Bound or free: interaction of the C-terminal domain of Escherichia coli single-stranded DNA-binding protein (SSB) with the tetrameric core of SSB.
Texte intégral
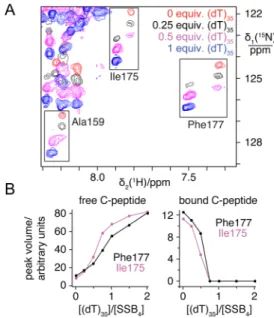
Figure
Documents relatifs
Under such conditions, the PAIN part from the simultaneous experi- ment has a negligible intensity loss compared to a separate PAIN experiment (\5%) and a good PAR spectrum is
Die Resultate der Studie zeigen, dass trotz einem erhöhten Risiko zu psychischen Folgen eines Einsatzes Rettungshelfer Zufriedenheit und Sinn in ihrer Arbeit finden können und
Nous souscrivons donc aux recommandations européennes les plus récentes concernant le traitement anticoagulant et antiagrégant des patients atteints de fibrillation auriculaire (et
Ein verwandt- schaftliches Empfinden ergab sich vor allem aus dem gemeinsamen Status einer Republik, aber auch aufgrund der ähnliehen politischen Institutionen und
C’était le moment choisi par l’aïeul, […] pour réaliser le vœu si longtemps caressé d’ accroître son troupeau que les sècheresses, les épizoodies et la rouerie de
L’archive ouverte pluridisciplinaire HAL, est destinée au dépôt et à la diffusion de documents scientifiques de niveau recherche, publiés ou non, émanant des
L’archive ouverte pluridisciplinaire HAL, est destinée au dépôt et à la diffusion de documents scientifiques de niveau recherche, publiés ou non, émanant des
- The proportionality between the small variations of total S-electron density and spin density at the site of nucleus location permits proposing a method of