L. Gourse. 1990. Guanosine 3'-diphosphate
5'-diphosphate isnotrequiredforgrowthrate-dependent
con-trolofrRNAsynthesisinE. coli.Proc. Natl. Acad.Sci. USA 87:5533-5537.
8. Gausing, K. 1974.Ribosomal protein in E. coli: rate of synthesis and pool size at different growth rates. Mol. Gen. Genet. 129:61-75.
9. Gausing,K. 1977.Regulation of ribosome production in Esch-erichia coli: synthesis and stability of ribosomal RNA and of ribosomal protein messenger RNA at different growth rates. J. Mol. Biol. 115:335-354.
10. Gille, H., J. B. Egan, A. Rothand, and W. Messer. 1991. The FIS protein binds and bends the origin of chromosomal DNA replication, oriC, of Escherichia coli. Nucleic Acids Res. 15: 4167-4172.
11. Gourse, R. L., H. A. de Boer, and M. Nomura. 1986. DNA determinants of rRNA synthesis in E. coli: growth rate depen-dentregulation, feedback inhibition, upstream activation, anti-termination. Cell 44:197-205.
12. Gourse, R. L., Y. Takebe, R. A. Sharrock, and M. Nomura. 1985. Feedback regulation of rRNA and tRNA synthesis and accumulation of free ribosomes after conditionalexpressionof rRNA genes. Proc. Natl.Acad. Sci. USA 82:1069-1073. 13. Jinks-Robertson, S., R. L. Gourse, and M. Nomura. 1983.
Expression of rRNA and tRNA genes in Escherichia coli: evidence for feedback regulation by products of rRNA operons. Cell 33:865-876.
14. Jinks-Robertson, S., and M. Nomura. 1987. Ribosomes and tRNA, p. 1358-1385. In F. C. Neidhardt, J. L. Ingraham, B. Magasanik, K. B. Low, and M. Schaechter (ed.), Escherichia coli andSalmonella typhimurium: cellular and molecular biol-ogy, vol. 2. American Societyfor Microbiology, Washington,
D.C.
15. Johnson, R.C., C. A. Ball,D.Pferrer, and M.I. Simon. 1988. Isolation of the gene encoding the Hin recombinational
en-hancerbinding protein. Proc. Natl. Acad. Sci. USA 85:3484-3488.
16. Johnson, R. C., and M. I. Simon. 1985. Hin-mediated
site-specific recombination requires two 26 bp recombination sites anda60bp recombinational enhancer. Cell 41:781-791.23. 17. Johnson, R. C., and M. I. Simon. 1987. Enhancers of
site-specificrecombination in bacteria. TrendsGenet. 3:262-267. 18. Josaitis, C. A., T. Gaal, W. Ross, and R. I. Gourse. 1990.
Sequences upstream of the -35 hexamer of rrnB P1 affect promoterstrength and upstream activation. Biochim. Biophys. Acta 1050:307-311.
19. Kahmann, R., F. Rudt, C. Koch, and G. Mertens. 1985. G inversion in bacteriophage Mu DNA is stimulated by a site within the invertase gene and a host factor. Cell 41:771-780. 20. Lindahl, L., and J. M. Zengel. 1982. Expression of ribosomal
genesin bacteria. Adv. Genet. 21:53-121.
21. Lindahl, L., and J. M. Zengel. 1986. Ribosomal genes in Escherichia coli. Annu. Rev. Genet. 20:297-326.
22. Messing, J. 1979. A multipurpose cloning system based on
single-stranded DNAbacteriophage M13. Recomb. DNA Tech. Bull. 2:43.
23. Miller, J. H. 1972. Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. 24. Miura, A., J. H. Krueger, S. Itoh, H. A. de Boer, and M.
Nomura. 1981. Growth-rate dependentregulation of ribosome synthesis inE.coli:expression of the lacZ and galK genes fused toribosomal promoters. Cell 25:773-782.
25. Nilsson, L., A.Vanet, E.V"genboom, and L. Bosch. 1990. The role of FIS in trans-activation of stable RNA operons of E. coli. EMBOJ. 9:727-734.
26. Nomura, M., R. L. Gourse, and G. Baugham. 1984.Regulation ofthe synthesis of ribosomes and ribosomal components. Annu. Rev.Biochem. 53:75-117.
27. Ross, W., J. F. Thompson, J. T. Newland, and R.L. Gourse. 1990. E.coli Fis protein activates ribosomal RNA transcription in vitro and in vivo. EMBO J. 9:3733-3742.
28. Ryals, J., R.Little, and H. Bremer. 1982.Control of rRNA and tRNA synthesis in Escherichia coli by guanosine tetraphos-phate. J. Bacteriol. 151:1261-1268.
29. Takebe, Y.,A.Miura,D. M.Bedwell,M.Tam,and M.Nomura. 1985. Increasedexpression of ribosomal genes during inhibition of ribosomalassembly in Escherichiacoli. J. Mol. Biol. 184:23-30.
30. Thompson, J. F.,L.MoitosodeVargas, C. Koch, R.Kahmann, and A.Landy.1987. Cellular factorscouplerecombination with growth phase: characterization of a new component in the
site-specific recombinationpathway. Cell 50:901-908.
31. vanDelft, J. H. M., B. Marion,D. S. Schmidt, andL. Bosch. 1987. Transcription of the tRNA-tuJB operon of Escherichia coli:activation,termination and antitermination. Nucleic Acids Res. 15:9515-9530.
32. vanDelft, J. H. M.,H. M.Verbeek, P. J.deJong,D.S.Schmidt, A.Talens,and L. Bosch.1988.Control of the tRNA-tufB operon inEscherichia coli. 1. rRNA gene dosageeffects and growth-rate-dependent regulation. Eur. J. Biochem. 175:355-362. 33. Verbeek, H., L.Nilsson, G. Baliko, and L. Bosch. 1990. Potential
bindingsitesof the trans-activator FISarepresent upstreamof all rRNA operons and of many but not all tRNA operons. Biochim. Biophys. Acta1050:302-306.
34. VUgenboom,E., L.Nilsson, and L. Bosch. 1988. The elongation factor EF-Tu fromE.coli bindstothe upstream activator region of thetRNA-tufBoperon. NucleicAcids Res. 16:10183-10197.