Mary
Ann I,¡chert.Inc.Penicillin and
Beyond:
Evolution,
Protein
Fold,
Multimodular
Polypeptides,
and
Multiprotein Complexes
JEAN-MARIE
GHUYSEN,
PAULETTECHARLIER,
JACQUES
COYETTE,
COLETTE
DUEZ,
EVELINE
FONZÉ,
CLAUDINEFRAIPONT,
COLETTEGOFFIN,
BERNARDJORIS,
and
MARTINE
NGUYEN-DISTÈCHE
ABSTRACT
As the
protein
sequence
and structuredatabases
expand,
therelationships
betweenproteins,
the notion of
protein superfamily,
and
thedriving
forces
ofevolution
are better understood.Key
steps
of
thesynthesis
of
the bacterial cell
wallpeptidoglycan
arerevisited in
light
of
theseadvances. The reactions
through
which
theD-alanyl-D-alanine depeptide
is
formed, utilized,
and
hydrolyzed
and
the sitesof action of
theglycopeptide
and
/3-lactam
antibiotics illustrate
theconcept
according
towhich newenzymefunctions
evolve asaresultof
tinkering
of
existing proteins.
This
occursby
theacquisition
of
localstructural
changes,
the fusion into
mul-timodular
polypeptides,
and theassociation into
multiprotein
complexes.
INTRODUCTION
THE
TRANSLATION OF GENETIC INFORMATION intobiological
activity
is achievedby
the conversion ofanewly
synthe-sized
polypeptide
chain into acompact,correctly
foldedpro-tein.8
Thefolding
code is still far frombeing
understood. Nascentproteins
foldrapidly,
in secondsorless,inspite
of thefact that the timethatwould be neededtosearch all
potentially
accessible conformations is astronomical. The solution tothisparadox
is that anessential step inprotein folding
is thefor-mation ofa"molten
globule."
Thisspecies
lacks thepersistent
tertiary
interactions characteristic of the native state,but ital-ready
possesses extensivesecondary
structuresthat aremajor
elements of the native
topology.
Fromthisintermediate,
there-maining
search forcorrectfolding
isonly
over alimitedcon-formationalarea.
Moreover,
thecells possessmanyfactors thatassist
folding
and minimize and/orcorrectmisfolding
events.The distribution of amino acid residue
types
along
thepolypeptide
chain isamajor
determinant ofsecondary
andter-tiary
structures.Yet,
the number of distinct foldsadopted
by
the
proteins
is limited. Proteinshaving
25%,
ormore,of their sequences incommonadopt
thesamefoldedstructures.But,
atthe same
time,
anincreasing
number ofproteins
arebeing
re-vealed that have similar folds and
statistically
insignificant
sim-ilarities.32
Hence,
proteins
unrelatedin sequence and functionmay
diverge
froma commonprotein
ancestorwhileretaining
thesamebasic
polypeptide
fold. It has beenreported32
that2511polypeptide
chains cluster into 212 amino acid sequence fam-ilies(25%,
ormore,identities)
and intoonly
80single
domainpolypeptide
foldfamilies. In consequence,aclassification hasbeen consideredthatextends the
sequence-based superfamilies
toinclude
proteins
with similar three-dimensionalstructuresbutnosequence
similarity.
One may alsonotethat ninesuperfolds
dominate theprotein
database,
representing
morethan 30% ofall determined
structures.32
Often,
evolution obscures the func-tion.The
pathway
of the bacterial cell wallpeptidoglycan
synthesis
shown in
Figure
1is that of Escherichiacoli.Itapplies
toall bac-teriapossessing
a wallpeptidoglycan.
From the MurAUDP-N-acetylglucosamine enolpyruvate
transferase,4
whichcatalyzes
thefirst committedstepof the
pathway
inthecytoplasm,
tothepeni-cillin-binding proteins
that assemble thepolymer
from thedisac-charide-pentapeptide-lipid
IIintermediateontheouterface of themembrane,
all the reactions arebacteria-specific.
Thelipid
IIin-termediate is a
ß-\,
4-linkediV-acetylmuramyl-iV-acetylglu-cosamine
disaccharide,
theN-acetylmuramic
acid of which is sub-stitutedby
aD-alanyl-D-alanine-terminated pentapeptide
via aD-lactyl-L-alanine
amide bondand theC-l atomis attachedtotheintracellular end ofatransmembrane
undecaprenyl lipid
carrier via apyrophosphate. Lipid
IIisakey
intermediate. Itis,
atthesameCentre
d'Ingénierie
desProtéines,Université deLiège,
InstitutdeChimie, B6,B-4000 Sart Tilman(Liège
1),Belgium.
Penicillin -* Receptor
vvc
/rv
^f
"'' COOH Peptidoglycan , hydrolases luí Induction/î-lactamase
synthesis/
Mia
. 1bljG
L-Ala-D-GlupL-Xaa-D-Ala-CO
2-M M/|
x IG
L-Ala-D-Glur—L-Xaa—O-Ala—CO-NH /I—I
\w
l-Ala-D-Glur-L-Xaa—D-AlaTCO-NH i KO¡¡J^ij-assembty
(Tpase: Tglyase)-S6-cell shape fc60 —cell septation »¿9 — recycling 42-cell shape 32 -recycling UDP-MurNAclMI
L-Ala¿
D-Glu—L-Xaa-'D-Ala-D-Alà'.
NH2 -"
ib-Ala-D-Ala/^
Ddl D-Ala+D-Ala MurF MurE L-Xaa\u>
NH, MurDL-Ala-LD-Glu-L-Xaa-'.p-Ala—D-Ala)
urAk'Ren0lpyrUVate
NH2 '"--• UDP-GUNAc-enolpyruvate MurB ^NADPH MurC D-Glu L-Ala UDP-MurNAcFIG. 1. Wall
peptidoglycan synthesis pathway.
The PBPpatternshown is that ofE. coli in whichcasethe diaminoacidresidueL-Xaais
meio-diaminopimelic
acid.G,
N-acetylglucosamine;
M,N-acetylmuramic
acid;
Tpase,
transpeptidase;
Tglyase,
transg-lycosylase.
Thesites ofcleavage
ofpeptidoglycan hydrolases
areshown:1,
N-acetylmuramidase;
2,
AZ-acetylmuramoyl-L-alanine
amidase; 3,
endopeptidase.
The PBPsare inactivatedby penicillin.
Penicillin ishydrolyzed
by
the/3-lactamases.
In somebacte-ria,
/3-lactamase
synthesis inducibility
is mediatedby
a receptor(see
Fig.
7).
ThePBPs,
themajority
of the/3-lactamases,
and theBlaR-type
penicillin
receptorsbelong
tothesuperfamily
ofpenicilloyl
serine transferases.time,
theproduct
of the"cytosolic"
stageand the substrate of the "wall"stageof thesynthesis.
The Ddl D-Ala-D-Ala
ligase,
the low-molecular-masspeni-cill-inbinding
proteins
(PBPs),
and the/3-lactamases
haveonesingle catalytic
function. Somepeptidoglycan hydrolases,
theBlaR-type penicillin
receptors, and thehigh-molecular-mass
PBPsaremultimodular
polypeptides.
Sets ofPBPs andnon-penicillin-binding proteins
associate intomultiprotein
com-plexes
(not
showninFig.
1)
andformmorphogenetic
networks. Thesesystemsofincreasing
complexity
are examinedsucces-sively.
With fewexceptions,
references are madeonly
topa-pers
published
from 1993 to 1995.D-ALA-D-ALA
ANDd-ALA-d-LACTATE
(ATP:ADP
+P¡)
LIGASES
In E.
coli,
thelipid
IIintermediateis formedby
the sequen-tial addition ofL-Ala, d-G1u,
meso-A2pm,
and apreformed
D-Ala-D-Ala to
UDP-/V-acetylmuramic
acidby
theMurC,
MurD, MurE,
and MurFadding
enzymes,respectively (Fig.
1).
TheD-Ala-D-Aladipeptide
issynthesized by
theDdladding
en-zyme. The MraY and MurG transferases
catalyze
theattach-ment of the
A'-acetylmuramyl pentapeptide
tothelipid
carrier and thesubsequent
addition ofA'-acetylglucosamine,
respec-tively.
The
synthesis
of the D-Ala-D-Aladipeptide (Fig.
2)
begins
with the attachment ofafirstD-alanine residueonthey-phos-phate
ofadenosinetriphosphate
(ATP)
toyield
anacyl
phos-phate,
followedby
attackby
the aminogroup of the secondD-alanine residue to
produce
a tetrahedralintermediate,
whichthen eliminates the
phosphate
group togive
the D-Ala-D-Aladipeptide.
The DdlB D-Ala-D-Alaligase
of E. coli is made ofthree
domains,14
each folded arounda4- to6-stranded/3-sheet
core,and theATP-binding
site issandwiched between the/3-sheets
ofthecarboxy-terminal
andcentral domains. A helixdipole
andthehydrogen-bonded
catalytic
triadEl5, S150,
andY216 assist
binding
anddeprotonation
steps.
The insertion of
lipid-transported,
butasyetnon-cross-linkeddisaccharide
pentapeptide
units,
inthegrowing
wallpeptido-glycan
mustbe achievedby transglycosylation
at the level of theglycan
chains andby
transpeptidation
at the level of thepeptide
chains if the process is toyield
an insoluble network.0>. ATP D-Ala AMP
Sl50'
--.H2N
R255-H3N
s/-H'
,G276
D-alanylphosphate
"-"-'R275 D-AlaH^_r^:H-NH--.N
V216.
\ ./^C°")hN-L2
s.H^C
y
,H0
S281~°?n
H3N*CH3
H2NX/CH3
0,P2~3 -0/f~-n
0 -,__*N^
*HN:
)=o H-NHI
hpo;/
rC02 H3CCOJ
TIH3C
D-Ala-D-AlaFIG. 2.
Synthesis
of the D-Ala-D-Aladipeptide
by
the Ddl(ATP:APP
+P¡)
ligase.
isareaction in which the
carboxy-terminal D-alanyl-D-alanine
moiety
ofapentapeptide
precursorserves ascarbonyl
donor. Theglycopeptide
antibiotics bindtightly
tothedipeptide
moi-ety
ofthelipid-transported disaccharide-pentapeptide
precur-sors,
preventing cross-linking.
Resistance to the
glycopeptide
vancomycin
in enterococci results fromchanges
in thepeptidoglycan
biosynthetic
path-way.37
In VanA and VanBstrains,
thedipeptide
D-Ala-D-Ala ofthepeptidoglycan
precursorisreplaced
by
thedepsipeptide
D-Ala-D-lactate. Thischange
doesnotlimit theactivity
ofthetranspeptidase
thatcatalyzes
cross-linking,
but it resultsin,
atleast,
a1000-folddecreasedbinding affinity
ofvancomycin
forthe
peptidoglycan
precursor. The D-Ala-D-Ala andD-Ala-D-lac-tate
ligases
have 30% of their sequences incommon.14 Hence,
conversion ofone
ligase
into theotherrequires
agreat
extentoflocal
changes
but it doesnotalterthefoldtopology.
Thecru-tial E15 and S150 are
conserved,
butnotable differences alsooccurin theactive
site,
themostsignificant
onebeing
the sub-stitution Y216-H.K.The DdlB D-Ala-D-Ala
ligase
andthey-glutamyl
cysteine-glycine ligase
(or
glutathione
synthetase)
couple
activation ofan
acyl
group andhydrolysis
of ATPintoADP andP¡
topro-vide the
thermodynamic
driving
force forpeptide
bond syn-thesis. The twosynthetases
perform
different functions(glu-tathione is the
major
determinant of the oxidation-reduction state of thecells).
They
lack amino acid sequencesimilarity
(10% identities).
Yetthey
showaremarkable foldsimilarity.15
Theircommon
signature
fold andcatalytic
site may becharac-teristics of a
particular superfamily
ofADP-forming peptide
synthetases.
TheMurC, MurD, MurE,
and MurFligases
and they-glutamic acid-cysteine ligase
(the
reactionproduct
of which is the substrate of theglutathione
synthetase)
also per-formpeptide
bond formation with concomitanthydrolysis
of ATP into ADP andP¡.
They
might
be other members of thesame
superfamily.
MONOFUNCTIONAL PENICILLIN-BINDING
PROTEINS AND
j3-LACTAMASES
Serine-assisted
transpeptidation
between aD-Ala-D-Ala-ter-minated
pentapeptide
precursoracting
ascarbonyl
donor andthew-amino group of theL-Xaaresidue of another
peptide
act-ing
asaminoacceptor
doesnotrequire
aninput
of energyand,
therefore,
can resultinpeptide
bondformation atexocellular siteswhereATPisnotavailable.The
transpeptidation
reactionrequires
aprecise
protonab-straction-donation
(Fig.
3).
Instep1,
the C-terminal D-Ala-D-Aladipeptide
moiety
ofapentapeptide
precursormustbindtothe active site of the enzyme ina
position
that allows thepro-ton ofthe
yOH
of the active-site serine(S*)
to beabstracted,
theactivated
OyS*
toattackthecarbonyl
of the D-Ala-CONH-D-Ala scissilebond,
and the abstracted protontobe back-do-nated totheadjacent nitrogen
atom. In step2,
the serine(S*)
ester-linked
peptidyl
enzyme mustadopt
aconformation thatallows the proton ofthe &j-amino group of the L-Xaa residue ofanother
peptide
tobeabstracted,
the activatedÑH
toattack thecarbonyl
of theesterbond,
and the abstractedprotontobeback-donatedtothe
OyS*
atom.Backbone amino groups of theenzyme
cavity
(denoted
E-NHinFig. 3)
polarize
thecarbonyl
of the D-Ala-D-Alapeptide
bond instep
1 and thecarbonyl
of thepeptidyl
enzymeesterbond instep
2.Because the
dipeptide
D-Ala-D-Ala(in
its extendedconfor-mation)
andpenicillin
arenearly
isosteric,
thetranspeptidase
also reacts with
penicillin.
But because the scissile/3-lactam
amidebond is
endocyclic,
the serine(S*)
ester-linkedpenicil-loyl
enzyme is verylong
lived. Thetranspeptidase
is inacti-vated and behaves as aPBP(Fig. 4).
An
evolutionary
scenariohasbeenproposed18
through
whichacquisition
of new functions from aputative
DD-transpepti-dase/PBP
ancestorisachievedby
localchanges (Figs.
3 and4).
Catalyzed hydrolysis
of theesterbond of thepeptidyl
enzyme with conservation of the inertness of thepenicilloyl
enzymegave rise to the monofunctional
DD-carboxypeptidases/PBPs.
They
may control the extent ofpeptidoglycan cross-linking.
Conversely,
catalyzed hydrolysis
of thepenicilloyl
enzyme with lossofpeptidase activity
gave risetothedefensive,
penicillin-hydrolyzing ß-lactamases.
The monofunctional PBPs and the
majority
ofthe/3-lacta-masesknown
today
areacyl
serinetransferases.They
fall intoseveral amino acid sequence classes
Similarity
amongmem-bers ofa
given
class(i.e.,
intraclasssimilarity)
forms acon-tinuum,
the cut-offpoints
being
> 20% identities. Interclasssimilarity
is almost nonexistent.Similarity
isnotalways
relatedtofunction. There ismore
similarity
between theStreptomyces
Carbonyl donor TI Peptidyt enzyme R~-D-Ala-C-NH-D-Ata-COO" n" 0
R~-D-Ala-C-vOr-5*-E
Ammo x_nh acceptor -R~D-Ala-C=01'-S~E Water HOHR—D-Ala-C-NH —D-Ala-COO" R~~D-Ala-C-0)-S-E
/ N *v. " .0. oí- X o H' ~H
I
I R~-D-Ata-C-OI'-S-E -0. NH-X N N D-Ala Product R^D-Ala—C-NH-X 0 DD-transpeptidase R^D-Ala-C-OH II 0 DD-carboxypeptidaseE-S^OI'H
FIG. 3.
Acyl
transfer reactions onD-alanine-D-alanyl-terminated peptides
via formation ofa serine-ester-linkedpeptidyl
en-zyme. Attack of the
peptidyl
enzymeby
anaminoacceptorleadstotranspeptidation
of thecarbonyl
donor. Attackby
waterleadsto
hydrolysis.
TI,
terahedral intermediate.between the
Streptomyces
R61DD-carboxypeptidase/PBP
and the class C/3-lactamases
thanbetween theclass Aand the class C/3-lactamases.
The K15
PBP,
the R61PBP,
andseveral class A and class C/3-actamases
are oftwo domains(Fig.
5).18
One domain is of a-helices and the other domain is a five-stranded/3-sheet
corethat is coveredby
additional a-helices. The active site thatis sandwiched between thetwo
domains,
isadensehydrogen-bonding
networkinterconnecting
watermoleculesand the sideCarbonyl donor Tl Pencilloyl enzyme
COO" 0=C tL y o—c^7 Osu 0?
\H
I H HOHA\ H I COO" I I /SZc C00" HOH OH C-HI C00" E-S-OiH/}-
lactamaseFIG. 4.
Acyl
transfer reactionsonpenicillin
viaformationofaserine-ester-linked
penicilloyl
enzyme. With thePBPs,
there-action
stops,
atleast foralong
time,
at the level ofthepeni-cilloyl
enzyme. With the/3-lactamases,
the reactionproceeds
to
hydrolysis
ofpenicillin.
chains of amino acid residues at the
boundary
of thecavity.
This foldtopology
accommodatesmany local structural varia-tions. Withxdenoting
anyamino acidresidue,
the tetrad S*xxKis located
centrally
atthe amino end ofana-helix of the all-a domain. The triad[K/H] [T/S]G
isonthe innermost/33
strandofthe
/3-sheet
ononeside of thecavity,
and the triad SGC(the
K15
PBP),
YxN(the
R61 PBP and the class C/3-lactamases),
orSDN
(the
classA/3-lactamases)
ison aloop
connecting
twohelices ofthe all-a domainonthe otherside of the
cavity.
TheclassA
/3-lactamases
haveanadditional active-sitedefining
mo-tif,
thepentapeptide
ExELN,
locatedattheentranceof thecav-ity
nearthe bottom of the/3-3
strand.Compared
with the K15PBPand the class A
/3-lactamases,
the R61 PBP and the class C/3-lactamases
have additionalloops
andsecondary
structuresaway from the active site.
In
spite
ofdifferences infunction,
the monofunctionaldd-peptidases/PBPs
and/3-lactamases
have retained much of thesame fold and much of the same active-site
signature
in the form of the motifsS*xxK,
[S/Y]xN
oranalogue,
and[K/H/R][T/S]G.
They
alsohaveretained much of thesamecat-alytic machinery.
They
eachcatalyze
rupture
of the/3-lactam
amide bond with transfer of the
carbonyl
carbon tothe serine(S*)
residue and formation ofa serine ester-linkedpenicilloyl
enzyme. On the basis of thiscommonproperty,
they
formasu-perfamily
ofpeniciloyl
serine transferases.They
alsocatalyze
acyl
transferonacyclic
carbonyl
donorsR1
-CONH-CH(R2)-COX-CH(R3)-COOH
where X is
NH,
O,
orS. The substituentsRl, R2,
andR3,
the natureof the scissile(peptide,
ester,thiolester) bond,
and the reactionpathways
areclass- andenzyme-specific.5-21
de-pend
onthe accuracy of fit of theligands
(peptide,
ester,thio-lester,
/3-lactam
carbonyl
donors;
aminoacceptors)
in theen-zyme
cavity. Catalysis
alsodepends
ontheefficacy
with whichamino acidresidues oftheactivesitefulfill the
required
func-tion ofgeneral
basecatalyst (abstracting
the proton of theyOHS*
instep1 and that ofwateror anaminoacceptorinstep2)
andprovide
anitinerary through
which the abstracted pro-toncanbe back-donatedtotheright
atomsin each stepof the reaction.Anextensive
study
of the PBPs and/3-lactamases
of known three-dimensional structureby
site-directedmutagenesis
and molecularmodeling
has failed toidentify,
withcertainty,
theroutethat the
proton
usesduring
catalysis.17
At this level of theinvestigation,
10~10
m,quantum
effects rule the nanoworld andtheprotonshuttlecanbedisclosed
only
by
themethods ofquan-tum
chemistry.
Such methodsarebeing developed.
Studiescar-riedouton
chymotrypsin
have ledtotheconceptthat thecharge
relay
ofanacyl
serine transferase iscreated,
de novo,by
theinteracting
partners,theenzyme,and theligand.6'7
The creation of thecharge relay
results from the combined effects of theac-tive-site
environment,
the deformationundergone by
the boundligand(s),
the relaxationundergone by
the enzymepolypeptide
backbone,
and the freedom ofone orseveral watermolecules.Hence,
enzymes ofasamefamily
or even a sameclasscan use morethanoneproton-shuttle
routedepending
onstructural fea-tures of the active siteand/or
the boundligand.
Thenaturally
occurring /3-lactamases
of classesA, C,
and D and theeasewithwhich
|8-lactamase
mutants emerge among clinical isolatessupportthis
concept.
In recentyears,atleast 26ß-lactamases
ofvarying specificities
have been identified.They
each aroseby
alteration of amino acid residues in the class A TEM-1ß-lactamase. Evolution is
occurring
beforeoureyes.In
conclusion,
the monofunctionalpenicilloyl
serinetrans-ferases have evolved and arestill
evolving
withpreservation
ofa characteristic
signature
fold and much of the sameser-ine-assisted
acyl
transfermachinery. They
illustrate theprin-ciple according
towhich evolution obscures the function. Thecatalytic properties
ofamember of thesuperfamily
cannotbe deduced from its amino acid sequence and even foldtopol-ogy. Direct biochemical evidence is
required.
Finally,
the monofunctionalpenicilloyl
serine transferases arehighly
adaptable
structures.The essential serine residue may beac-tivated
by
differentgeneral
bases and several proton shuttleroutesmay be used.
KTGS
KTGS EPELN
FIG. 5.
Peptide
fold of monofunctionalpenicilloyl
serine transferases. TheStreptomyces
K15DD-peptidase/PBP
functionsas atranspeptidase
onD-alanyl-D-alanine-terminated
peptides.
TheStreptomyces
R61DD-peptidase/PBP
functionsmainly
as acar-boxypeptidase.
The E. coliTEM-1/3-lactamase
andE.cloacae P99/3-lactamase
aremembers of the amino acid sequence classAand class
C,
respectively.
The active-sitedefining
motifsare indicated. Forreferences,
seeGhuysen.18
The atomic coordinates of the K15PBPwill bepublished
shortly.
MULTIMODULAR WALL PEPTIDOGLYCAN
HYDROLASES
MonofunctionalPBPsof E. coliare, atthesame
time,
DD-car-boxypeptidases
andpeptidoglycan
hydrolases. They
hydrolyze
thecarboxy
terminalD-alanyl-D-alanine peptide
bond(made
by
the D-Ala-D-Alaligase)
of thepentapeptide
precursors.They
alsohy-drolyze
thecarboxy
terminalD-alanyl-(D)-meio-diaminopimelic
acidbond
(made
by
thetranspeptidase)
thatcross-links the pep-tide unit in thecompleted
peptidoglycan.
Streptomyces
albus Gsecretes anon-penicillin-binding
dd-carboxypeptidase/peptidoglycan
hydrolase.
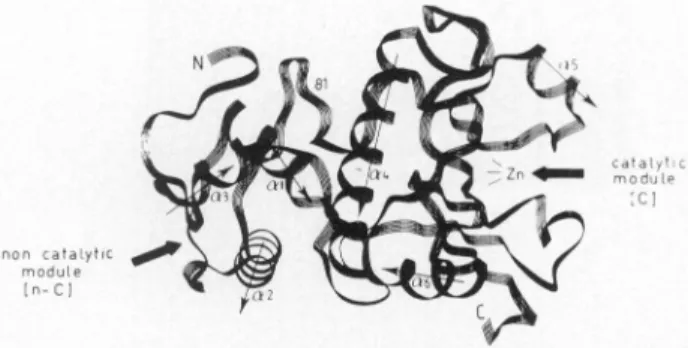
Thismétallo(zinc)
enzyme is constructed oftwomodules
(Fig.
6).19
Thearrow onthe
right points
toward thezinc-containing
active siteborneby
the 132 amino acid residue
carboxy-terminal,
catalytic (C)
mod-ule.Thearrow onthe left
points
toward thecavity
borneby
the81 amino acid residue
amino-terminal,
noncatalytic
(n-C)
mod-ule. The crevice(18.6
Â/13.5
À)
of the n-C moduleis definedby
two a-helical repeats(a2
anda3)
each 16 amino acidresidues
long,
connectedby
aheptapeptide loop.
The n-Cmodule of theZn
DD-peptidase
is theprototype
ofanamino acid sequence
family
ofn-C modules also foundin the Bacillus subtilis CwlA and B.licheniformis
CwlLA'-acetyl-muramoyl-L-alanine
amidases and theCorynebacterium
aceto-butylicum N-acetylmuramidase (lysozyme). Similarity
between the n-C modules ishigh (30-50% identities),
and, therefore,
one canbe confident that
they
have the same fold. However(with
IPdenoting
anintervening peptide)
themodulardesign
ofthe
peptidoglycan hydrolases
is different:NH2-[n-C]-[C]-COOH
for theZnDD-peptidase
NH2-[C]-[n-C]-COOH
forthe CwlA amidaseNH2-[C]-[n-C]-[n-C]-COOH
for thelysozyme
NH2-[C]-[n-C]-IP-[n-C]-COOH
for the CwlL amidaseDepending
ontheenzymes,the n-C moduleoccursatthe aminoorthe
carboxy
end of the C module inone ortwocopies
and thecopies
are eithercontiguous
orconnectedby
aninterven-ing peptide.
Acquisition
ofasubstrate-binding
module fused tothecat-alytic
module is anevolutionary advantage
forexocellularen-zymes that interact with and
hydrolyze
bonds in apolymeric
substrate.
Peptidoglycan
hydrolases
have achieved this featFIG. 6.
Peptide
fold of the bimodularZnDD-carboxypepti-dase/endopeptidase
ofStreptomyces
albus G.Reproduced
fromGhuysen
etal.,19
withpermission
ofElsevier.through
theinterchange
and local structural alterations of spe-cialized modules. Thelikely
functionof these modulesassub-strate
recognition/binding
sites woulddepend mainly
on theconserved amino acid residues. Substrate
specificity
and di-rectedtopological activity
woulddepend
ontheoccurrence ofnonconserved amino acid residues and the location and copy
number of then-C modules.
BIMODULAR PENICILLIN
RECEPTORS
Thebacterial cellsare sensitive to
virtually
everyaspect of their environment and theseenvironmentalchanges
aremoni-tored
by
specialized
sensory transducers. The dominantforms ofsignal
transductionproceed
viaphosphoryl
transferpathways
(the
so-calledtwo-component
regulatory
systems)
or areasso-ciated with sitesof
methylation
anddemethylation.
Bacteriahave
developed unique
transductionpathways
that detect/3-lactam
molecules in the environment of the cell and switch on thetranscription
of the/3-lactamase encoding
gene. Induction of/3-lactamase synthesis
in a number ofgram-nega-tive bacteria istheresult of
/3-lactam
antibiotic-induced andpep-tidoglycan hydrolases-mediated deregulation
of the cell wallre-cycling
process.26-27
Inthegram-positive
Bacilluslicheniformis,
the inductionof/3-lactamase synthesis
restsupon the presence inthe
membrane,
ofapenicillin
receptorthat results fromafusionevent
through
whichapenicilloyl
serinetransferasepolypeptide
islinkedtothecarboxy
endofasignal transducer.29
Regulation
oftranscription
of the/3-lactamase-encoding
blaP gene inB.licheniformis
involves three-chromosome-borneregulatory
genes, blal encodesa 15-kDa repressor; blaRlen-codesa
penicillin
receptor;theproduct
of blaR2 is of unknownfunction. Membrane
topology experiments, predictional
stud-ies,
andconformationalanalyses29
(unpublished
data from thislaboratory
andR.Brasseur)
strongly
suggestthat the 601amino acid residue blaR1 -encodedpenicillin
receptor BlaR has themultipartite
organization
shown inFigure
7. Central to this model is afour a-helix bundle definedby
four transmembranesegmentsTM1 toTM4. A 63 amino acid residue extracellular domainconnectsTM2 and TM3.A189amino acid residue in-tracellulardomainthat possesses the
signature
ofametallo-pep-tidase
(Zn2+
binding
site)
connects TM3 and TM4. A 261 amino acid residue extracellulardomain,
thepenicillin
sensor,is fusedtothe
carboxy
end ofTM4.Thissensorpossesses theactive-site
defining
motifs(S*TYK,
YCN, KTGT)
of thepeni-cilloyl
serinetransferasessuperfamily.
The sensor canbepro-duced
independently
from the restofBlaRas a water-solublepolypeptide
in theperiplasm
ofE. coli. Theisolatedpolypep-tide binds
penicillin
and behavesas ahigh
affinity
PBP.On thebasis of this
model,
alikely
andtestable mechanism ofsignal
transmissionby
BlaR may beputforward. Penicillin-inducedconformationalchanges
in thepenicillin-bound
sensorand the
interacting
63 amino acid residue extracellular domain would be transmitted via the four a-helix bundleto the intra-cellular domain withconcomitant activation of theputative
met-allopeptidase. Degradation
of the Blal repressoror release of anantirepressor
in thecytosol
by
the "activated"peptidase
would result inderepression
of/3-lactamase synthesis.
BlaR is the
prototype
ofanamino acid sequencefamily
ofSensor
K539TG Y476GN
S*¿02TYK
601-_I_I_L
Zn
binding
siteSignal
emissionFIG. 7. Schematic
representation
ofthe bimodularpenicillin-receptor
BlaR of B.licheniformis.
Thepenicillin
sensorbelongs
tothe amino acid sequence classD/3-lactamases.
and
low-affinity
PBP2'synthesis
inS.aureus.29
The S342-R601polypeptide
sensorof BlaRis, itself,
amember of the amino acidsequence class D
/3-lactamases (32%
identities withtheOxa-2/3-lactamase),
suggesting
a commonsignature
fold.Hence,
apeni-cilloyl
serine transferase ofagiven
class(a
/3-lactamase)
mayac-quire
a new property(penicillin sensing)
through
localchanges
and fusiontoanother
polypeptide,
andtheresulting
chimeric pro-tein mayacquire
a newfunction(gene
regulation).
MULTIMODULAR
PENICILLIN-BINDING
PROTEINS
The
high-molecular-mass
PBPsarealso of modulardesign'2
(Table
1,
Fig.
8).
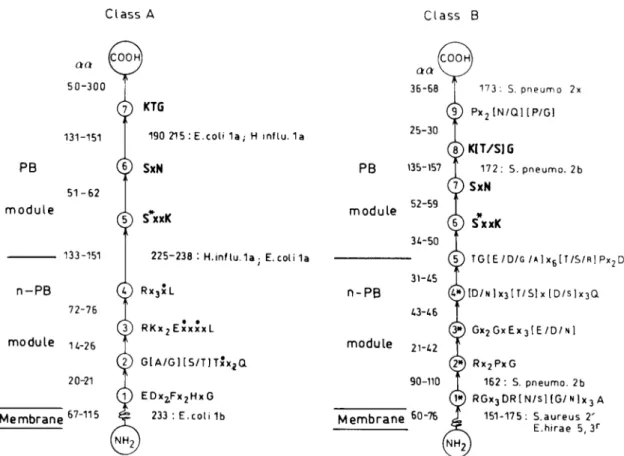
The amino acid sequence dataare from theliterature.2,12'34
A
penicillin-binding
(PB)
module thatbelongs
to andpos-sesses the
S*xxK,
SxN,
andK[T/S]G
markers of thepenicill-loyl
serine transferasessuperfamily
is fusedtothecarboxy
end ofanon-penicillin-binding
(n-PB)
module inasingle
polypep-tide chainthatfoldsontheouterface ofthe
plasma
membrane. Thepolypeptide
itself is fused to an amino-terminaltrans-membrane-anchoring
module. Insertsmayoccurthatarelarge
enough
toform additional modules.By analogy
with themono-functional
PBPs,
the PB modulesareassumedto startabout 60 amino acid residuesupstream
fromthe S*xxK motif andtoter-minate about 60 amino acid residues downstream from the
K[T/S]G
motif.The n-PB modules also possessspecific
amino acidmarkers,
but thesignature
of the n-PB modules of the classAmultimodular PBPs is different from that of the n-PB mod-ules of theclass B multimodularPBPs.
Asa
polypeptide
chain increases inlength,
finding
theright
fold becomes more difficult because the
possibilities
ofmis-folding
increase.Study
of derivatives of multimodular PBPsoverproduced
fromappropriate expression
(and secretion)
vec-tors shows that the
acquisition
ofastable,
penicillin-binding
fold
topology
by
the PB module is membrane-anchorindepen-dent but
requires
concomitantbiogenesis
of then-PBmodule. As shown with the E. coliPBPlb31
andPBP316
(unpublished
data),
the S. aureusPBP2',4041
and the E. hirae PBP5(un-published
data),
replacement
of the membrane anchorby
acleavable
signal peptide
orsubstitution of thegenuine
anchorby
another transmembraneanchoring
device has noeffect onthe
thermostability
andpenicillin-binding capacity
of the PBPmutants.
However,
elimination of both the membrane anchor and the n-PB module(or
partofit)
orelimination of then-PBmodule with conservation of the membrane anchor isnot
tol-erated.
Moreover,
expression
of E. coliftsl
genesencoding
PBP3 mutants in which El93 of motif 3* of the n-PB module is
replaced
by
DorNgives
rise tomembrane-boundproteins
that are veryunstable,
suggesting
that motif 3*plays
arole inthe
folding
process(unpublished
datafromthislaboratory
andJ.
Ayala).
Intraclass
similarity
between then-PB modules and the PBmodules,
respectively,
is acontinuum with acut-offpoint
ofabout 20% identities
(Table
2).
Interclasssimilarity
between then-PBmodules is nonexistent and interclass
similarity
between the PB modules isvery low. The PB modules of the classA PBPsandthoseof classBPBPs havediverged
sofarthattracesof
similarity
other thantheactive sitedefining
motifshaveal-most
completely disappeared.
Then-PB modules of the classA PBPs and those of the class B PBPs formtwodistinct
fam-Table 1. Sizeofthe MultimodularPBPs in NumberofAmino AcidResidues andPosition of theActive Site Serine S* ClassA ClassB S* B. subtilis
la/b
H.influenzae
la E. coli la E. coli lb M.leprae
1 S. aureus2x S.pneumoniae
la B.subtilis la S. oralis la 10. B.subtilis 4 914 864 850 844 821 727 719 714 637 624 390 452 465 510 398 398 370 359 371 388 1. S.pneumoniae
2x 2. S.pneumoniae
2b 3. E. hirae5,3r
4. S.aureus 2' 5. E. coli 2 6. E. coli 3 7. N.gonorrhoeae
2 8. N.meningitidis
750 680 678 667 633 588 582 582 337 385 422 405 330 307 312 310ClassA Class B
7) KTG
190215:E.coli1a; H inftu.la
SxN
(5)
S*xxK
225-238:H.influ.1a;
E.cotila ORx3xL
3)
RKx2ExxxxL
p
G[A/G][S/T]Txx,2Q
l)
EDx2,Fx2HxG
233:E.coli 1b PB module n-PB(cooh)
36-68 173: S. pneumo 2x9)
Px2[N/Q][P/G)
25-30(6.
135-157 172: S.pneumo. 2b SxN 52-59 34-50(5) TGtE/D/G/*lx6[T/S/H]Px2D
31-45(4<9[D/N]x3lT/S]x[D/Slx3Q
43-46(3*)
Gx2GxEx3(E/D/Nl
module 2i-42(2»)
Rx2PxG
90-110 162: S. pneumo.2b(]»)
RGx3DR[N/s][G/N]x3
A 151-175: S.aureus 2' E.hirae 5,3r Membrane 6o~76 NH,FIG. 8.
Design
and amino acid sequencesignatures
of the multimodularPBPs of classesAand B. Thedistribution of thecon-served
motifs,
the averagelength
of theintermotif sequence, and the occurrenceof inserts(expressed
innumber of amino acid residuesaa)
areshown. The data derive from the amino acid sequences of the PBPs listed in Table 1.S*,
active-siteserine;
x,variableaminoacid
residue;
x,hydrophobic
amino acid residue.Table 2. Identities
(%)
between theAligned Amino AcidSequences
ofHigh-Molecular-Mass PBPsa n-PB module PBP Class A 1. M.leprae
1 2. E. coli la 3. H.influ-enzae la 4. E. coli lb Class B 5. E.coli2 6. E. coli 3 7. E. hirae 3r 8. E. hirae 5 9. S. aureus 2' Class A 100 26 27 24 100 57 100 22 24 100 Interclass PB module Class A 100 17 16 18 100 52 100 24 24 100 Interclass 10 11 12 11 10 10 11 12 12 11 11 14 13 13 11 10 12 10 10
10
100 Class B 24 28 17 21 20 100 85 100 100 23 17 39 40 100ilies. The two families have a characteristic amino acid
se-quence
and,
presumably,
foldsignature,
suggesting
thatthey
have evolved from differentpolypeptide
ancestors.Asann-PBmodule of class A is linkedtoaPBmodule of class Aandan
n-PBmodule of class Bis linkedtoaPB module of class
B,
alikely
corollary
of thisclass-specific
modulardesign
is that the class A PBPsperform
a different function(or
differentfunc-tions)
from that(those)
of the classB PBPs. In consequence,the effects of the inactivation ofthe class A PBPs onthe cell
viability
should be different from those of the classB PBPs. Invitro,
thepurified
class A PBPla and PBPlb of E. coli(and,
perhaps,
by
extension other classAPBPs)
are wallpep-tidoglycan synthetases.
They catalyze
glycan
chainelongation
andpeptide
cross-linking
from thelipid
II intermediate.Inhibition of the
n-PB/transglycosylase
module ofPBPlbby
moenomycine
preventspeptide cross-linking
whileinactivation of thePB/transpeptidase
moduleby
penicillin
enhancesglycan
chainelongation,
showing
that thetwomodules interactwith each other.Moreover,
PBPlb containsamembrane associationsite in addition to the transmembrane
anchor31
and dimeric forms of PBPlb are in close association withthepeptidogly-can.42
Invivo,inactivation of thePBmodules ofPBPlaandPBPlb
of E. coli
by
j3-lactam
antibioticscausespeptidoglycan
hydro-lases-induced cell
lysis.
Deletion of the genesencoding
PBPla and PBPlb isfatal,
but deletion of either of the PBPla-orthePBPlb-encoding
geneistolerated,
suggesting
thatonePBPcancompensate
for another.In
vitro,
thepurified
classBPBP3 of E.coli21
(unpublished
data)
and PBP2x ofS.pneumoniae29
catalyze
serine-assistedhydrolysis
andaminolysis
of thiolestercarbonyl
donors. Bacterial strainshaving
a reducedaffinity
forpenicillin
andpossessing
one or several classB PBPs with reducedaffinity
for thedrug synthesize
a wallpeptidoglycan
with a differentpeptide moiety
from that of the wildtype.36
Hence,
thePBmod-ules of the class B PBPs are
involved,
oneway oranother,
inpeptidoglycan
cross-linking.
However,
theprecise
natureof thecatalyzed
reactions remains tobe elucidated.Indeed,
the iso-lated PBP3 ofE.coli is inertonthelipid
IIintermediate1
(un-published
results from thislaboratory
and J. VanHeijenoort).
Moreover,
evolution may obscure the function ofaprotein
andfusion between
polypeptides
may result in theacquisition
ofa new function(see
preceding
sections).
Given that then-PBmodules oftheclassB PBPshavea
dif-ferent amino acid
signature
fromthatof the n-PB modules of the classAPBPs,
they
maynothaveatransglycosylase
activ-ity.
Theacyl
transferaseactivity
of their PBmodulesmight
becoupled
with thetransglycosylase activity
of the n-PB modules ofsomeclass A PBPs. Such a situationimplies
that the classA and class B multimodularPBPsinteract with eachother. One
may also note that the E. coli PBP3 forms dimers
(personal
communication from N.Nanninga
andJ.Ayala).
In
vivo,
theprimary
effects of the inactivation of the PB mod-ules of the classBPBP2 and PBP3 of E. coliby
/3-lactam
an-tibioticsare
morphological
abnormalities,
followedby
cellly-sis. Inactivation of PBP2 results in
growth
asspherical
cells(the
rod-shaped
maintenancemachinery
is nolonger
func-tional).
Inactivation ofPBP3 results ingrowth
as filamentouscells
(the
septation machinery
is nolonger functional).
Inactivation of either of thePBP2-or
PBP3-encoding
genes ofE. coliand of the PBP2b-or
PBP2x-encoding
genes of S.pneu-moniae isnottolerated.
Like the monofunctional PBPs and
/3-lactamases,
the multi-modular PBPs arehighly adaptable
structures. Multimodular PBP-mediated resistanceto/3-lactam
antibiotics amonggram-positive pathogens
has becomea serious healthproblem.
In S.
pneumoniae,i0,23,3S
the first PBPs tobe affecteddur-ing
selection oflaboratory
mutantshaving
a reducedaffinity
for cefotaxime and
piperacillin
are the class B PBP2x andPBP2b,
respectively.
The modified PBP2x and PBP2b confer resistance upon transformation. Lowaffinity
forms of PBP2x and PBP2b and the class A PBPlaarepresent
inhighly
resis-tantclinical isolates. Reducedaffinity
is the result of structuralchanges affecting
either a limited number of amino acidresiduesor
large
blocks of amino acid sequences of thePBmod-ules.
Interspecies
recombinationaleventsoccurthatreplace
partofa
PBP-encoding
gene withthecorresponding
partsfromho-mologous
PBP-encoding
genes ofclosely
related,
naturally
re-sistant
species.
The
low-affinity
class BPBP2' ofS.
aureus3
and PBP5 and PBP3r ofE.hiraeu'33
allow the strains thatproduce
them togrow and manufacture a wall
peptidoglycan
under conditionsin which all the monofunctional PBPs andthe PB modules of all the other multimodular PBPs areinactivated
by penicillin.
The
staphylococcal
PBP2' is chromosomaland,
insomestrains,
its
penicillin-induced synthesis
isBlaR-mediated.29
Theente-rococcal PBP5 is also chromosomal but its
homologue
PBP3r isplasmid
borne in E. hirae S185R(unpublished
data).
Thisplasmid
carries,
atleast,
twocopies
of thePBP3r-encoding
gene,onecopy of thestreptomycin-resistance
markerstr,onecopy of the
erythromycin-resistance
marker erm, and severalcopies
oftheinsertion moduleIS/276,
asituationeminently
fa-vorablefor the
spread
of multiresistanceamongenterococci and related bacterialspecies.
Asthe
origin
of thelow-affinity
classBPBPz, PBPs,
and PBP3r isunknown,
the structural featuresresponsible
for the lowaffinity
are also unknown.Compared
tothe other classBmultimodular
PBPs,
thelow-affinity
PBPshave anextendedn-PB module because of the presence ofa«100 amino acid
residue insert
immediately
downstream from the membrane anchor. These inserts are related in amino acid sequence.Presumably, they
have a samefold.Expression
of genesen-coding low-affinity
PBP derivatives in which the insert ismissing
or truncatedgives
rise toproteins
that are inert in terms ofpenicillin-binding41
(unpublished
data),
showing
that the insert
plays
arole inbiogenesis.
The active-sitetopol-ogy of the PB module confers resistance to the usual
penicillins
andcephalosporins
butnotnecessarily
toall/3-lac-tam
compounds. Cephalosporin
derivatives arebeing
devel-oped
that are activeagainst
methicillin-resistant S. aureusstrains.22
A
plausible picture
that arises from the aboveanalysis
is that the enzyme activitiesrequired
tobuild thepeptidoglycan
poly-merwould be
provided by
then-PB/transglycosylase
modules of the class A PBPs and thePB/acyl
transferase modules of both class A and class B PBPs.Regulation
ofthese activities in acell-cycle-dependent
fashion would be mediatedby
then-PB modules oftheclassB PBPs. Recent
genetic
studies havebrought
tolight
the existence ofmorphogenetic
networks. These networksaremultiprotein
complexes,
theconstitutive elementsofwhicharesetsofmultimodular
PBPs,
monofunctionalPBPs,
and
non-penicillin-binding
proteins.
MORPHOGENETIC NETWORKS
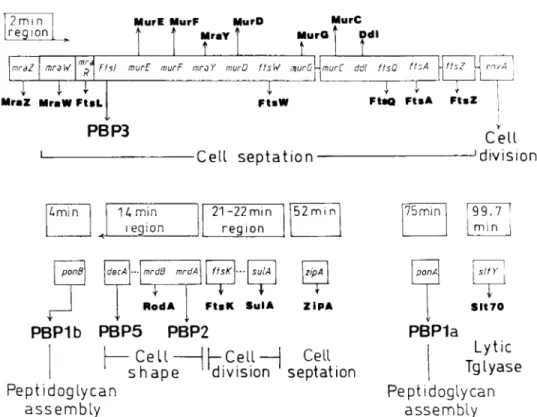
Figure
9 shows partof the geneorganization
in thechro-mosomeof E. coli. The 4 and 75 min
regions
contain the genesencoding
thebienzyme
(transglycosylase/transpeptidase)
class A PBPlb andPBPla,
respectively.
The 2 min
region,
which contains the geneencoding
the mul-timodular classBPBP3,
also contains genes that encode othercell-septation
proteins.1-9
Figure
10 shows the cellularlocal-izationofsomecomponentsof the cell
septation
network(also
called divisómeor
septator). FtsQ
is of unknown functionandFtsL containsa leucine
zipper
motif. Each isontheouterfaceof the
plasma
membrane. FtsW is anintegral
membranepro-tein.
FtsA,
anATP-binding
protein
of theeukaryotic
actinfam-ily,35
isexposed
on the inner face of theplasma
membrane. MraW is aprotein bearing
anS-adenosyl methionine-binding
motif and FtsZ isa
GTP-binding/GTPase protein.
Each iscy-toplasmic.
FtsZ contains a shortglycine-rich
segment
that isstrikingly
similarto theGTP-binding
motif oftheeukaryotic
cytoskeletal
tubulin.13
About10,000
molecule of FtsZoccurpercell.
They
formaring
at the future divisionsite39
in the form ofa*=AO-pm
filamentthat islong enough
tosurround theIN
^©0/
FIG. 10. Cellularlocalization of
proteins
of the "cellsepta-tion"
morphogenetic
network inE. coli.cell 20 times. These genes form a cluster the
expression
ofwhichis controlled
by
agearbox
and metabolicpromoters.Genes located outside the 2min
region
of the chromosome also encode "cellseptation" proteins.
SulA isacell-divisionin-hibitor.
ZipA
is anotherintegral
membraneprotein.
Slt-70isan/V-acetylmuramidase
thatcatalyzes
thehydrolysis
of theglyco-sidic bond with transfer of the
carbonyl
to the C-6hydroxyl
group,yielding
a(
1-6)anhydrornuramic
acid.25
FtsK possessesprobably
an N-terminal domain with severalmembrane-span-ning
helices andalarge cytoplasmic
domain withanATP/GTP-r-Lipid
II intermediatesynthesis
-i2min
region
MurE MurF MurD
MraY
Î
MurC MurG Odl mraZ mraWIT
mr4T
F1st murE murF mraY murD ftsW tnurG MraZ MraW FtsL
PBP3
~1T~
FtsW-Cell
septation
murC ddl flsO ftsA UftsZ
~i
i
r
FtsQ FtsA FtsZCell
Jdivision
imin 1 ¿, minregion
21-22mmregion
52min 75 min 99.7 minpond decA mrdd mrdA
1
RodA ftsK3-
sulA FtsK SulA zipA ZipAPBPlb
PBP5
PBP2
I—Cell-11—Cell—1
Cell
shape
'division septation
Peptidoglycan
assembly
ponA sit Yr
SIt 70PBPla
Lytic
Tglyase
Peptidoglycan
assembly
FIG. 9.
Organization
of the genes in thechromosome ofE. Coli involved inlipid
IIintermediatesynthesis, peptidoglycan
as-sembly,
cellseptation,
cellshape,
and cell division.ponA
andponB
arealso called mrcA andmrcB,
respectively. Tglyase,
binding
motif(L.
Begg,
S.Dewar,
and W.Donachie; report
presented
attheworkshop
"Structure,
Function andControlsinMicrobial Division." InstituteJuan
March, Madrid,
May
1995).
A
single
basechange inftsK
causes aconditionalblock in cell division that issuppressed by
deletion of thePBP5-encoding
dacA.A
battery
oftechniques
isbeing
usedtostudy
protein-pro-teininteractions within the "cell
septation
network."According
to reports
presented
at theworkshop
mentionedabove,
PBP3interacts,
presumably
through
its intracellular aminoend,
withFtsA,
SulA,
and FtsZ in thecytoplam;
FtsZ itselfinteracts with theintegral
membraneprotein ZipA;
and PBP3interacts,
pre-sumably through
itsperiplasmic
module,
withPBPlb,
PBP7(an
endopeptidase),
and Slt-70.At thesame
time,
the structuralrequirements
of PBP3 forin vivoactivity
arebeing
studiedby complementation
experi-ments. InE. coli
RP41,
thetemperature-sensitive-/ttl
2158-en-codedPBP3 has two mutations: G191-D in motif 3*ofthen-PB module and D266-Natthe
junction
between then-PB and PBmodules.1
At42°C,
E. coli RP41 grows as filaments andlyses
butrod-shaped,
cell division andviability
arerestoredby
transformation with a
plasmid carrying
thewild-type ftsl.
Incontrast,
complementation
isnotachievedby ftsl
genesencod-ing
PBP3mutantsthat either lack the membraneanchor,
havea membrane anchor different form the
genuine
one,orhave a17 amino acid residue insert
immediately
upstreamfrom R60 of motif1*(the
insertbeing
theR60-D76 sequence,exceptthat D76 is mutated into N;unpublished
datafrom thislaboratory
and J.
Ayala). Complementation
doesnotoccurinspite
of thefact that the
produced
PBP3mutantshave thesamethermosta-biltiy
andpenicillin-binding capacity
asthewild-type
PBP3. Theacquisition
ofapenicillin-binding
foldtopology by
thePBmodule of PBP3
depends
on anintact motif 3* ofthen-PBmodule but it is
membrane-anchor-independent.
Itnowappearsthat the in vivo
activity
of PBP3requires
notonly
a correctpenicillin-binding
foldtopology,
but,
inaddition,
the presence of thegenuine
membraneanchorandanintactenvironment ofmotif 1*of the n-PB module.Ithas also been
reported
that theP565-G571 sequenceatthe end
(V577)
of thematurePBP3 isnot
required
forpenicillin-binding
but is essentialfor in vivoactivity.20
Likely,
the membraneanchor,
features ofthe n-PBmodule,
and thecarboxy
end of thepolypeptide
chainaresitesthrough
which PBP3mayinteract with othercomponentsof the "cell division"morphogenetic
network.The 14 min
region
of theE. colichromosome,
whichcon-tains the gene
encoding
the multimodular class BPBP2,
also contains genes that encode the"cell-shape"
PBP5 andRodA,
aprotein
very similartoFtsW. Thiscomplex
and ribosomalac-tivities appeartobe coordinated
by
achain ofinteracting
ele-ments,oneof which is
regulated by
the nucleotideguanosine
5'-diphosphate, 3'-diphosphate (ppGpp,
an RNApolymerase
effector),
itselfsynthesized
by
theSpoT/RelA, proteins.
Remarkably,
the "cellshape"
and "cellseptation"
morpho-genetic
networks areconnected. PBP2 isnotrequired
forsep-tum
synthesis.
However,
loss of PBP2activity
results inablockof cell division
and,
in the absence ofPBP2,
celldivision andviability
are restoredby
increasing
thepool
ofppGpp
or thelevel of
FtsQ-A-Z.30
The "cell
septation"
and "cellshape"
networksareprobably
ubiquitous
in the bacterialworld,
with manyindividualvaria-tions.
Likely,
other networks remaintobe identified. Thestep-wise increased resistanceto
/3-lactam
compounds
of S.pneu-moniae
laboratory
mutantsdoesnotalways
correlate withPBPchanges,
but correlates withgenetic
competencedeficiency.23
In a cefotaxime
laboratory
mutant, increased resistance andcompetence
deficiency
aremediatedby
apoint
mutation inahistidine kinase
CiaH.23
CiaH and CiaRaremembers ofsignal
transduction
pathways including
theEnvZ/OmpR
osmoregula-tion inE. coli and theVanS/VanR
vancomycin
resistancein-ducibility
in E.faecium.
The
morphogenetic
networksare still far frombeing
under-stood.
They probably
possess severalphosphoryl
transferpath-ways.
They
maybe components ofmultiple parallel,
overlap-ping,
andinteracting
signal
transductionsystems.
There is cross-talk between thepathways.
These characteristicsaretyp-ical of
"(phospho?)neural"
networks.24
CONCLUSION
The
synthesis
andassembly
of the bacterial cell wallpepti-doglycan
require proteins
theprimary
functions ofwhich arethe chemical transformation of metabolite
intermediates,
thebuilding
ofacellular structure, and the transfer andprocessing
ofinformation
through integrated
biochemical "circuits" thatare abletotransform an
input signal
intoanoutputsignal
andanoutput
signal
intoaninput signal.
The wall
peptidoglycan
is abacterium-specific polymer.
Empirical
approaches
to thediscovery
of "bacterial-cell-wall" inhibitors and theimprovement
ofexisting
drugs
werejustified
when basicresearchwas still
struggling
tocope with thecom-plexity
of the process and the molecularstructures. The situa-tion ischanging.
With thepresentadvancedstateofourknowl-edge
and theavailability
ofexperimental
and theoretical tools ofeverincreasing
incisiveness,
future antibacterialchemother-apy
strategies
arelikely
todepend
ontheunderstanding
of thefunctioning
ofexisting
targetsatthe atomic levelallowing
newdrugs
tobedesigned,
andonthe identification of theconstitu-tive elements and the
"wiring"
of themorphogenetic
networksallowing
newtargetstobe discovered.ACKNOWLEDGMENTS
This workwas
supported
inpartby
theBelgian
programonInteruniversity
Poles of Attraction initiatedby
theBelgian
State,
Prime Minister's
Office,
Servicesfédéraux des affairesscien-tifiques, techniques
et culturelles(PAI 19),
the Fonds de la RechercheScientifique
Médicale(contract 3.4531.92),
and the Fonds de la Recherche Fondamentale Collective(contract
2.4534.95).
REFERENCES
1.