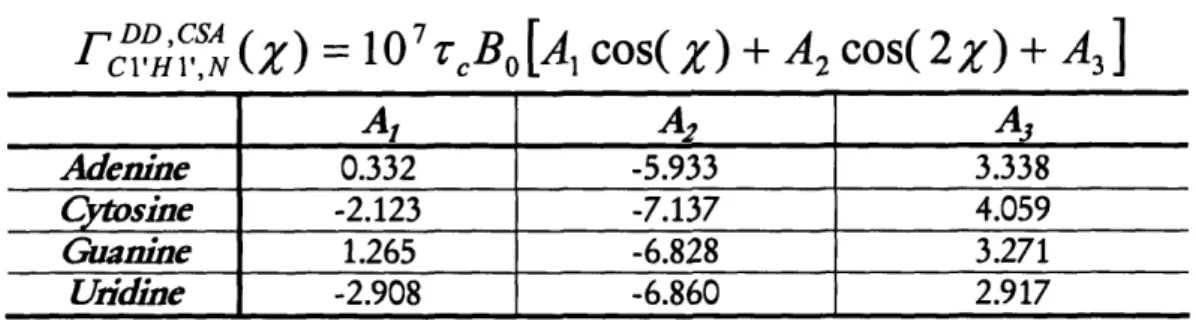Development of new Parameters for
Structure Determination and Dynamic Investigations
on
Biomacromolecules by NMR
by
Elke Duchardt
Dipi
phil mzt, Bidriste
Unirsity fFrankfut Gernd 1999
SUBMITtED TO THE DEPARTMENT OF CHEMISTRY IN PARTIAL FULFILL OF THE
REQUIREMENTS FOR THE DEGREE OF
DOCIOR OF PHILOSOPHY IN BIOLOGICAL CHEMISTRY
AT THE
MASSACHIUSETTS
INSTITUTE OF TECHNOLOGY
Febary 2005
© 2005 Massachusetts Institute of Technology. All rights reserved.
1.
Author
/'7
/
x/ 1! VDepartment of Chemistry
September 17, 2004 ,,Certified by
Harald J. SchwalbeProfessor of Chemistry
Thesis SupervisorAccepted by
Chairman, Departmental
Robert W. Field
Department of Chemistry
Committee on Graduate Students
MASSACHUSETTS INSTITUTE, OF TECHNOLO", MAR 2 5 2005
LIBRARIES
...- ;.%,njv-zs j"
-Z4
This doctoral thesis has been examined by a committee of the Department of Chemistry as follows:
•I r ) /
Professor Robert G. Griffin
kk,
Chair
Professor Harald J. Schwalbe
Thesis Supervisor
Development of new Parameters for Structure Determination and
Dynamic Investigations on Biomacromolecules by NMR
by
Elke Duchardt
Submitted to the Department of Chemistry in Partial Fulfillment of the Requirements for the Degree
of Doctor of Philosophy
ABSTRACT
Nuclear magnetic resonance (NMR) spectroscopy is unique in the content of structural as well as dynamic information that it can provide at atomic resolution. The aim of this PhD-thesis was to contribute to the understanding of biochemical processes by means of NMR-spectroscopic techniques, targeting specific problems as well as contributing to the general understanding and providing new, widely applicable methods. The main focus was on the structural as well as dynamic study of ribonucleic acids (RNA).
A new structural NMR method was developed aimed at the determination of the glycosidic
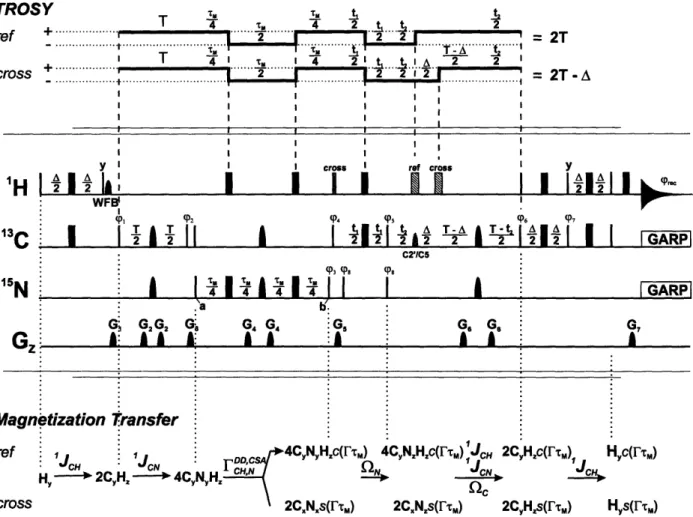
torsion angle X in RNA, which defines the relative orientation of the nucleobases in respect to the ribose moiety (Duchardt et al., 2004). X was derived from the angle dependence of carbon-hydrogen dipole-dipole, nitrogen chemical shift anisotropy cross-correlated relaxation rates (F-rates). Method development comprised the design of a novel NMR experiment, the (HCN), as well as the introduction of F versus parameterization curves. The novel method provides an accuracy of
around 10 degrees or better, comparable to the precision of conventional angle determination
techniques. In contrast to conventional methods, the F(HC) is sensitive to molecular size and will therefore proof beneficial in the investigation of larger RNAs by NMR
Apart from this methodological contribution to RNA structure determination, the dynamic
properties of the abundant YNMG RNA tetraloop motif (with Y=C or U; N= any base; M=C or A)
were studied in a residue specific manner by means of 3C NMR relaxation measurements. The dynamics of the extraordinarily stable cUUCGg motif were compared to the less stable uCACGg hairpin, which forms the stem-loop D (SLD) in the regulatory 5'-cloverleaf of coxsackievirus 3B. Measurements were carried out at 25°C, at which both motifs are stable, as well as at close to the
melting point of SLD (43°().
Ribose and base moiety specific amplitudes and time-scales of motion were extracted from R1,
R
1p and the heteronuclear nOe of Q6 andCQ.
in pyrimidines andCQ
andCQ.
in purines by application of the model-free formalism (Lipari and Szabo, 1982). The application of the model-free formalism toC
1.,
and C6 which possess an additional adjacent carbon spin, was examined for the uniformly isotope labeled RNA hairpins investigated in this study. In addition, the relaxation data analysis was optimized for the 13C chemical shift anisotropy based on chemical shift tensors available to date (Stueber and Grant, 2002;Fiala et al., 2000). While at room temperature, the dynamics closely follow the structural features of both hairpins, residues in the loop closing as well as in the adjacent base-pair exhibit highly increased flexibility at temperatures close to the melting point of SLD. In contrast, loop dynamics remain unperturbed. In SLD, the residues close to the loop are conserved among thefamily of rhino- and enteroviruses, indicating a sequence based mechanism of decoupling loop structure and stability in order to adjust to the twofold requirement of a defined structure for protein recognition and low stability to ensure efficient melting within the genomic transcription process.
In addition to investigations on RNA, structural studies were also conducted on the C-chain of the T-cell receptor (TCR) in order to contribute to the elucidation of the early events in T-cell activation. According to earlier studies (Aivazian and Stem, 2000), the extracellular encounter of the T-cell receptor with an antigen is transmitted into the T-cell via a lipid induced structural transition
of the TCR C-chain cytoplasmic domain. In order to lend detailed structural support to the proposed
mechanism, the structure of TCR 4 was studied in free solution and in the presence of detergent micelles. By means of standard NMR-techniques, the backbone assignment of both forms of 4 was achieved and secondary structural information was obtained. In addition, the binding site of the protein Nef of simian immunodeficiencyvirus (SIV) onto TCR ~ was determined.
Thesis Supervisor. Harald Schwalbe
Title: Professor of Chemistry
References:
Aivazian, D. and Stem, L. J. (2000) Nat Sta Bio, 7, 1023-6.
Duchardt, E., Richter, G, Ohlenschlager, O., Gorlach, M, Wohnert, J. and Schwalbe, H. (2004) J
Am(banSr, 126, 1962-70.
Fiala, R., Czernek, J. and Sklenar, V. (2000)
J
Bind NMR, 16, 291-302.Lipari, G. and Szabo, A. (1982) JAm CaenSoc 104, 4546-4559. Stueber, D. and Grant, D. M. (2002) JAmCIemSc, 124, 10539-51.
"Them
are a
rmillion
zs
to get no signal"
Harald Schwalbe
"7n
ee
just muke nore scans"
Christian Richter
'"Still,
Me ill er gt as
rch
signl as on a Varian"
THANKS TO....
.... THE SUPERVISORS
I am deeply indepted to my thesis supervisor, Prof. Harald Schwalbe. He taught me NMR, encouraged, supported my projects with his expertise, and especially in the last years of my thesis supported my increasing independence with occasional insightful discussions.
Jens W6hnert, who would never allow me to call him anything close to my supervisor, has been of indispensable help to me ever since he joined the group. I would like to acknowledge his critical support concerning both specific as well as very fundamental aspects of my work his patience in telling me the same things over and over again (it did go both ways after all) as well as the readiness
in supplying me with any type of literature. I also appreciate his cynical sense of humor. Thanks for reading my thesis so carefully and partly even twice.
I would like to warmly thank Christian Richter for our inspiring collaboration and his continued support. It was due to his careful and friendly supervision during a lab-rotation that I decided to work in the field of NMR Later, collaborating with him in the field of pulse programming proofed
very fruitful for both of us. In addition, his great memory for whatever he had ever done made my
life so much easier, as did his easy manners and his continuous kindness, no matter if a shaped pulse was required or a ride home.
.... THE THESIS COMMITTEE
A am grateful to Prof. Bob Griffin, Prof. Barbara Imperiali and Prof. Joanne Stubbe for their
support in the course of this thesis, in respect to science as well as in coordinating the whole procedure.
.... THE JAN GUY
I could not be more grateful for your cheerful temper during this last month of writing. Thanks for being there, making me worry less, getting me out for a coffee break, feeding me with junk-food and reading this stuff over and over again. You are the best.
....THE FAMILY
I am deeply grateful to my mother, who always wanted me to go study and become a PhD. I am even more grateful that in the end she really believed, I would make it. Thank you for starting to study medicine, so you finally realized that I actually learnt something in the last ten years.
My brother has been always there for me during all this time. He makes me ever so happy to have a sibling that I always walk around telling people they should have more than just one child.
.... THE SISTERS IN ARMS
Luckily for me, Joanna & Suman both started writing up when I did to share good sessions in the library, deep depressions about unproductiveness and nice coffee-breaks. Thanks guys, long live the Tripod!
....THE OTHER PHD-STUDENT
For five years, I sat next to Julia, first in Boston then in Frankfurt. That's why we had plenty of time discussing almost everything from science to an appropriate retirement plan. I cherish your honest
opinions and the occasional apple that I got from you.
....THE GIRLS-LAB
I had a great time with Julia, Emily, Karla and Aph. Thanks for the special, girly atmosphere. Where else could I share my waist-line concerns and learn about tanning solutions? My special regards go
to Aph, who became a friend and impressed me both by her dedication to her work and her great
fashion sense. Thanks for your constant open ear for my worries. I also want to acknowledge Karla,
who is trying so hard to make sense of this thing called thesis at the moment. Don't worry, I did it, so you can, too.
.... THE GROUP
The continued good accord in the whole Schwalbe group rendered this PhD period a very pleasant time for me. The aptitude to help other group members, the sense of a common goal and the extension of the work group into private activities were really exceptional. Among all these people my special regards go to Christian Schlbrb, who was a very friendly and good humored fellow system administrator and will certainly do a great job in the future, if he doesn't decide to finish his PhD,
Boris FiIrtig and Jonas Noeske, who were very good undergraduate researchers and fortunately
decided to join the group and Elke Stirnal, who patiently purified my RNA on the HPLC Last but
definitely not least, I would like to thank Frau Paulus, who is simply the best secretary ever and the
gute Geist of the work group. The same goes for Zim, who was always there for helping out with the
spectrometers or anything technical, which is amazing considering his difficult working environment.
.... THE BOSTON PEOPLE
My deep-feklt gratitude goes to my friends in Boston, who helped me to find a start in this new environment. You contributed a lot to the great experience that these two years abroad constituted for me, which I still remember in happy melancholy. Especially, I would like to thankJulie, the social wizard, who introduced me to squash as well as to her very diverse circle of friends and thus gave me
an easy start. In addition, I want to acknowledge Peter and his great explorer spirit. If it wasn't for
him, I would have never seen half as much of New England. He was also a great supporter during
my time at MIT. I would also like to thank the people in the Magnet Lab for their enthusiasm and
their friendliness, especially Ajay and Peter. I feel especially grateful to Chris Turner, who became a special supporter to me during my time in Boston and is one of the funniest people I have ever met.
TABLE OF CONTENTS
Abstract ... 5
Acknowledgement ... 8
Table of Contents ... 10
List of Abbrev iations ... 13
CHAPTER 1: INTRODUC ON...15
The power of NMR 16 Basic principles ... 17
Structure determination Structure dtermination...1 19 Dynamic investigations ... 26
References-30
... CHAPTER 2: F(HCN)-EXPERIMENT ... 37 INTRODUC17ION 40 I ... The glycosidic torsion angle X 40Determination of X by NMR
41
Determination of X from cross-correlation relaxation rates 42 Parameterization of F(X) 44 Definition of X in NMR structure determination 45 Molecules under investigation: The 14-nt and the 30-nt RNA 45MATERIAL AND METHODS ..
47
Sample preparation 47
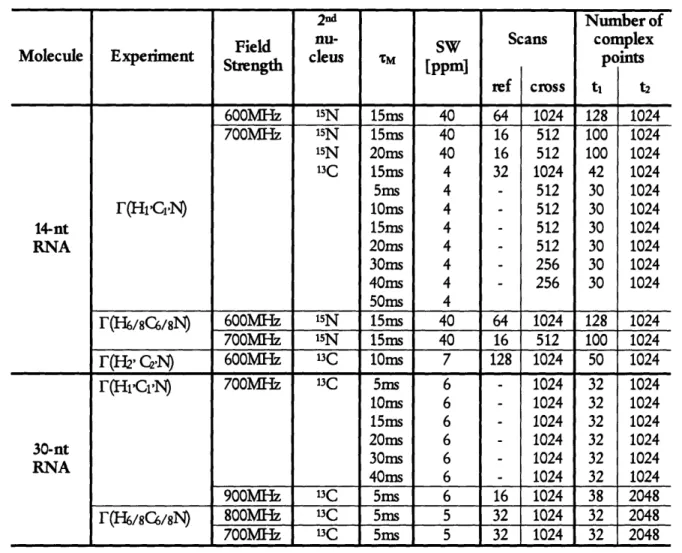
NMR spectroscopy
47
F-rate calculation 49 RNA structure calculation 49 R E SU LTS ...51The quantitative F(HCN-experiment 51 The quantitative F(H2'C2'N-experiment 68 CONCLUSIONS ... 1
Pulse Sequence Development 71 Experimental Quality 72 REFERENCES ... 75
CHAPTER 3: RNA TETRALOOP DYNAMICS ... 79
INTRODUCTION
,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,82
The tetraloop RNA secondary structure motif
82
Dynamic investigations on RNA by NMR
85
Autocorrelated relaxation rates 88
Model-free formalism 89
Motional models 92
MATERIALS AND METHODS...94
MATERIALS AND METHODS ~~~...9NMR spectroscopy
94
Relaxation data analysis 96
RESULTS ... 99
Temperature dependence of the imino proton resonances 99
13C relaxation analysis 101
DISCUSSION
.
... 131
Model-free analysis 131
Dynamics of the YNMG tetraloop motif
134
OUTOOK .139
REFERENCES
,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,140
CHAPTER 4: STRUCTURE OF THE TCR -CHAIN ...
145
INTRODUCTION.148
INTRODUCTION ~~...4T-Cell Activation 148
The TCR ;-chain 149
TCR q /Nef interaction 150
Protein backbone resonance assignment 152
MATERIALS AND METHODS
MATERIALS AND METHODS ~~~...15154
Protein expression 154
Backbone assignment of free TCR q 154
Concentration dependent experiments 155
Temperature dependent experiments 155
~/Nef interaction studies 155
TCR q in LMPG micelles 156 RESULTS RESULTS ... ...5 157 TCR ~ in solution 157
TCR bound to IMPG-micelles
169
CONCLUSIONS ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,174
REFERENCES
.,,,,,,,,,,,,,,,,,,,,,,,,,,176
APPEN D IX
...
179
A1. CHAPTER 2: F(HCN)-Experiment----Pulseprograms F-rate calculation References
A2. CHAPTER 3: RNA Tetraloop Dynamics. Pulse sequences
Extraction of peak intensities
... 181 181 184 195 ... 196 196 202
A3. CHAPTER 4: Structure of the TCR c-chain ... 203
Assignment of free TCR 4 at 278K 203
Assignment of free TCR ~ at 317K 205
Assignment of TCR tLMPG at 310K 207
LIST OF ABBREVIATIONS
13C carbon- 13 15N nitrogen- 151H
hydrogen- 1 (proton)
31pphosphorous-31
A
adenine
C
cytosine
CD
circular dichroism
cm
centimeter
CS
chemical shift
CSA
chemical shift anisotropy
CSI
chemical shift index
CV
coxsackie virus
DNA
desoxyribonucleic acids
F
cross-correlated relaxation rate
G
Gauss
G
guanine
hetnOe
heteronuclear nuclear Overhauser enhancement
HIV
human immunodeficiency virus
HSQC
heteronuclear single quantum correlation spectrum
Hz
Hertz
INEPT
insensitive nucleus enhancement bypolarization transfer
ITAM
immunoreceptor tyrosine-based activation motif
J
scalar coupling constant
K
Kelvin
KD
dissoziation constant
kDa
kilodalton
LMPG
myristoyl-phosphatidyl choline
MHz
megahertz
mM
millimolar
PM
micromolar
MQ
multiple quantum
MRI
magnetic resonance imaging
Nef
negative factor
nOe
nuclear Overhauser enhancement
ns
nanoseconds
NS
number of scans
ppm
parts per million
ps
picoseconds
R
1longitudinal relaxation rate
R2
transversal relaxation rate
RDC
residual dipolar coupling
RMSD
root mean square deviation
RNA
ribonucleic acid
S
2square of the generalized order parameter
SIV
simian immunodeficiency virus
SLD
stem-loop D
TPPI
Time Proportional Phase Incrementation
SW
spectral width
Tc
molecular rotational correlation time
TCR
T-cell receptor
TMSP
3-trimethylsilyl propionic-2,2,3,3-d
4acid sodium salt
TOCSY
total correlation spectroscopy
TROSY
transverse relaxation optimized spectroscopy
U
uridine
Chapter 1: Introduction
CHAPTER
1
Introduction
TABLE OF CONTENTS
1.1 The power of NMR ...16 1.2 Basic principles ... 17 1.3 Structure Determination... 191.3.1 Structure determination process ...19
1.3.2 Structural Parameters ...19
1.3.3 RNA structure determination ...22
1.4 Dynamic Investigations ...26
1.4.1 The importance of dynamics in the structure-function relationship . ...26
1.4.2 Dynamic information from NMR ...28
Chapter 1: Introduction
1.1 The power of NMR
Nuclear magnetic resonance (NMR) spectroscopy constitutes a non-invasive technique that allows for the spatially resolved investigation of objects of the size of a view Angstrom up to the entire human body (Figure 1). Two fundamentally different techniques have emerged: While the observation of the same spin in different environments yields super-molecular spatial resolution in magnetic resonance imaging (MRI) and NMR niCat, the investigation of different spins in the same environment constitutes the basis of NMR sptrcpy. In MRI, the spatial distribution as well as the specific properties of water hydrogens is obtained within different tissues or cell compartments through the application of magnetic field gradients, thus creating highly resolved, non-invasive two- or three-dimensional images from opaque, water-containing objects such as the human body. MRI therefore constitutes a powerful medical tool both for research and for diagnostic purposes. NMR microscopy represents the high spectral resolution variant of NMR, allowing for the detailed study of objects of the size of a view millimeters down to individual cells. In contrast to other microscopy techniques, NMR microscopy does not require transparent objects and can be extended to the observation of specific compounds such as metabolites.
-- '.':-- :' " I. 'i::.' ... . .. i : . '.- -. 'MRi.:: .... ... spe
'-..i'::.'.:'
"?'.: , O ,-.:'
,... ?....-...;.0,'-. .: '!'::.':; -::.' .'. . -': ;: :'.''.:.. i-:1 '~-" :,? ~'-!' -.?/..: mm <.::' : . > i . :-'-~~~~~~~~~~...
"':.-103' "i '-<'f.:.".:''.:.:".:....':~i 1.:.":~.".'.':"-~~~~~~~~~~~~~~...
' i .'; '..'. :. : . .';.L. .. :' ...;'i ... , ,:.;!;.:.;;! ' ' :",,i'v .;',. D;: ! ..:: * _ 1 0** *-- 'N A X-','. ' V ' :.' "''''" i^. ': f'' s 2 inm : !;;, :; ,.NM; : ,,- -, I; ,..'.: '~SS4Oj~
nm ;
Id t
,N:--V N-
. ...
....-
i; ;..
Phttoy~
pmu-
....
i- o
.i: . ... ,.:,Chapter 1: Introduction
The major disadvantage of MRI techniques lies in their relative insensitivity, which they share with all NMR-techniques. Hence, if substances other than the bulk water are to be observed, appropriate isotope labeling schemes have to be applied, while substances of low in ziw concentrations may not be detectable at all.
In contrast to MRI techniques, which directly obtain an image of the investigated object from the spatial distribution of one kind of spins, NMR spectroscopy produces an indirect image of its object
through the observation of different kinds of spins and their interactions. From a network of
spin-spin interactions, the image of the object is then reconstructed. The observation of different spin-spins
allows for the investigation of objects on the molecular level. Due to the lack of spatial resolution
and sensitivity of this technique, homogenous sample conditions are required. Hence, while it is possible to conduct these studies under physiological conditions within the living cell [7], sample inhomogeneities within the cell render this method unfavorable. In addition, NMR spectroscopy is a
rather insensitive technique and therefore requires high sample concentrations. Consequently, the lion's share of NMR spectroscopic investigations so far have been conducted in zto. To date, the use of isotope labeling techniques with 13C, 5N [8] and 2H [9-11], the development of sensitivity and resolution enhanced NMR techniques [12-14], the provision of novel structural NMR parameters
[15, 16] and the use of high field magnets have been combined to render the study of proteins of
around 30kDa and oligonucleotides of around 10kDa a routine task, while molecules of up to 80kDa
[17] can be studied in high detail in special cases. Due to their unfavorable relaxation behavior, membrane proteins are still difficult to study in solution state NMR spectroscopy. In contrast, solid
state NMR allows for the study of membrane proteins within the physiological environment of lipid
bilayers [18, 19]. This technique, however, still suffers from line broadening and low sensitivity compared to liquid state NMR.
1.2 Basic principles
In the following, a brief introduction to the basic principles of NMR is given. For a general
introduction to NMR, see ref. [20].
NMR experiments are based on the manipulation and detection of weak sample magnetizations that arises from the interaction of spin-carrying nuclei with a static magnetic field. The simplest NMR active nuclei are those with a spin of ½. Spin h nuclei can occupy either of two distinct energy levels or spin-states. In the presence of an external magnetic field, the degeneracy of these spin-states is removed. The spins will align either parallel (lower energy state) or antiparallel (higher energy state)
Chapter 1: Introduction
to the magnetic field. In the classical picture, the spins can be imagined as precessing around an effector axis oriented along or opposite to the magnetic field. The precession frequency
(Larmor-frequency) is the product of the magnetic field at the position of the spin and a proportionality
constant y, the gyromagnetic ratio, which reflects the magnetic properties of the nucleus. Since the magnetic field experienced by a given nucleus constitutes the sum of the external magnetic field and the field generated by the surrounding electrons, resonance frequencies reflect the chemical composition as well as the conformational arrangement. Under equilibrium conditions, the higher population of the lower energy spin-state results in a weak excess magnetization oriented along the magnetic field, which can be manipulated by the application of radio-frequency pulses at a frequency
close to the Larmor-frequency of the nuclei. By application of the appropriate pulses, the
magnetization can be rotated into the plane transversal to the magnetic field, where its precession frequency can be detected. Due to their different Larmor-frequencies, spins in different chemical environments are detected at different frequencies, and spectra can be obtained. Since the
Larmnor-frequency or resonance Larmnor-frequency of a spin is dependent on the spectrometer field-strength, the
frequencies are normalized to the magnetic field and then called the chemical shift, which is quoted in parts per million (ppm).
Perturbation of the equilibrium magnetization also constitutes the starting point for spin-spin
interactions through different through-bond as well as through-space mechanisms (section 1.3.2). These effects give rise to spectral resolution in multidimensional spectra from the correlation of different chemical shifts. In addition, spin-spin correlations relate spins in space or through bonds and therefore constitute the basic principle of any structural NMR spectroscopic study.
Spin 1h2 particles are the ones most widely studied by NMR. Apart from 1H (protons) and 31p, the
spin ½ isotopes of most elements occurring in macromolecules are not very abundant. In the case of
carbon and nitrogen, isotope enrichment techniques with 13C (natural abundance 1%) and "5
N
Chapter 1: Introduction
1.3 Structure Determination
1.3.1 Structure determination process
Structure determination by NMR is based on the sequence of procedures depicted schematically in Figure 2. At first, a sample of the molecule of interest needs to be prepared in quantities of several milligrams featuring the desired isotope labeling scheme. In general, several samples with different labeling schemes are required. After preparation, the desired NMR spectra are obtained from the samples. Spectra analysis requires the resonance assignment in order to connect the resonances within the spectra to the spins in the molecule. The NMR parameters, on which the structural investigation is based, are then obtained from the assigned spectra and subjected to a structure calculation procedure. ,.
-.
....mplo
.
'~., '.-.
: '-:;.: .:. .Spla-a .
$^
". "./. -;r"paratlon. : Acquisition ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~....
...
. Structure .I . . . Structure .. .. .:... --* Calcultion 6616h~on -Figure 2: Stmawue d im by NMR.1.3.2 Structural Parameters
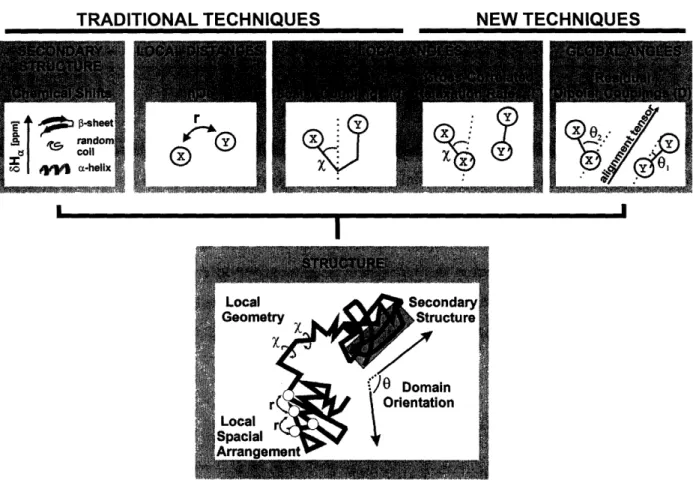
The acquisition of structural parameters from NMR-experiments is possible due to distance or angle dependent spin-spin interactions. The most common structural parameters are summarized in Figure 3 and briefly discussed in the following:
In general, distance information is obtained exclusively from dipolar through space interactions
(nuclear Overhauser effect or nOe [24). The intensity of a nOe between two nuclei is dependent on
the inverse sixth power of the internuclear distance. Proton-proton nOes constitute the main source
of information in NMR structure determination [25]. They, however, only report on distances smaller than 6
A
and therefore fail to report on global structural arrangements.Chapter 1: Introduction
TRADITIONAL TECHNIQUES
I
Figure 3: Structural NMR paranreters and dxheir inpliatin instmcture detemina
The same applies to scalar coupling constants (J-couplings) [26]. J-couplings are angle dependent on through-bond interactions. While three-bond vicinal J-couplings (3J-couplings), which report on dihedral angles, represent the most widely used structural parameters [27], also geminal (2J) and one-bond (J) couplings can contain a measure of structural information [28-30]. The structural content of scalar couplings cannot be extracted directly but relies on semi-empirical parameterizations, the so-called Karplus-equations [31]. For a number of structurally interesting J-couplings, e. g. for 3
J(HN-H-) in the protein backbone, these Karplus relations have been developed extensively[32-35].
Cross-correlated relaxation rates (F-rates) have only recently been introduced as structural NMR parameters [15] and since their introduction have found wide-spread application in structural studies on proteins and as well as nucleic acids [12, 36-41].
F-rates arise from the angle dependent through space interaction of two dipoles (Figure 3), a dipole and the chemical shift anisotropy (CSA) of a nucleus or two CSAs. Although the chemical shift anisotropy is averaged out in solution by rapid tumbling and only the isotropic chemical shift is observed, the CSA results in relaxation relevant fluctuating magnetic fields at the position of
NEW TECHNIQUES
Chapter 1: Introduction
J-couplings is redundant. Since F-rates are, however, through space in nature, they do not depend on through bond correlations and are in principal distance independent. Furthermore, while scalar coupling constants are independent of molecular size, F-rates are sensitive to the size and shape of the molecule as well as its dynamics. While these dependencies can complicate the structural interpretation of F-rates, in the study of larger systems their size dependence constitutes a major advantage compared to J-couplings.
Residual dipolar couplings (RDCs) represent the most easily accessible and therefore most widely
used long-range structural NMR parameter. RDCs report on the orientation of two dipoles in
respect to a molecular orientation tensor (Figure 3) [16, 42, 43]. The dipolar interaction between two nuclei depends on the angle between the internuclear vector and the magnetic field and can be of the order of several Kilohertz. In isotropic solution, dipolar interactions are averaged out due to rapid
molecular reorientation and are only maintained in the nOe and their contribution to nuclear
relaxation. Weak alignment by orienting media such as phage or lipid bilayers leads to the adoption of a preferential orientation of a fraction of the molecules, thus rendering the measurement of scaled
or 'residual' dipolar couplings possible. These RDCs are of the order of several Hertz and can be translated into structural information through a molecular alignment tensor, which defines the extent
of alignment and the orientation of the molecule with respect to the magnetic field. Since RDCs describe the orientation of each dipole in respect to this alignment tensor, relative orientations between different dipoles can be obtained irrespective of the distance. Similar information to the ones obtained from RDCs can also be extracted in the same fashion from the CSA [44-46].
Apart from these spin-spin interactions, the chemical shift of a spin by itself is sensitive to its environment and therefore carries structural information. In proteins FL, Cu, Co and C chemical
shift deviations from standard random coil chemical shifts measured semi-quantitatively in the form
of the so-called chemical shift index (CSI) constitute a very sensitive probe for the presence of
a-helical of 13-sheet secondary structural elements [47-49]. In addition, chemical shift changes constitute sensitive probes for structural rearrangements involved in conformational transitions or interactions.
In this thesis, a structural investigation of the C-chain of the T-cell receptor has been carried out restricted to standard protein assignment experiments in order to obtain chemical shift based
secondary structure information as well as to study the interaction site with the HIV-protein Nef
Chapter
1: Introduction1.3.3 RNA structure determination
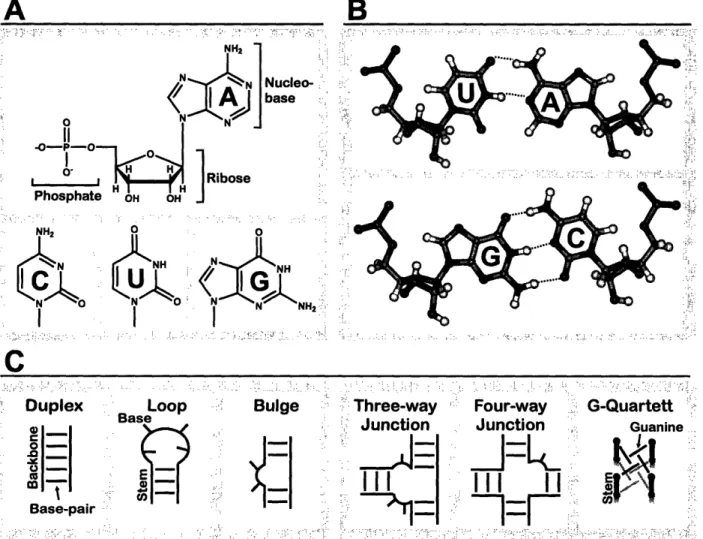
Ribonucleic acids (RNAs) are composed of a sequence of four different nucleotides, adenosine (A), cytosine (C), guanosine (G) and uridine () (Figure 4A), linked by phosphodiester bonds. The ability of the nucleobases to form hydrogen bonded pairs constitutes the basis of secondary structure formation. While regions with Watson-Crick base pairs (A paired to U, C paired to G; Figure 4B) fold into helical duplex structures, loops are formed when the RNA strand folds back on itself and bulges arise from unpaired regions within the stem (Figure 4). These basic secondary structure elements are structured to a varying degree, involving non-canonical (non-Watson-Crick) base-pairing as well as hydrogen bonding interactions of the ribose moiety and the phosphodiester-backbone.
A
",~i'
~~
-.NH
2
N Nucleo-:(IA]abase
: P' 0 NJ
III
0~ i m.i.JI
RiboseI
H I I
"H
Phosphate OH OH i r .':... ... . .. ' ".:"": f-:" '". --- ' = :::.-:B
-.. ... :' , .:.: - .... . : I.' : ... '.' ''-.. , ':'.,: .. ... ::" :- ..,:' .. . .. . ,:.!):.i'" -. :i.:.
.--. NH2 u NH2 NH N' N I I I
C
:: -. --/..-~ ... ' .- "-:..-'-- .... " ... .. '- .. ::.. -:...-..":.. .. ..:~. ::!: ... .. :..: .. : '.- !.-'':': .:.. .:.. ''.:..'.. '=.- ;'' ~.: "i' :'."'..l', .:.-:';.. ".:..Duplex Loop -- Bulge ''Three-way ' Four-way ' G-Quartett ',
,
..
Base
'
IJunction Junction
Guanine.
0--.0
-I-I
|
2 J~l 00S_10
'I~~~~~~~~~~~~~~~~~7
J=QI0l
Base-pai
II
;
=
.--7?
i
Figure 4:RNA srumo A Nudetde sm
B: WatscnCride
basepas. C Semdarystru h
bo
xks (left)
and nve owplwaiud
senivysctim nag (nit).
. ...
:'. I I
Chapter 1: Introduction
Apart from a small number of special secondary structure motifs, like the G-quartett, more complicated secondary structure arrangements such as three- or four-way-junctions (Figure 4C) can be derived from these three basic secondary structure motifs.
Since the investigation of RNA by multidimensional heteronuclear NMR methods was made
possible due to the development of techniques for the preparation of
3C and "
5N isotope labeled
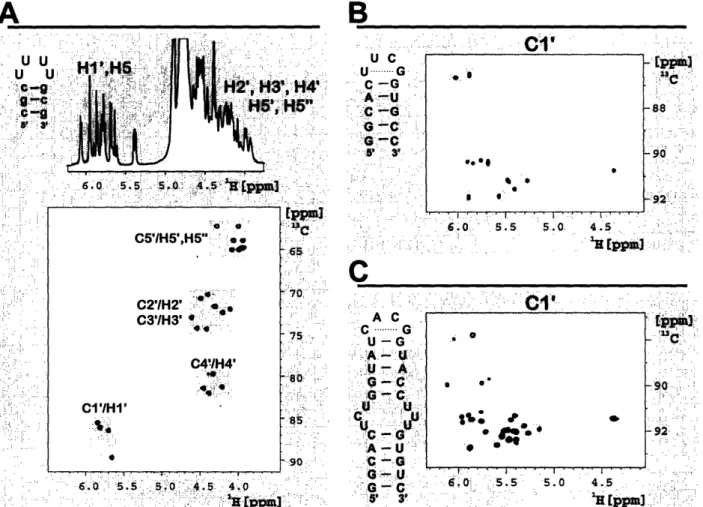
RNA in 1992 [21-23, 50], a large number of NMR experiments has been devised [51, 52] aimed at improving the accuracy of NMR-derived RNA structures. However, the size-limit of RNAs studied routinely by NMR is still restricted to around 10kDa (- 30nt). This size limitation is due to the inherent low spectral resolution of oligonucleotides. As depicted in Figure 5A for a 10-nt RNA with selective 3C labeling of the four loop residues [53, 54], the poor spectral resolution of proton 1D spectra can be overcome by 2-dimensional correlations with the attached carbon nuclei.
. . . ;,..- .-.' . . .' - .. .. i' .;:.'HS";:'.: 6.0 .5 5 4 1
H--
-s , .... ., ., ,..,.. - - : ~C5'/H5',H5" ---N.
. : : : ~~~~~~~~~65-C2'/H2' C3'/H3'...-'=75 -
.... ..7 .: ' C4!IH4';
..~~~~~~~~~~~~~~~~~~~~~~~[:
.";".-i:
..
/
::
-C'/H1'.. :
-1-4.~~~~~~~~~~~~~~~~~~."...
85 ; ' ,-! .-. :'- ,' " ... ... .. ... ...--:....
.:.0 5 .
:5, 0'4.
4
4:-.-'
: :.
.. ~~~~~~~~~~ ...: .^.,^.,... .--u;* --- ,.B
Cl,
U C u c -. U G-,-C-G
!' .- '.:---U -' ., .;.'-.. .. . : . ° ... a. .... G C .....
.--. : :.
C:"...
... .A ".. U" :'C :G '.
..'. :
-
G-G' '-:
.: '
~., ': ';,-1:
6 a:- .' '.'. :1,
55'J .'.' 1.':..
50 .... . '. ... .s.':
'- ".
-3:
'
.'.:
"..
-
'.'
!
.:
".:
:- . .... . . ... -AC -GUA ::-G-C - . . . 5' 3' .. . . . . '- .. -'' .. ' .' .... .. .: .. ,.45
-.';.:.
.'.-... .. .;-..:... -sc .... . '.... ... :.... ... . -.. ....... :92 " . -: .'. 4 5~~~~~ - :- ; . .. ;:-.. " ... .: ....; 4.5-4,5 .. ..'..: :..'Figure 5: 1D prnm spenn
ad
(t)
'H,
1CI (tqfd'
e risbe
ramties
f
a 10-nt hapn ith sdect
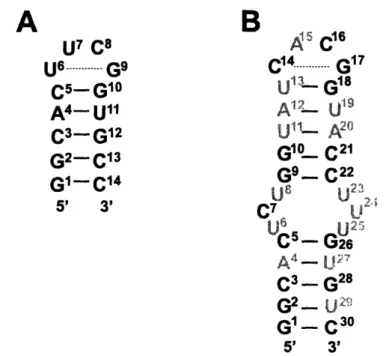
13C laktbeling in thef lct residues (). 'H, 13C-ordati (H, C- rgi of a fidly '13C lali 14-nt (B) ard 30-nt (C) RNA haitpin The lo ard dosing basepair residues are higblild in de speanan
A
IJ 11]IFtnhII
| '-U U.' u^J f ui. .. ... .: .:i.i. .-.. !. .. - * : 1..C....t-... . t-...
.
Chapter 1: Introduction
In contrast, for larger molecules, like a fully 13Glabeled 14-nt (Figure 5B) and a 30-nt hairpin RNA (Figure 5QC, the spectral resolution becomes increasingly poor. Dispersion is obtained mainly for the non-canonical loop regions, which exhibit structure specific resonance patterns (see grey shaded resonances in Figure 5B,C), while the resolution of canonical duplex structures resonances is small.
Spectral overlap arises from the small number of building blocks of partly similar or even identical chemical constitution as well as the limited diversity in secondary structural motifs. Although spectral crowding can be overcome by the application of elaborate selective isotope labeling schemes [53], this is work as well as cost intensive.
A
B
1
C
Figure 6: Tosion angl (4) and NMR-atew mndez (B) in dte phapt ter dei ck and e riboie ty of RNA. C
Bases in RNA. The mdedzsf are pai it tir pesiz WatsmoCride Kdren b g panrs; amiei (A) it
uradle (U) and guarni (G) ith cyeid (C). Pnta are shded in gre prom, ubich arm ecang uith the sdolet O er
am addioUy mk ed by ba str . ... .: .. , . , .. '. :' "'. , ., . l . .. I . ... , : ,: o
Chapter 1: Introduction
In addition to the problem of spectral resolution, the density of structural NMR parameters in
RNA is small compared to its degrees of freedom In the absence of conformational averaging, the structure of a single oligonucleotide can be described by six dihedral angles (-c) in the backbone, two parameters for the cyclic ribose (the pseudorotation phase P and the maximum amplitude v-)
and the glycosidic torsion angle X, which defines the orientation of the nucleobase in respect to the
ribose (Figure 6A). Due to the depletion in NMR-active nuclei (Figure 6B), the backbone conformation cannot be sufficiently described by conventional NMR parameters such as 'H-H nOes and lHI,1H-, 13G and 1H,3 1P-coupling constants. In particular, the angles a and ( cannot be
determined. Only recently, a method based on GH dipole-dipole, 31P-CSA cross-correlated relaxation has been developed to address a and 4 [55].
In contrast, the ribose conformation can be determined from homonuclear
3J(H,) coupling
constants [56-58] or GH dipole-dipole, C-H dipole-dipole cross-correlated relaxation rates [59, 60].In the case of conformational averaging, heteronuclear 3J(CH) and 2J(C,) coupling constants [2, 58, 61, 62] can complement the structural characterization of the ribose.
The glycosidic torsion angle X, is conventionally determined from nOe intensities between base and ribose protons [63] or 3J(C,) coupling constants [64-66]. The advantages of a new method based on GH dipole-dipole, N CSA cross-correlated relaxation rates [1] developed in the course of this thesis, are discussed in Chapter 2. Within the planar nucleobases, the number of protons is also scarce (Figure 6). In addition, the solvent exchangeable amino and iino protons cannot be detected unless stabilized by hydrogen bonding. The participation of a base imino proton in a hydrogen bond is hence demonstrated by the appearance of its resonance in the NMR-spectrumr In addition, the hydrogen bonding partner can be characterized by exploitation of the partial through-bond nature of hydrogen through-bonds, which gives rise to a scalar coupling between the donor and the acceptor nucleus [67]. As described in detail above, further interresidual structural parameters can be
Chapter 1: Introduction
1.4 Dynamic Investigations
1.4.1 The importance of dynamics in the structure-function relationship
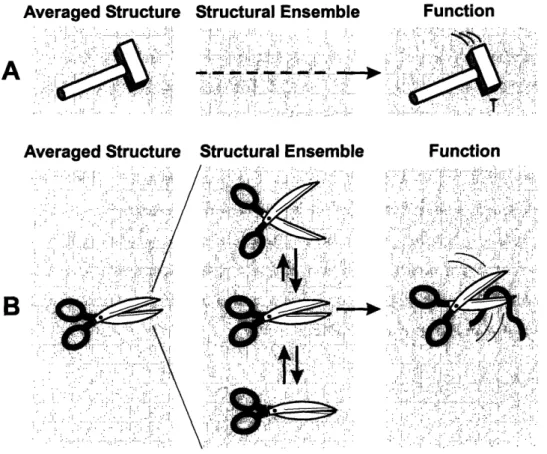
In the enterprise of understanding physiological processes, it is in general easier to find out bat something is doing than to elucidate how it exerts its function. In order to understand molecular function, detailed structural knowledge is indispensable. It is, however, a common feature of all structure elucidation techniques like NMR, X-ray or electron microscopy that they report only on either the averaged structure or the most populated structure, depending on the time-scale of interconversion. However, due to their intrinsic flexibility, molecules do not exist in one conformation alone but in an ensemble of structures, which is characterized by the populations of the respective members as well as the rates of interconversion. While in some cases the limited information obtained from an averaged structure proofs sufficient to understand a mechanism (Figure 7A), in others the average structure fails to explain the function (Figure 7B).
Averaged Structure
... . ...
Structural Ensemble
.. - -. -. .
Averaged Structure Structural Ensemble
~~~~b.
... .
B
f : ..- <:: /: :- -_ * ! ;;; '" / , -Ai ; 44A Function Function . -: - a'-- -, '' -,':,'',',!''': .... - ,-, , .'- :.,: " . '.. - ' 1. . , .- .'. ..7, .; : ,, ... : .:.:, , ... -, :, : ... :. :.: ' : ', '., , ' .. 1, .,,: A- - hi:, * ., ., a.,,.,,-. ,2.-, ., ,...:, . s . ' '.' ' ' < ' '' ' . '' '' .'1: ' .:" ': .' '. .: ..--. . .. . , ., -: . : .; ss .. ,: -: .-:-- §. 'a', ,,, .': -' .,. '- .'^' ''. ... : .. -': - ' ,' _JJj,,_','. i, ' . ' ,' ': . -,'M§,-',x'''. d',' I'd:'. . -''; ,: ''''.'' ".,''''''k'' ^ '., .. f....: .)...:,:.-.. :.- :: : A- .':. S . . .. - ' . ... ,,: . .-' ' '-.' -' '-... ., ' -. ' , - .' . ' . :-,' -! -- 9'; -- -; '' ', ' '';: 1 ' ' .' :' ', -. ', ,: '^ .- .. ^' ., .: . :',, -.'. :. : j ', .:"..' ,'. Thou',:... ... :. , - --- : . - - -.:, - -- :. ,Chapter 1: Introduction
A very prominent example for the break-down of the simplifying structure-function relationship
can be found in the catalysis mechanism of peptide bond formation in the ribosome. Despite the
availability of high resolution crystal structures [68, 69], the exact mechanism of catalysis is still highly under debate [68-73]. Also for a number of enzymatically active RNA molecules (the hammerhead ribozyme [74-76] and the lead dependent enzyme [77, 78), the averaged structure alone was
insufficient to understand the proposed catalytic mechanisms. In these cases, the active form of the
molecule, in which it exerts its function, constitutes a state within the structural ensemble, which is not observed in the structural investigation.
In order to detect this active conformation, the equilibrium within an ensemble can be perturbed in a way that the active species become directly observable (Figure 8). An example for this strategy is the trapping of enzyme transition states by binding to substrate analogs mimicking the transition state. Apart from that, a possibility of assessing the entire conformational space accessible to a molecule is to study its dynamics (Figure 8). Dynamic investigations can be carried out without
perturbation of the system and provide indirect information about the structural ensemble through
amplitudes and frequencies of deviations from the averaged or most populated structure. Therefore, dynamic information constitutes the key for deconvoluting of the structural average into an ensemble of states (Figure 7B).
Trapping
Dynamics
: ' '' -. ' 1. '' -- ;-,. ' ... i. -.- '. ,: - - . - --/ ... --!5;.."-; -- .'
--i : A d . ; ,, , . , . . -S . . ..
.-.' f-.Ad J
Figure 8:Medxs fr
st
om f emenii.er.For proteins, the importance of non-equilibrium conformations for exertion of function has been derived from the correlation of enzymatic activity and conformational flexibility in enzymes from thermophilic and mesophilic organisms [79, 80]. While these homologues exhibit widely different temperature optima, internal dynamics are similar at maximum activity. The requirement of a certain measure of flexibility for enzyme activity was assessed from the dynamic information obtained by
Chapter 1: Introduction
various techniques such as neutron scattering [81-83], NMR hydrogen-deuterium exchange [80, 81, 84-90], NMR relaxation measurements [79, 91] and molecular dynamics simulations [92-95].
For RNA, the binding mode of TAR RNA to the Tat protein in HIV could be identified as a
conformational capture rather than an induced-fit mechanism by demonstrating that the bound form
of the RNA already constitutes a part of conformational ensemble of the free molecule [96]. In this case, the structural ensemble was studied by the dynamic information included in residual dipolar couplings. This finding indicates that ground state flexibility might constitute the key precursor of adaptive binding, which is ubiquitously observed for RNA-protein interactions, and demonstrates the importance of dynamic investigations for the understanding of fundamental biochemical processes.
1.4.2 Dynamic information from NMR
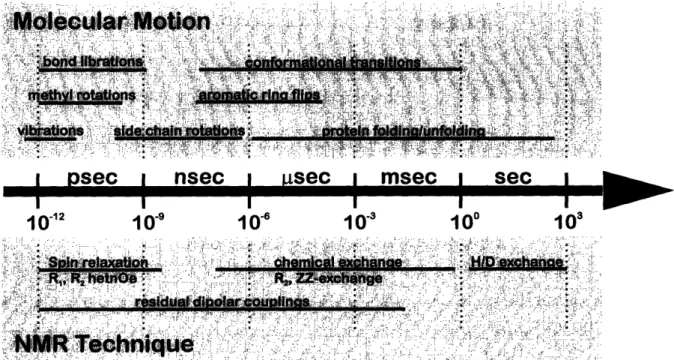
NMR-spectroscopy is sensitive to motions within a wide range of time-scales (Figure 9). In principle, every structural parameter also contains dynamic information due to its inherent sensitivity to structural changes. Apart from these side-effects of mainly structural parameters, a number of purely dynamic techniques have been developed.
1012
10-
91o-,
10-3
10
°103
Figure 9: Timsades qfndear nar and NMR-MJ e.
, r ... I Z" I-Ai;i
Chapter 1: Introduction
Different techniques exhibit sensitivity to different motional time-regimrnes. Thus, hydrogen-deuterium exchange rates report on the dynamics of solvent-exchangeable protons such as amide-and imino-protons on the time-scale of seconds to days. Autocorrelated spin relaxation rates (R1, R2
and the hetnOe) are sensitive to motions in the picosecond up to nanosecond range. In addition, R2
rates as well as ZZ-exchange spectroscopy contain information on slower dynamics in the millisecond regime. Furthermore, scalar couplings, cross-correlated relaxation rates, the nOe and residual dipolar couplings are all sensitive to motions in the millisecond regime.
Due to their sensitivity to a wide range of time-scales, autocorrelated relaxation rates constitute a
valuable tool for dynamic investigations, especially for the study of fast internal motions. In contrast
to dipolar couplings, they can be measured without perturbation of the sample with aligning media. In this thesis, autocorrelated relaxation parameters have been obtained for an abundant RNA
secondary structure motif and translated into time-scales and amplitudes of motion in order to gain
Chapter 1: Introduction
1.5 References
1. Duchardt, E.; Richter, C; Ohlenschlager, O.; Gorlach, M.; Wohnert, J.; Schwalbe, H. (2004).
Determination of the glycosidic bond angle chi in RNA from cross-correlated relaxation of
CH dipolar coupling and N chemical shift anisotropy. JAm ChemSoc, 126, 1962-1970.
2. Duchardt, E.; Richter, C; Reif, B.; Glaser, S. J.; Engels, J. W.; Griesinger, C; Schwalbe, H. (2001). Measurement of 2J(HC)- and 3J(HQC)-coupling constants by alpha/beta selective
HC(C)H-TOCSY.
J Bir NMR, 21,117-126.
3. Duchardt, E. and Schwalbe, H. (2004). Residue specific dynamic investigation of the cUUCGg tetraloop motif by NMR 13C relaxation. JAm Cheman So, submitted.
4. Albert, M.; Seebach, D.; Duchardt, E.; Schwalbe, H (2002). Synthesis and NMR Analysis in Solution of Oligo(3-hydroyAmnoic acid) Derivatives with the Side Chains of Alanine, Valine and Leucin(beta-Depsides): Coming Full Circle from PHB to beta-Peptides to PHB.
Hdreta (tinia A
ta, 85, 633-658.
5. Gee, P. J.; Hamprecht, F. A.; Schuler, L. D.; van Gunsteren, W. F.; Duchardt, E.; Schwalbe, H; Albert, M.; Seebach, D. (2002). A Molecular-Dynamics Simulation Study of the Conformational Preferences of Olig(3-hydoxyalkanoic acids) in Chloroform Solution.
Hdia
Cbii
A aa, 85, 618-632.
6. Waser, P.; Rueping, M; Seebach, D.; Duchardt, E.; Schwalbe, H. (2001). On the Solution Structure of Phb: Preparation and NMR Analysis of isotopically Labeled
Olig[(R)-3-hydroxybutanoic Acids] (OHBs). Heutica AiniAa, 84, 1821-1845.
7. Serber, Z.; Keatinge-Clay, A. T.; Ledwidge, R.; Kelly, A. E.; Miller, S. M.; Dotsch, V. (2001). High-resolution macromolecular NMR spectroscopy inside living cells.
J
A m Chem Sc, 123,2446-2447.
8. McIntosh, L. P. and Dahlquist, F. W. (1990). Biosynthetic incorporation of 15N and 13C for
assignment and interpretation of nuclear magnetic resonance spectra of proteins. Q Rev
Bi/phys, 23, 1-38.
9. Venters, R A.; Huang, C C; Farmer, B. T., 2nd; Trolard, R; Spicer, L. D.; Fierke, C A (1995). High-level 2H/1 3C/ 5N labeling of proteins for NMR studies.
J
Biond NMR, 5,339-344.
10. Leiting, B.; Marsilio, F.; O'Connell, J. F. (1998). Predictable deuteration of recombinant proteins expressed in Escherichia coli. And Bihm 265, 351-355.
11. Gardner, K. H. and Kay, L. E. (1998). The use of 2, 13C, "5N multidimensional NMR to
studythe structure and dynamics of proteins. A nnru
RevBiop)s Biond Stt,
27, 357-406.
12. Riek, R; Wider, G.; Pervushin, K.; Wuthrich, K. (1999). Polarization transfer by cross-correlated relaxation in solution NMR with very large molecules. Prc Nad A cad Sci U S A, 96, 4918-4923.
13. Meissner, A; Duus, J. .; Sorensen, . W. (1997). Spin-State-Selective Excitation.
Application for E.COSY-Type Measurement of JHH Coupling Constants. J Magn Resorn, 128,
Chapter 1: Introduction
14. Pervushin, K.; Riek, R; Wider, G.; Wuthrich, K. (1997). Attenuated T2 relaxation by mutual
cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Prrc Nat A cad Sci U S A, 94, 12366-12371.
15. Reif, B.; Hennig, M.; Griesinger, C (1997). Direct measurement of angles between bond vectors in high-resolution NMR. Saene, 276, 1230-1233.
16. Tjandra, N. and Bax, A. (1997). Direct measurement of distances and angles in biomolecules byNMR in a dilute liquid crystalline mediunm Se , 278, 1111-1114.
17. Korzhnev, D. Mv; Kloiber, K.; Kanelis, V.; Tugarinov, V.; Kay, L. E. (2004). Probing slow dynamics in high molecular weight proteins by methyl-TROSY NMR spectroscopy
application to a 723-residue enzyme.JAmCrnSc, 126, 3964-3973.
18. Watts, A.; Burnett, I. J.; Glaubitz, C; Grobner, G.; Middelton, D. A.; Spooner, P. J. R.; Watts, J. A.; Williamson, P. T. F. (1999). Membrane protein structure determination by solid
state NMR Naural PuctReport, 16, 15-19.
19. Griffin, R. G. (1998). Dipolar recoupling in MAS spectra of biological solids. Nat Strua Bid, 5 Suppl, 508-512.
20. Levitt, M. (2001). Spin Dynamics-Basics of Nuclear Magnetic Resonance. Chichester Wley. 21. Batey, R. T.; Inada, M.; Kujawinski, E.; Puglisi, J. D.; Williamson, J. R. (1992). Preparation of
isotopically labeled ribonucleotides for multidimensional NMR spectroscopy of RNA. Nudeic
A cids Res, 20, 4515-4523.
22. Batey, R T.; Battiste, J. L.; Williamson, J. R. (1995). Preparation of isotopically enriched
RNAs for heteronuclear NMR Mexs Enyrd, 261, 300-322.
23. Nikonowicz, E. P.; Sirr, A.; Legault, P.; Jucker, F. M.; Baer, L. M.; Pardi, A. (1992). Preparation of 3C and '5N labelled RNAs for heteronuclear multi-dimensional NMR studies.
NudeicA C Res, 20, 4507-4513.
24. Noggle, J. H and Shirmer, R E. (1971) The Nuclear Overhauser Effect: hemical
Applications. Academic Press: New York p. 1-259.
25. Wagner, G. (1997). An account of NMR in structural biology. Nat Stn Bid, 4 Suppl,
841-844.
26. Bax, A.; Vuister, G. W.; Grzesiek, S.; Delaglio, F.; Wang, A. C; Tschudin, R.; Zhu, G. (1994). Measurement of homo- and heteronuclear J couplings from quantitative J correlation.
Me
rds
Enzymd, 239, 79-105.
27. Bystrov, V. F. (1976). Spin-spin coupling and the conformational state of peptide systems.
P
esin NudearMagetic Resonance
Speacopy, 10, 41-81.
28. Juranic, N. and Macura, S. (2001). Correlations among (1)J(NQ' and (h3)J(NC)' coupling
constants in the hydrogen-bonding network of human ubiquitin.
J
Am Chen Soc, 123,
4099-4100.
29. Wrmer, J. and Schwalbe, H (2002). Angular dependence of 1J(Ni,Calphai) and
2J(Ni,Calpha(i- 1)) coupling constants measured in J-modulated HSQCs.
J
Biord NMR, 23,47-55.
30. Mierke, D. F.; Grdadolnik, S. G.; Kessler, H (1992). Use of one-bond Calpha.-Halpha coupling constants as restraints in MD simulations. JAm ChxmSoc, 114, 8283-8284.
Chapter 1: Introduction
31. Karplus, M. (1959). Contact Electron-Spin Coupling of Nuclear Magnetic Moments. (em P/os, 30, 11-15.
32. Vuister, G. W. and Bax, A. (1993). Quantitative
J
correlation: a new approach for measuring homonuclear three-bondJ
(HNEHlalpha.) coupling constants in 5N-enriched proteins.J
Am CJimSoc, 115, 7772-7777.33. Pardi, A.; Billeter, M.; Wuthrich, K. (1984). Calibration of the angular dependence of the amide proton-C alpha proton coupling constants, 3JHN alpha, in a globular protein. Use of
3JHN alpha for identification of helical secondary structure. J Mol Bid, 180, 741-751.
34. Ludvigsen, S.; Andersen, K. V.; Poulsen, F. M. (1991). Accurate measurements of coupling constants from two-dimensional nuclear magnetic resonance spectra of proteins and
determination of phi-angles.JMdl Bid, 217, 731-736.
35. Smith, L. J.; Sutcliffe, M. J.; Redfield, C; Dobson, C M. (1991). Analysis of phi and chi 1 torsion angles for hen lysozyme in solution from 1H NMR spin-spin coupling constants.
Bi/ety, 30, 986-996.
36. Schwalbe, HI; Carlomagno, T.; Hennig, M.; Junker, J.; Reif, B.; Richter, C.; Griesinger, C (2001). Cross-correlated relaxation for measurement of angles between tensorial interactions.
Metxs E nzyr, 338, 35-81.
37. Carlomagno, T.; Blommers, M. J.; Meiler, J.; Cuenoud, B.; Griesinger, C (2001).
Determination of aliphatic side-chain conformation using cross-correlated relaxation: application to an extraordinarily stable 2'-aminoethox3ymodified oligonucleotide triplex. JAm
JimSoc, 123,7364-7370.
38. Boisbouvier, J. and Bax, A. (2002). Long-range magnetization transfer between uncoupled nuclei by dipole-dipole cross-correlated relaxation: a precise probe of beta-sheet geometry in
proteins.JAm(emSoc, 124, 11038-11045.
39. Riek, R (2001). Characterization of hydrogen bond lengths in Watson-Crick base pairs by cross-correlated relaxation. JMagnReson, 149, 149-153.
40. Bertini, I.; Kowalewski, J.; Luchinat, C; Parigi, G. (2001). Cross correlation between the dipole-dipole interaction and the Curie spin relaxation: the effect of anisotropic magnetic susceptibility.
J
Magn Reso, 152, 103-108.41. Banci, L.; Bertini, I.; Felli, I. C.; Hajieva, P.; Viezzoli, M. S. (2001). Side chain mobility as monitored by CH-CH cross correlation: the example of cytochrome b5. J Biwri NMR, 20,
1-10.
42. Tjandra, N.; Omichinski, J. G.; Gronenbomrn, A. M.; Clore, G. M.; Bax, A (1997). Use of dipolar H-15N and H-13C couplings in the structure determination of magnetically oriented macromolecules in solution. Nat Stru Bid, 4,732-738.
43. Prestegard, J. H (1998). New techniques in structural NMR--anisotropic interactions. Nat
Stma Bid, 5 Suppl, 517-522.
44. Comilescu, G. and Bax, A. (2000). Measurement of proton, nitrogen and carbonyl chemical shift shielding anisotropies in a protein dissolved in a dilute liquid crystalline phase. J Am
CbemSc, 122,10143-10154.
45. Wu, Z.; Tjandra, N.; Bax, A. (2001). 3 1p chemical shift anisotropy as an aid in determining nucleic acid structure in liquid crystals. JAm CbemSoc, 123,3617-3618.
Chapter 1: Introduction
46. Choy, W. Y.; Tollinger, M.; Mueller, G. A.; Kay, L. E. (2001). Direct structure refinement of high molecular weight proteins against residual dipolar couplings and carbonyl chemical shift changes upon alignment: an application to maltose binding protein.
J
Biori NMR, 21, 31-40.47. Wishart, D. S.; Sykes, B. D.; Richards, F. M. (1992). The chemical shift index: a fast and
simple method for the assignment of protein secondary structure through NMR
spectroscopy. Bikxhenitry, 31, 1647-1651.
48. Wishart, D. S. and Sykes, B. D. (1994). The 3C chemical-shift index: a simple method for the identification of protein secondary structure using 13C chemical-shift data.
J
Biand NMR, 4,171-180.
49. Wishart, D. S.; Bigam, C. G.; Holm, A.; Hodges, R. S.; Sykes, B. D. (1995). 1H, 13C and 15N random coil NMR chemical shifts of the common amino acids. I. Investigations of
nearest-neighbor effects.
J
Biond NMR, 5, 67-81.
50. Varani, G.; Aboul-ela, F.; Allain, F. I-L-T. (1996). NMR investigation of RNA structure.
Prgmss in Nudear Magc Resoma e Spoctpy, 29, 51-127.
51. Varani, G. and Tinoco, I., Jr. (1991). RNA structure and NMR spectroscopy. Q Rev Biophs,
24,479-532.
52. Cromsigt, J.; van Buuren, B.; Schleucher, J.; Wijmenga, S. (2001). Resonance assignment and
structure determination for RNA. M os Enzyd, 338, 371-399.
53. Quant, S. S. P.; Wechselberger, R.; Woker, M. A.; Wbmer, K.; Schell, P.; Engels, J. W.; Griesinger, C; Schwalbe, H. (1994). Chemical Synthesis of 3C labeled RNA and DNA
monomers for the solid phase and template controlled enzymatic synthesis of DNA and
RNA oligomers. TetrahinLett, 35,6649-6652.
54.
Richter, C, En ilung eier NMR-Spektrokqisher Man
zur Bestig
der Koafim
in
i& ed in Facbbh Ceie. 1999, Frankfurt: Frankfurt, Germany.
55. Richter, C; Reif, B.; Griesinger, C; Schwalbe, H. (2000). NMR Spectroscopic Determination of Angles alpha and zeta in RNA from CH-Dipolar Coupling, P-CSA Cross-Correlated
Relaxation.JAmGJienSox 122, 12728-12731.
56. Van de Ven, F. J. and Hilbers, C. W. (1988). Nucleic acids and nuclear magnetic resonance.
EurJBike 178, 1-38.
57. Schwalbe, H; Marino, J. P.; Glaser, S. J.; Griesinger, C. (1995). Measurement of H,H-Coupling Constants Associated with Nu-1, Nu-2, and Nu-3 in Uniformly C- 13-Labeled Rna
by Hcc-Tocsy-Cch-E.Cosy.
Jul
theA nrcan Chenil Soty, 117,7251-7252.
58. Marino, J. P.; Schwalbe, H; Glaser, S. J.; Griesinger, C (1996). Determination of gamma and
stereospecific assignment of H5' protons by measurement of (2)J and (3)J coupling constants
in uniformly CG 13 labeled RNA. Janl ftheArrican ncl Society, 118, 4388-4395.
59. Felli, I. C; Richter, C; Griesinger, C; Schwalbe, H. (1999). Determination of RNA Sugar Pucker Mode from Cross-Correlated Relaxation in Solution NMR Spectroscopy. J A m GChem
Soc, 121, 1956-1957.
60. Richter, C; Griesinger, C; Felli, I.; Cole, P. T.; Varani, G.; Schwalbe, H (1999).
Determination of sugar conformation in large RNA oligonucleotides from analysis of
Chapter 1: Introduction
61. I-fines, J. V.; Varani, G.; Landry, S. M.; Tinoco, I., Jr. (1993). The stereospecific assignment of H5' and H5" in ma using the sign of two-bond carbon-proton scalar couplings.
J
AmChn
Soc, 115, 11002-11003.
62. H-fines, J. V.; Landry S. M.; Varani, G.; Tinoco, I., Jr. (1994). Carbon-Proton Scalar Coulings in RNA: 3D Heteronuclear and 2D Isotope-Edited NMR of a 3GLabeled Extra-stable
Hairpin.JA m C nSoc, 116, 5823-5831.
63. Wijmenga, S.; Mooren, M. M. W.; Filbers, C W. (1995) NMR of Macromolecules. Oxford University Press Inc.: New York p. 217-288.
64. Schwalbe, IH; Marino, J. P.; King, G. C.; Wechselberger, R.; Bermel, W.; Griesinger, C
(1994). Determination of a complete set of coupling constants in C-labeled
oligonucleotides.
J
Biord NMR, 4, 631-644.
65. Munzarova, M. L. and Sklenar, V. (2003). DFT analysis of NMR scalar interactions across
the glycosidic bond in DNA. JAm CInmS, 125, 3649-3658.
66. Trantirek, L.; Stefl, R.; Masse, J. E.; Feigon, J.; Sklenar, V. (2002). Determination of the glycosidic torsion angles in uniformly "3C-labeled nucleic acids from vicinal coupling constants 3J(C2)/4-H1' and 3J(C6)/8-H1'.
J
Bionr NMR, 23, 1-12.67. Dingley, A. and Grzesiek, S. (1998). Direct Observation of Hydrogen Bonds in Nucleic Acid Base Pairs by Intemucleotide 2JNN couplings. JAm
CnemSoc,
120, 8293-8297.68. Ban, N.; Nissen, P.; Hansen, J.; Moore, P. B.; Steitz, T. A. (2000). The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Saene, 289, 905-920.
69. Nissen, P.; Hansen, J.; Ban, N.; Moore, P. B.; Steitz, T. A. (2000). The structural basis of ribosome activity in peptide bond synthesis. Sdere, 289, 920-930.
70. Xiong, L.; Polacek, N.; Sander, P.; Bottger, E. C; Mankin, A. (2001). pKa of adenine 2451 in the ribosomal peptidyl transferase center remains elusive. RNA 7, 1365-1369.
71. Barta, A.; Domrner, S.; Polacek, N. (2001). Mechanism of ribosomal peptide bond formation.
Sdexe, 291, 203.
72. Polacek, N.; Gaynor, M.; Yassin, A.; Mankin, A. S. (2001). Ribosomal peptidyl transferase can withstand mutations at the putative catalytic nucleotide. Natue, 411, 498-501.
73. Thompson, J.; Kim, D. F.; O'Connor, M.; Lieberman, K. R.; Bayfield, M. A.; Gregory, S. T.; Green, R.; Noller, H. F.; Dahlberg, A. E. (2001). Analysis of mutations at residues A2451 and G2447 of 23S rRNA in the peptidyltransferase active site of the 50S ribosomal subunit. Pxmc
Natd A cad Sci U S A,
98,9002-9007.
74. Pley, H. W.; Flaherty, K. M.; McKay, D. B. (1994). Three-dimensional structure of a hammerhead ribozyme. Nature, 372, 68-74.
75. Scott, W. G.; Finch, J. T.; Klug, A. (1995). The crystal structure of an all-RNA hammerhead ribozyme: a proposed mechanism for RNA catalytic cleavage. CQl, 81, 991-1002.
76. Scott, W. G.; Murray, J. B.; Arnold, J. R.; Stoddard, B. L.; Klug, A. (1996). Capturing the structure of a catalytic RNA intermediate: the hammerhead ribozyme. Sdeaze, 274, 2065-2069.
77. Hoogstraten, C G.; Legault, P.; Pardi, A. (1998). NMR solution structure of the
![Figure 2 shows the angle distribution for the different nucleotides within the large ribosomal subunit [2]](https://thumb-eu.123doks.com/thumbv2/123doknet/14733885.573692/40.949.135.810.163.438/figure-shows-angle-distribution-different-nucleotides-ribosomal-subunit.webp)