Biochemical and Biophysical Investigations of N-linked Glycosylation Pathways in Archaea
by
Michelle M. Chang B.S. Chemical Biology
University of California, Berkeley, 2008
MASSACHUSETTS INSTITUTE OF TECHNOLOLGY
MAY 192015
LIBRARIES
Submitted to the Department of Chemistryin Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy
at the
Massachusetts Institute of Technology
December 2014 [Febrtary 2c1 2014 Massachusetts Institute of Technology
All rights reserved
Signature of Author: Certified by:
Signature redacted
Department of Chemistry December 11, 2014 __Signatureredacted
Barbara Imperiali Class of 1922 Professor of Chemistry and Biology Thesis SupervisorAccepted by:
Signature redacted_____
Robert W. Field Haslam and Dewey Professor of Chemistry Chairman, Committee on Graduate Students
This doctoral thesis has been examined by a committee of the Department of Chemistry as follows:
Signature redacted
Alice Y. Ting Ellen Swallow Richards Associate Professor of Chemistry Thesis Chair
Signature redacted
Barbara Imperiali Class of 1922 Professor of Chemistry and Biology Thesis Supervisor
Signature redacted
Alexander M. Klibanov Novartis Professor of Chemistry and Bioengineering
Biochemical and Biophysical Investigations of N-linked Glycosylation Pathways in Archaea
by
Michelle M. Chang
Submitted to the Department of Chemistry on December 11, 2014 in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy
Abstract
Asparagine-linked glycosylation is an abundant and complex protein modification conserved among all three domains of life. Much is known about N-glycan assembly in eukaryotes and selected bacteria, in which the oligosaccharyltransferase (OTase) carries out the
en bloc transfer of glycans from polyprenyl-PP-linked donors onto asparagine side chains of
acceptor proteins.
The first aim of this thesis is to elucidate the biochemical details of archaeal N-linked glycosylation, specifically through in vitro analysis of the polyprenyl-P-dependent pathway of the methanogenic archaeon Methanococcus voltae. The archaeal OTase, known as AglB, utilizes a-linked dolichyl-P-trisaccharide substrate as the glycosyl donor for transfer to the acceptor protein. This dolichyl-P-glycan is generated by an initial retaining glycosyltransferase (AglK) and elaborated by additional glycosyltransferases (AglC and AgIA) to afford Dol-P-GlcNAc-Glc-2,3-diNAcA-ManNAc(6Thr)A. Despite the homology to other bacterial or eukaryotic OTases that exploit polyprenyl-PP-linked substrates, the M. voltae AglB efficiently transfers disaccharide to model peptides from the Dol-P-GlcNAc-Glc-2,3-diNAcA monophosphate. While this archaeal pathway affords the same asparagine-linked
P-glycosyl
amide products generated in bacteria and eukaryotes, these studies provide the first biochemical evidence revealing that despite the apparent similarities of the overall pathways, there are actually two general strategies to achieve N-linked glycoproteins across the domains of life.A second focus of this thesis involves biophysical studies to probe structural features and conformational dynamics of AglB. An intramolecular LRET experimental system was developed to report on substrate binding and the resulting structural transformations in AgIB. There is a strong need for detailed studies on the mechanistic and functional significance of archaeal adaptations of N-linked glycosylation, especially exploring differences between AglB and other OTases that allow AglB to utilize these unique polyprenyl-P-linked substrates.
Lastly, a cell-free expression system was established for the efficient synthesis of Alg5, a yeast dolichyl-phosphate glucosyltransferase that shares high sequence similarity to AglK, the first glycosyltransferase in the M. voltae pathway. Dol-P-Glc was generated and examined to unambiguously characterize the stereochemistry of the product of Alg5.
Thesis Supervisor: Barbara Imperiali
Acknowledgements
I would like to start by thanking my advisor, Barbara Imperiali, for giving me the incredible opportunity to be a part of her lab and for her guidance, patience, and support throughout my graduate studies. Thank you for assembling a wonderful group of people who are always willing to collaborate and help one another become better scientists. Thank you also to my thesis committee members, Professors Alice Ting and Alex Klibanov, for their help and advice through the years.
I have had the pleasure of working with a fantastic group of labmates over the past six years. I am thankful for everyone who welcomed Mike and me into the lab in our first year. Even though I didn't overlap very long with them, Galen, Elvedin, Mark, and Nelson all helped make the lab a wonderful place in my first year. Angelyn, Meredith, Wendy, and Brenda were the four fourth-year students who helped me in countless ways as a new graduate student. I am especially thankful to Angelyn for taking the time to mentor me and teach me about the world of membrane protein preparations, OTase assays, HPLC maintenance, and so much more. I am also grateful to Meredith for her generous and sage advice and for teaching me about nanodiscs and the joy of volleyball. I am thankful for Wendy's help with my fluorescent peptide experiments. I worked on PglB with Marcie, and I am so thankful that I had her to discuss science with and for all of her help with LBTs. I appreciate being able to walk through grad school with Mike as a classmate and labmate, and I love that he hosted an annual Thanksgiving meal for our classmates. I am happy to have shared most of my grad school duration with Vinita, who has been a great friend and someone to commiserate with in purifying our sugars and lipid-linked oligosaccharides. I am thankful to Stephanie for helping me with yeast expression and for deciphering my horrible drawings. I am delighted that Sonya joined our lab and enjoyed our time working on cell-free translation. Austin always spoke with great enthusiasm for science and never failed to ask about my day, and I am sorry that his time in our lives was cut short.
I also need to acknowledge the many postdocs throughout my time, starting with Matthieu, Jay, and Cliff who were always generous with their advice. I am particularly thankful to Matthieu for teaching me about peptide synthesis. I would also like to thank postdocs Andrew, Elke, Laura, Marthe, Julie, Joris, Debasis, Silvano, and Cristina who have been brilliant scientists and great resources throughout the years. I am glad to see Monika continuing the LBT-PglB project, and I am excited to see how it compares to AglB. I am thankful for Garrett's help with Dol-P synthesis and NMR and for being my hockey and early lunch buddy. Joris has been my late lunch partner. James, Philipp, and Carsten have each been my podmate at some point, and I appreciate all of their vastly different personalities and value our various conversations about science and life.
I would like to thank members of the Stubbe lab for allowing me to use their French press, as well as members of the Drennan lab for helping me with crystallography. I would also like to thank Traci, Daniel, Chiara, Seymour, and Kaspar for their time working in our lab. I have also enjoyed the time Will, Thais, Pi, Hui, and Natalie have spent in our lab as undergrads. And I would be remiss if I did not thank Debby for her work in the BIF and Elizabeth for all her work to keep the lab running.
Lastly, I would never have made it this far if it weren't for the love and support of my family and friends. Thanks to my mom and dad who are always proud of me and concerned for my academic and personal well-being without ever pressuring me with their expectations. Thanks to my amazing sister, Liana, who is always there for me whenever I need to talk. Thanks
to the MIT Graduate Christian Fellowship, Hope Fellowship Church, and Tang small group communities for fortifying my faith and for all the adventures we've had together. And thank you to my current and former roommates (Stephanie, Heather, Wei-Shan, Nan, Ange) who have made Tang a special place filled with laughter, shared experiences of failures and triumphs, rousing conversations about God and relationships, and jigsaw puzzles.
Table of Contents
Abstract ... 3
Acknow ledgem ents ... 4
Table of Conttents ... 6 List of Figures ... 9 List of Tables... 12 List of A bbreviations... 13 Chapter 1: Introduction...15 Introduction ... 16
Eukaryotic N -linked glycosylation... 17
Bacterial N -linked glycosylation... 19
A rchaeal N -linked glycosylation... 20
Thesis objectives ... 28
References ... 29
Chapter 2: Identification and characterization of archaeal glycosyltransferases...33
Introduction ... 34
Results and D iscussion... 41
AglH is not the first enzyme in the M. voltae N-linked pathway... 41
A glK is a specific D ol-P-GIcN Ac synthase ... 42
D ol-P-GIcN A c exhibits an a-glycosidic linkage ... 47
A giK m utagenesis and inhibition studies... 50
Com parison of A glK with other glycosyltransferases ... 51
Observation of Dol-P-glycan in M . maripaludis cells ... 53
A gIC is a U D P-Glc-2,3-diN A cA glycosyltransferase... 56
Identification of D ol-P-glycans in other archaeal species... 60
Conclusions ... 62
A cknow ledgem ents ... 63
Experim ental M ethods ... 64
Cloning of A glH , A glK and AgiC ... 64
Protein expression and purification... 64
Synthesis of (C55-60) (S)-dolichols... 65
Synthesis of (S)-dolichyl-phosphate ... 66
Synthesis of (C55-60) Dolichyl-PP-GlcNAc and Dolichyl-PP-GaINAc ... 67
Synthesis of U D P-[3H ]G lc-2,3-diN AcA ... 67
A glK and A glC glycosyltransferase assays ... 67
A glK reaction product identification... 69
A glK D X D m utagenesis... 69
N M R and M S characterization of Dol-P-glycans ... 70
M ethanococcus maripaludis LLO preparation ... 70
Pyrococcusfuriosus LLO preparation ... 71
References ... 72
Chapter 3: Characterization of an archaeal oligosaccharyl transferase ... 76
Introduction ... 77
Results and discussion...82
Characterization of oligosaccharyl transferase activity ... 82
AgIB and its lipid-linked glycosyl donor substrate... 87
Investigation of the role of AgIB His-597... 89
AgLB crystallographic studies ... 92
Conclusions ... 97
Acknowledgem ents ... 98
Experim ental m ethods... 99
AglB expression and purification... 99
Oligosaccharyl transferase assay... 100
Purification of glycopeptides ... 101
AglB H597 m utagenesis... 101
Purification of AglB for crystallography ... 101
AglB peptide substrates... 102
AgIB crystallography ... 103
References ... 104
Chapter 4: Efforts towards establishing an experimental system for investigating AgiB conform ational dynam ics by LRET ... 108
Introduction ... 109
Results and discussion... 115
Generation of LBT-AgIB construct...115
Generation of LBT-AgIB cysteine mutants ... 117
LRET with fluorophore-labeled LBT-AgIB ... 121
Lum inescence studies with LBT-AgIB and peptide substrate ... 124
N ew peptide library for LRET experim ents... 127
Luminescence studies with LBT-AgIB and short acceptor peptides ... 131
Conclusions ... 132
A cknowledgem ents...133
Experim ental m ethods... 133
LBT-AglB construct cloning, expression, and purification ... 133
LBT-AglB and LBT-Ub lum inescence experim ents ... 135
LBT-AgIB Cys m utagenesis ... 135
LBT-AgB fluorophore labeling...136
AglB peptide substrate synthesis... 137
Km determ ination of AglB peptide substrates ... 138
References ... 139
Chapter 5: Characterization of Alg5 activity and product stereochemistry ... 141
Results and discussion... 145
A lg5 expression in E. coli ... 145
Alg5 expression in S. cerevisiae ... 150
Cell-free translation of Alg5 ... 151
Conclusions ... 156
Acknowledgem ents ... 157
Experim ental m ethods... 157
A lg5 expression in pGEX-4T3, pET-47, and pE-SUM O vectors... 157
Alg5 expression in pBAD vector ... 158
A lg5 expression from pET(Alg5) plasm id... 160
Alg5 expression in S. cerevisiae ... 160
Alg5 activity assay ... 162
Cell-free expression of Alg5 ... 162
Dol-P-Glc synthesis and analysis... 164
List of Figures Chapter 1
Figure 1-1: Schematic overview of the five different types of protein glycosylation. ... 17
Figure 1-2: N-linked glycosylation pathway in S. cerevisiae. ... 18
Figure 1-3: N-linked glycosylation pathway in C. jejuni...20
Figure 1-4: N-glycan structures from Halobacterium salinarum, Thermoplasma acidophilum and Pyrococcusfuriosus... 22
Figure 1-5: N-linked pathway in H. volcanii in high and low salinity. ... 23
Figure 1-6: N-linked glycosylation pathway in M. voltae. ... 25
Figure 1-7: N-linked glycosylation pathway in M. maripaludis...26
Figure 1-8: N-glycosylation pathway in S. acidocaldarius. ... 27
Chapter 2 Figure 2-1: Representative structures of the two main GT folds...35
Figure 2-2: Catalytic m echanism s in GTs... 37
Figure 2-3: N-linked glycosylation in Methanococcus voltae. ... 39
Figure 2-4: General assay for PGT and GT activity.. ... 41
Figure 2-5: Topology prediction for M. voltae proteins AgIK and AglC ... 43
Figure 2-6: SDS-PAGE and Western blot analysis of purified AglK and AglC proteins ... 43
Figure 2-7: A glK activity assay. ... 45
Figure 2-8: Normal phase HPLC purification of Dol-P-GlcNAc. ... 45
Figure 2-9: ESI-M S of purified Dol-P-GlcNAc... 46
Figure 2-10: Capillary electrophoresis of AglK reaction products... 46
Figure 2-11: Alignment of AglK from M. voltae with S. cerevisiae Alg5 ... 47
Figure 2-12: 1H-NMR spectrum of Dol-P-GlcNAc... 48
Figure 2-13: Activity assay for AgIK DI05A and D107A mutants...50
Figure 2-14: AgiK inhibition observed with UDP-(5F)-GlcNAc...51
Figure 2-15: Alignment of AglK from M. voltae with AgIJ from H. volcanii or MMP 1170 from M m arip alud is. ... 53
Figure 2-16: N-linked glycan structures from M voltae and M. maripaludis ... 54
Figure 2-17: Mass spectrometry analysis of M maripaludis total lipid extraction ... 55
Figure 2-18: Biosynthetic pathway of UDP-ManNAc(3NAc)A ... 57
Figure 2-19: AglC donor and acceptor substrate specificity...58
Figure 2-20: Normal phase HPLC purification of Dol-P-Glc-2,3-diNAcA ... 58
Figure 2-21: ESI-MS of Dol-P-GIcNAc-Glc-2,3-diNAcA ... 59
Figure 2-22: Reverse phase LC/MS analysis of P. furiosus whole cell extractions confirms the presence of Dol-P-heptasaccharide and other Dol-P-glycan intermediates...61
Figure 2-23: M voltae N-linked glycosylation pathway enzymes ... 63
Chapter 3 Figure 3-1: N-linked glycosylation pathway in Saccharomyces cerevisiae. ... 77
Figure 3-3: N-linked glycosylation pathway in Campylobacterjejuni...80
Figure 3-4: Sequence alignment of 16 archaeal Stt3 homologs, depicting the region surrounding the critical W W D X GX m otif... 82
Figure 3-5: Comparison of the archaeal OTases expression by Western blot...82
Figure 3-6: Topology prediction for M voltae AgiB... 83
Figure 3-7: Purified AglB analyzed by SDS-PAGE and Western blots ... 83
Figure 3-8: Standard assay for OTase activity... 84
Figure 3-9: AglB activity assay using the peptide Ac(YKYNESSYKpNF)NH 2 as the acceptor substrate and Dol-P-GlcNAc or Dol-P-disaccharide as the donor substrates ... 85
Figure 3-10: Dependence of AglB activity on divalent metal cations ... 86
Figure 3-11: Reverse-phase HPLC purification of peptide and glycopeptide product...86
Figure 3-12: ESI-MS of the glycopeptide produced by AgIB ... 87
Figure 3-13: Sequence alignment of the WWDXGX motif of representative OTases...90
Figure 3-14: Activity of AglB His-597 mutants with Dol-P-GIcNAc-[3H]Glc-2,3-diNAcA and Ac(YKYNESSYKpNF)NH 2 as the donor and acceptor substrates. ... 91
Figure 3-15: AglB WT, H597A, H597F, H597N, H597Y expression in membrane fractions .. 91
Figure 3-16: AglB wild type and AgIB H597Y mutant activity with Dol-P-GlcNAc-Glc-2,3-diNAcA and Und-PP-Bac-GaINAc... 92
Figure 3-17: Gel filtration chromatography of AgIB with Triton X-100 and LMNG...94
Figure 3-18: Structure of LM NG amphiphile. ... 95
Figure 3-19: AglB acceptor peptide library screen ... 96
Figure 3-20: Crystallization of AgIB in the presence of peptide ... 97
Figure 3-21: M voltae AglB catalyzes the oligosaccharyl transfer from a Dol-P-glycan onto the Asn residue within the NXS/T sequon of acceptor proteins. ... 98
Chapter 4 Figure 4-1: N-linked glycosylation across the three domains of life ... 110
Figure 4-2: Jablonski diagram of a ligand-lanthanide complex ... 111
Figure 4-3: Lanthanide binding tag... 112
Figure 4-4: Intram olecular LRET of AglB ... 114
Figure 4-5: LBT-AglB purification and luminescence of LBT-AglB upon Tb3+ titration ... 116
Figure 4-6: Membrane topology prediction plot for AgIB, illustration of Cys to Ser mutants, and structure of BODIPY-TM R m aleim ide... 118
Figure 4-7: LBT-AglB-C35S/C590S purification and OTase assay... 119
Figure 4-8: LBT-AglB-C35S/C590S/P356C and LBT-AgIB-C35S/C590S/T367C purification and labeling with BODIPY-TM R fluorophore. ... 120
Figure 4-9: Emission spectrum of LBT-AglB-T367C-BODIPY-TMR with or without gating 121 Figure 4-10: Time-gated emission spectra of BODIPY-TMR labeled P356C upon Tb3+ titration. and luminescence decay measured at 543 nm ... 123
Figure 4-11: Time-gated emission spectra of BODIPY-TMR labeled T367C upon Tb3+ titration and lum inescence decay measured at 543 nm ... 124
Figure 4-12: Emission spectra and luminescence decay of LBT-AglB-T367C-BODIPY-TMR and Tb3+ ion exhibit quenching upon addition of Ac(YKYNFTSYKRR)NH 2 peptide...125
Figure 4-13: Loss of luminescence signal with addition of peptides Ac(YFNFTGRR)NH2 and Ac(YKYNFTSYKRR)NH 2to LBT-Ub... 127
Figure 4-14: AgiB acceptor peptide library screen with peptides having N-terminal acetylation and or free N -term inal am ines... 130 Figure 4-15: Emission spectra of LBT-Ub with Tb3 and addition of peptides Ac(TFNFTS)NH
2
and Bz(TFNETS)NH2.... ... . . . ... 132
Chapter 5
Figure 5-1: Dolichol pathway of N-linked glycosylation in Saccharomyces cerevisiae...143 Figure 5-2: Alignment of AgiK from M. voltae with S. cerevisiae Alg5 ... 144 Figure 5-3: Topology prediction for S. cerevisiae Alg5. ... 145 Figure 5-4: GST-Alg5 and His6-Alg5 expression detected by SDS-PAGE and Western blot.
Activity assay of GST-Alg5, His6-Alg5 constructs ... 146
Figure 5-5: Expression of His6-SUMO-Alg5 in C41 (DE3), C43 (DE3), BL21 (DE3) RIL,
Rosetta2 (D E3), and Lem o2l (DE3) cell lines ... 146 Figure 5-6: Alg5-Hisio expression levels detected at various times after induction with variable
concentrations of L-arabinose ... 147 Figure 5-7: Activity of cell lysates Alg5-Hisio expressed from pBAD vector in BL21 (DE3) RIL cells at various times after induction with 0.2% L-arabinose. ... 148 Figure 5-8: TRX-His6-Alg5 expression is detected by Western blot analysis...148
Figure 5-9: Alg5 expression as CEF from pET(Alg5) under different conditions. Activity assay of A lg5 C E F s... 149 Figure 5-10: HPLC fractions of the organic extraction of the Alg5 reaction products... 150 Figure 5-11: Alg5 yeast expression in yeast strains INVSc1, W303a, or W303c...151 Figure 5-12: Glucosyltransferase activity assay with membrane fractions of Alg5 expression in
IN V Sc1, W 303a, and W 303c ... 151
Figure 5-13: Western blot showing cell-free expression of Alg5-NT and Alg5-CT ... 153 Figure 5-14: Western blot of Alg5-NT and AglC expression with additives, or Alg5-NT and
A lg5-CT solubilization after expression. ... 154 Figure 5-15: Glucosyltransferase activity assay with Alg5-NT and Alg5-CT ... 155 Figure 5-16: 1H-NMR and 3'P decoupled 1H-NMR spectrum of Dol-P-Glc... 156
List of Tables Chapter 1
Table 1-1: Features of archaeal N-linked glycosylation in select archaeal species. ... 28 Chapter 2
Table 2-1: 1H- and 13C-NMR assignments of Dol-P-GlcNAc structure ... 49
Chapter 4
Table 4-1: Amino acid preference at each position of XO-X1-X2-X3-X4-X5 peptide. ... 131 Table 4-2: Apparent Km values of best AglB peptide substrates. ... 131
List of Abbreviations
Standard 3-letter and 1-letter codes are used for the 20 natural amino acids. Standard 1-letter codes are used for the 4 common DNA bases.
AcCoA ATP B-OG BODIPY CAM CE CEF CHAPSO DDM DIPEA DMF DMSO Dol-P Dol-PP DMPC DTT EDTA ER ESI-MS Fmoc Fos-12 FRET GDP-Man GST GT IPTG HEPES Km Kd LB LBT LDAO LLO LMNG LRET NDP Ni-NTA NMR NP-HPLC OTase acetyl-coenzyme A adenosine 5'-triphosphate n-octyl
P-D-glucoside
4,4-difluoro-4-bora-3a,4a-diaza-s-indacene ceric ammonium molybdatecapillary electrophoresis cell envelope fraction
3-[(3-cholamidopropyl)dimethylammonio]-1 -propanesulfonate n-dodecyl--D-maltoside diisopropylethylamine dimethylformamide dimethylsulfoxide dolichyl-phosphate dolichyl-diphosphate 1,2-dimyristoyl-sn-glycero-3-phosphocholine dithiothreitol
extinction coefficient for molar absorptivity ethylenediaminetetraacetic acid
endoplasmic reticulum
electrospray ionization-mass spectrometry 9-fluorenylmethoxycarbonyl
n-dodecyl phsophocholine
fluorescence resonance energy transfer guanosine diphosphate mannose glutathione S-transferase
glycosyltransferase
isopropyl f-D-1-thiogalactopyranoside
4-(2-hydroxyethyl)- 1 -piperazineethanesulfonic acid Michaelis constant
dissociation constant Luria-Bertani broth
lanthanide binding tag lauryl dimethylamine oxide lipid-linked oligosaccharide lauryl maltose neopentyl glycol
luminescence resonance energy transfer nucleotide diphosphate
nickel nitrilotriacetic acid nuclear magnetic resonance
normal phase high performance liquid chromatography oligosaccharyltransferase
PAL PDB PEG-PS PGT pNF POPC Pren-P Pren-PP PSUP PyBOP Ro RP-HPLC SDS-PAGE T-TFA TIS TLC TMHMM Triton X-100 TRX TUPS UDP UDP-Bac UDP-Glc UDP-GlcNAc UDP-Gal UDP-GalNAc UDP-Glc-2,3-diNAcA UMP Und-P Und-PP Xaa 5-(4'-aminomethyl-3',5'-dimethoxyphenoxy)valeric acid Protein Data Bank
polyethylene glycol-grafted polystyrene phosphoglycosyltransferase
4-nitrophenylalanine
1 -palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine polyprenyl-phosphate
polyprenyl-diphosphate pure solvent upper phase
benzotriazol- 1 -yl-oxytripyrrolidinophosphonium hexafluorophosphate
FOrster distance
reverse phase high performance liquid chromatography sodium dodecyl sulfate polyacrylamide gel electrophoresis lifetime
trifluoroacetic acid triisopropylsilane
thin layer chromatography
Tied Mixture Hidden Markov Model for transmembrane prediction t-octylphenoxypolyethoxyethanol
thioredoxin
theoretical upper phase with salt uridine 5'-diphosphate UDP-2,4-diacetamido-2,4,5-trideoxy-ax-D-glucose UDP-glucosamine UDP-N-acetyl-D-glucosamine UDP-galactosamine UDP-N-acetyl- D-galactosamine UDP-2,3-diacetamido-2,3-dideoxy-D-glucuronic acid uridine 5'-monophosphate undecaprenyl-phosphate undecaprenyl-diphosphate
Introduction
Post-translational modifications of proteins help to expand the proteome far beyond what is directly encoded by the genome.' Glycosylation is one of the predominant and most complex protein modifications found in nature, occurring in all three domains of life - Eukarya, Bacteria, and Archaea.2 The enormous diversity of sugars, sugar linkages, sugar-protein linkages, and glycan size leads to a seemingly endless number of possible protein-linked glycan structures. Glycosylation can profoundly impact protein function, playing important roles in biological processes such as protein folding and stability, intracellular targeting, cellular signaling and adhesion, and the immune response.3
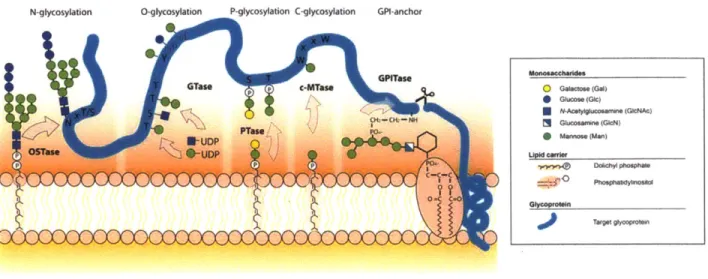
Five major types of protein glycosylation have been observed in nature and are classified based on the manner in which the glycan is linked to the modified protein (Figure 1-1). N-linked glycosylation involves the en bloc transfer of an oligosaccharide from a lipid carrier onto the side chain amide nitrogen of asparagine residues within a conserved sequon of acceptor proteins.4-6 O-linked glycosylation involves the transfer of monosaccharide units onto the side chain hydroxyl oxygen atom of either serine or threonine residues, and to a lesser extent, other residues such as tyrosine, hydroxylysine and hydroxyproline. Glycosylphosphatidylinositol (GPI) anchoring involves the transfer of a GPI moiety to the C-terminus of a target protein, tethering it to the cell membrane.' Phosphoglycosylation involves the addition of sugar I -phosphates to the hydroxyl oxygen atom of serine or threonine residues.9 C-mannosylation involves the transfer of mannose to the indole C2' carbon atom of tryptophan residues resulting in a C-C linkage.10'11 N-linked, 0-N-linked, and GPI-anchored glycosylation have been well-studied biochemically, while
0-glycosylation P-glycosylation C-glycosylation GPM-anchor
zon W
Figure 1-1: Schematic overview of the five different types of protein glycosylation. Taken from Jarrell, et al.'
Eukaryotic N-linked glycosylation
N-linked glycosylation is an essential part of the protein maturation process in eukaryotes and is estimated to occur in more than half of all eukaryotic proteins.'2 Genetic and biochemical characterization of this process has provided the best understanding of this pathway in
Saccharomyces cerevisiae (Figure 1-2A), but N-glycosylation is remarkably conserved in all
eukaryotes, from yeast to man.'3 The first phase of N-glycosylation occurs on the cytoplasmic face of the endoplasmic reticulum (ER) membrane. The assembly of the lipid-linked oligosaccharide (LLO) is initiated by a phosphoglycosyltransferase, followed by a series of membrane-bound glycosyltransferases in the asparagine-linked glycosylation (Alg) family that catalyze the transfer of each monosaccharide unit from nucleotide sugar donors onto a dolichyl-diphosphate (Dol-PP) carrier. Dolichol is a long chain, a-saturated linear polyisoprene that typically varies in size between 14 and 21 units depending on the cell type and species.'4 After the completion of the branched heptasaccharide, the Dol-PP-GlcNAc2Man5 intermediate is
flipped across the membrane to face the ER lumen in an ATP-independent manner. Yeast genetic Man-4cwwif o Galrncto. (Gd * kn(I * N'A*My~uCOGsamN(G4A SGuWsmri (GICN) *Muano (an Upid asaw S DO Gooh ftycopm-.i Torg~t gtyoop-ew
analysis previously implicated Rftl as the purported flippase, but recent studies show that Rftl-null cells do not stall at the flippase step, and Rftl may instead act as a chaprone.15"1 6 The flipped LLO is further elaborated by the transfer of mannose and glucose residues from Dol-P-mannose and Dol-P-donors, respectively. The fully assembled branched tetradecassachride, Glc3Man9GlcNAc2, is transferred by the oligosaccharyltransferase (OT) onto asparagine residues
within the N-X-S/T sequon, where X can be any amino acid except proline (Figure 1-2B).
A
CTP UDP1 Sec59 Alg7 Cytoplasm ER Lumen nascent gycoprotein OT ILAiglO UDP-U Alg13I14 GDP-U A g2I
'3 a.j
GDP-U A g2I A,
IrA
I
A1g98 a Ag6 a A1IgQ CLa A1g12
T 4 . 1. -4a
i
L
0-
Ci
IIcLc
rl" N-Ma Glu d_ DolB
HO OH HO OH HO 0 OH HO HO - 0O HO OH OH 0 LHO HO OH ix;;4.l0o OH HO 0 OH 0 'A '0 HO OH HO. OH -Z~o0 0 H% OH HO H OH H0 AcHN AcHN GDP- Y GDP-0UYA1 A
Rfti? IAlg5 O-P 8 GDP-U Dpm1 a. a. " 4- a. a,,d acetylglucosamine (GIcNAc) nnose (Man) cose (Glc) 0. 0. o-P-O-P-o-ichyl-diphosphate =-
- 1~ 2Figure 1-2: (A) N-linked glycosylation pathway in S. cerevisiae. (B) Structure of the eukaryotic N-linked glycan, Glc3Man9GlcNAc2.
1--GDP-11 A191
Newly formed glycoproteins are processed by glycosidases and glycosyltransferases in the ER and Golgi apparatus before being secreted or distributed to various cellular locations. This downstream modification results in the vast diversity of N-linked glycans observed in eukaryotes." Although the Glc3Man9GlcNAc2 tetradecasaccharide core is widely conserved in
eukaryotes, several species of protists have been found to assemble only truncated forms of the glycan.18 N-glycans play an important role in the quality control of protein folding in the ER, serve as identity and localization tags to direct proteins to the proper cellular destination, help regulate the mobility of cell membrane proteins through interactions with lattice-forming lectins, impart structural rigidity, and may provide protection against proteolysis.19
Bacterial N-linked glycosylation
The first bacterial N-linked glycosylation system was discovered in Campylobacter
jejuni, a gram-negative human gut mucosal pathogen.2 0'2 1 Glycan assembly occurs on the cytoplasmic face of the periplasmic membrane by the protein glycosylation (Pgl) enzymes (Figure 1-3A). This pathway resembles the first half of the dolichol pathway in S. cerevisiae, but with several differences. A phosphoglycosyltransferase initiates the LLO assembly and then a series of glycosyltransferases add monosaccharide units from nucleotide sugar donors onto an undecaprenyl-diphosphate (Und-PP) carrier rather than onto Dol-PP. Undecaprenol is a long chain, unsaturated polyisoprene comprising 11 isoprene units.2 2 Instead of the GlcNAc residue found in eukaryotic N-glycans, the first sugar in the C. jejuni glycan is N,N'-diacetylbacillosamine (Bac). After the completion of the heptasaccharide, the Und-PP-BacGalNAc5Glc intermediate is flipped across the membrane to the periplasmic face by an
(PglB) onto asparagine residues within the extended sequon, D/E-X1-N-X2-S/T, in which an
acidic residue occupies the -2 position (Figure 1-3B).
A
PgIF PgIE
UDP-U - UDP-8 UDP-O Cytoplasm PgIC P91A
Perplasm
B
OH UDP-O PglJ PgIH CLI
UDP-U Pg i 'UI
]
PgKK PglB HO OH o O MAcH I ACOHO 0 21 HO~ AHNO4 CH3 H H ACHN' 0 A~ HO 0 N-acetylglucosamine (GlcNAc) N,N'-diacetylbacillosamine (Bac)o
N-acetylgalactosamine (GalNAc) Glucose (Glc) Undecaprenyl-diphosphate Undecaprenol n-OH n = 7 (length < 50 A)Figure 1-3: (A) N-linked glycosylation pathway in C. jejuni. (B) Structure of the N-linked glycan, GlcGalNAc5Bac.
Over one hundred glycoproteins have been identified in C. jejuni and are associated with a wide range of cellular functions.2 4 Mutations of the pgl locus resulted in impaired host cell
adhesion and colonization, establishing a link between N-glycosylation and pathogenicity.20,25,26 Other important roles for N-linked glycosylation may include modulation of protein-protein interactions, enhancement of protein stability, and protection of C. jejuni from osmolytic stress.27 28
Archaeal N-linked glycosylation
In 1977, recognition of major differences in the structure and genetics of prokaryotes led to the classification of archaea and bacteria into two distinct domains.29Archaea share common
features with both bacteria and eukaryotes while also possessing several unique characteristics. Like bacteria, archaea are single-celled organisms, have circular chromosomes and no nucleus or membrane-bound organelles, but they also possess several genes and metabolic pathways that are more closely related to those of eukaryotes, especially the enzymes involved in transcription and translation. Archaea contain ether-linked phospholipids in their cell membranes instead of the ester-linked phospholipids found in bacteria and eukaryotes. In addition, archaea are able to
30 utilize a wide range of energy sources and frequently inhabit extreme environments.
N-linked glycosylation is predicted to occur in nearly all of the archaeal species with sequenced genomes.3 1 While many archaeal N-glycans have been identified in S-layer,
archaellin, and other cell-surface glycoproteins, not many N-glycan structures have been fully characterized and even fewer biosynthetic pathways understood.. The structures that have been characterized over the years demonstrate the incredible variety of archaeal N-glycans (Figure 1-4).3-3 There is a high diversity of archaeal N-linked glycan structures in terms of size, the degree of branching, the identity of the linking sugar, the presence of unique sugars, and further modification of sugar moieties by methylation, sulfation, and even addition of pendant amino acids. Such diversity is far beyond that found in bacteria and eukaryotes. However, the chemical structure of the glycan and the nature of the glycan linkages to protein have often remained undetermined and the pathways for N-glycan assembly unclear. The recent availability of complete genome sequences for nearly 200 archaeal species and the development of appropriate molecular tools for genetic manipulation have facilitated the detailed delineation of several of these pathways.31
,36 Since the first discovery of archaeal N-linked glycoproteins in
limited to the euryarchaeal Haloferax volcanii, Methanococcus voltae, and Methanococcus
maripaludis, and the crenarchaeal Sulfolobus acidocaldarius.
A B HO 0 0>(. 0 0 H H1CO R'H $O3 RAO R- RO H R30 30 R30 330 RO ~ .~ R=H, SOO A2N -o IN0 1 H 0 AcN HO
/10-13
C D S O 0" OH HHutOH eI N ~ HO~HO . 0 00 HO HO YHOOH HIH 6ic OH 040
TOHOH OH OilH~
high-mannose-type linked glycan of Thermoplasma acidophilum. (D) Structure of the N-linked heptasaccharide in Pyrococcusfuriosus.
There are two pathways of N-glycosylation in the extreme halophile Haloferax volcan ii. Under normal high salt conditions, the process begins with the sequential addition of the first four sugar subunits of the pentasaccharide from nucleotide-activated donors to a Dol-P carrier at the cytoplasmic leaflet of the cytoplasmic membrane (Figure 1 -5A).38-44 In contrast to the
eukaryotic (C70-Cl o) Dol-PP lipid carriers, (C5 5-C6o) Dol-P serves this role in H. volcanii. The
Dol-P-tetrasaccharide intermediate and a separate Dol-P-mannose are flipped across the membrane, and the tetrasaccharide is transferred by the oligosaccharyltransferase (AgiB) to the asparagine residue within the N-X-S/T sequon of the target S-layer glycoprotein or archaellins. The final mannose subunit is then delivered from its flipped Dol-P carrier onto the glycoprotein-linked tetrasaccharide, resulting in the final glycoprotein N-glycoprotein-linked to the pentasaccharide. At higher salinities, both Dol-P charged with the first four pentasaccharide sugars and Dol-P charged with final mannose sugar have been observed by mass spectrometry analysis.
A NDP-U
AgIM NDPU NDP-U NDP-U AgIF
AgIJ q AgIG AgIl
Cytoplasm a. - a.- C. ON C Exteror Dolichol OH - 7-8 2 NDP-U AgIE -+ I SAM AgIP -- + o-Agli2 Ag Il AIj~iAg l2 AgI5 P AgI8 Ag l3
Ag6 qtAplasmss Ag7Ag19 i Agli4 !soi
Cytoplasm a. - a.- e L . -. C.. - a - a. Exteror Ag ? AI HH, 115 0 , NDP-u AgID
!
AgIR AgIS H3 0 Hexose (Hex)N Hexuronic acid (HexA) * Mannose (Man)
O Rhamnose (Rha)
I
Dolichyl-phosphate (Dol-P)Figure 1-5: (A) linked pathway in H. volcanii cells grown in 3.4 M NaCl. (B) A second N-linked pathway in H. volcanii cells grown in 1.75 M NaCl. The exact structures of the N-N-linked glycans on S-layer glycoproteins or archaellins have not yet been determined.
When H. volcanii cells are grown at lower salinity, S-layer glycoproteins display a distinct tetrasaccharide, comprising a sulfated hexose, two unmodified hexoses, and a rhamnose. Dol-P charged with this tetrasaccharide has also been observed by mass spectrometry.43 The current working model in this low-salt environment implicates an entirely different set of enzymes for glycan assembly (Figure 1-5B). A currently unidentified oligosaccharyltransferase, not AglB, is believed to be responsible for the transfer of this glycan to a different asparagine residue on the S-layer glycoprotein. This second pathway appears to be an adaptive response and is recruited when the first pathway is compromised.45
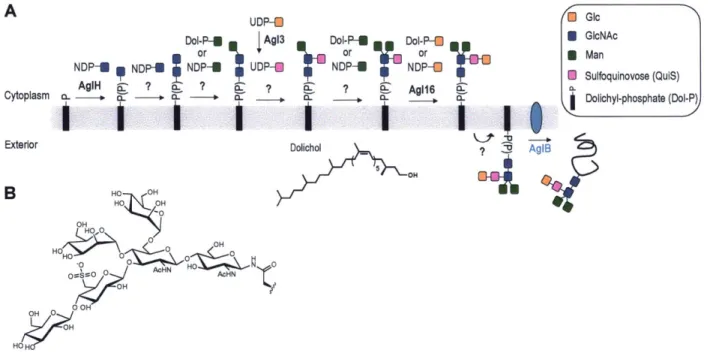
Chapters 2 and 3 of this thesis detail the biochemical characterization of the N-linked glycosylation pathway of Methanococcus voltae, a marine methanogenic mesophile. This pathway is initiated at the cytoplasmic face of the cell membrane by a series of glycosyltransferases that add monosaccharide units from nucleotide-activated sugar donors onto
a Dol-P carrier (Figure 1-6A).46 49 The Dol-P-trisaccharide is then flipped across the membrane
to the exterior surface of the cell, where AglB transfers the trisaccharide onto the asparagine residue within the N-X-S/T sequon of target proteins. In our studies, AgIB efficiently transfers a disaccharide from a Dol-P carrier to acceptor peptides, but the glycan that is natively N-linked to archaellins and S-layer proteins is a trisaccharide, ManNAc(6Thr)A-Glc-2,3-diNAcA-GIcNAc (Figure 1-6B).
A UDP -U UDP-U UDP--U GlcNAc
Cytoplasm cL A1K AgIC Ag1A Glc-2,3-diNAA
[LC
ManNAc(6Thr)A Exterior[Dolichyl-phosphate
AgIB Dolichol B HO 0 OH O~NHAC 0 0 OH RO -O 0 N 0 0 AcH AcHNFigure 1-6: (A) N-linked glycosylation pathway in M voltae (B) Structure of the N-linked
archaellar glycan, ManNAc(6Thr)A-Glc-2,3-diNAcA-GlcNAc.
The N-linked glycosylation pathway in another methanogen, M maripaludis, shares similarities with that of the closely related M voltae (Figure 1-7A).50 Chapter 2 of this thesis
posits that the first glycosyltransfer step of the M maripaludis pathway is initiated by MMP 1170, an enzyme that has yet to be characterized but exhibits extremely high homology to
AglK, the first glycosyltransferase of the M voltae pathway.49 Three other glycosyltransferases
add monosaccharide units from nucleotide-activated sugar donors onto a P carrier. The Dol-P-tetrasaccharide is then flipped from the cytoplasmic face of the cell membrane to the exterior, where AglB transfers the tetrasaccharide onto the asparagine residue within the N-X-S/T sequon of acceptor proteins. Strong evidence for a Dol-P carrier comes from our mass spectrometry analysis of M maripaludis cells, in which we observed the presence of a Dol-P-trisaccharide intermediate bearing the first three sugars of the N-linked glycan identified in M maripaludis archaellar and pilin proteins, Sug-ManNAc(3NAm6Thr)A-Glc-2,3-diNAcA-GaNAc, where the terminal Sug is 2-acetamido-2,4-dideoxy-5-0-methylhexosulo-1,5-pyranose (Figure 1-7B).
A U GaINAc NDP-U NDP-U NDP-U NDP- M Glc-2,3-diNAcA
MMP1170?
!
AglO AgIA AgIL I ManNAc(3NAm6Thr)ACytplsm .. -- c. - . 0- -- + O. - + O. c 2NAc(4deoxy)hexulose Dolichyl-phosphate (Dol-P) Exterior AgIB Dolichol OH B HO 0 OH NHAc 0 --<NHc ~CiO 0 OH OH
OCH3 AcHN AcHN
Figure 1-7: (A) N-linked glycosylation pathway in M maripaludis (B) Structure of the N-linked
archaellar glycan, 2NAc(4deoxy)hexulose-ManNAc(3NAm6Thr)A-Glc-2,3-diNAcA-GaNAc.
Crenarchaeal N-linked glycosylation has been best studied in the thermophilic Sulfolobus
acidocaldarius, but the pathway enzymes are only partially identified (Figure 1-8A).5
2-54 The actions of a phosphoglycosyltransferase and subsequent glycosyltransferases are thought to build up a tribranched glycan on a Dol-PP carrier, which is flipped from the cytoplasmic to the exterior surface of the cell membrane, before transfer of the glycan onto the asparagine residue of target proteins. S-layer, archaellin, and cytochrome b5 5 8/5 6 6 proteins are N-glycosylated with
heterogeneous family of glycans, with the largest comprising GlcNAc2GlcMan2 and an unusual sulfated sugar called sulfoquinovose (Figure 1-8B). The dolichols observed in S. acidocaldarius are unusually short and display a higher degree of saturation of internal isoprene units than eukaryotic or other archaeal dolichols. While genetic complementation data suggest that Dol-PP is the lipid carrier, only Dol-P and Dol-P charged with different hexoses have ever been detected
by mass spectrometric analysis of lipid extracts.5 3 No Dol-P or Dol-PP bearing complex glycans
the M voltae pathway illustrates that genetic complementation studies alone are insufficient in characterizing enzyme activity.47'49 This ambiguity suggests that the S. acidocaldarius pathway could in fact be more like the euryarchaeal pathways, with Dol-P being the lipid carrier instead of Dol-PP. Interestingly, S. acidocaldarius is the only archaeal organism studied thus far in which N-linked glycosylation appears to be essential.54
A UDP-* Gic
Dol-P-U Agl3 Dol-P-U DoI-P-U U G2cNAc
or ,, I or or EM an
NDP-1 qNDP- NDP-U UDP-U NDP-U NDP-U 0 Sulfoquinovose (QuiS)
Cytoplasm 0 A CL (L a. Dolichyl-phosphate (Dol-P)
Exterior Dolichol AgIB
OH B HO OH HO H OH H010 0 OH HHO O OH OH 0O 'jOH HO H
Figure 1-8: (A) Current understanding of the N-glycosylation pathway in S. acidocaldarius. (B) Structure of the tribranched hexasaccharide N-glycan containing the 6-sulfoquinovose.
Progress has been made in recent years to identify highly unusual archaeal N-glycans and the enzymes involved in their biosynthesis and transfer. Much of this progress has come through deletion of genes potentially involved in a pathway and analysis of the downstream effects on protein glycosylation through mass spectrometry. Unfortunately, there is a dearth of biochemical validation of proposed pathways and in vitro enzymatic assays are difficult when the identities of the substrates are often unknown. Even when the glycan identity is known, questions remain about the lipid-linked carrier (Dol-P vs. Dol-PP) and the mechanism of transfer to acceptor
proteins. Archaeal oligosaccharyltransferases (designated as AglBs) are easily identifiable in archaeal genomes, and many archaeal species encode multiple AgiBs. The physiological significance of this observation remains unclear, as multiple AgiBs could reflect differences in substrate or target preference, possibly as a function of local growth conditions. A few AglB crystal structures have been published, but there are still major gaps in our knowledge of the AgiB mechanisms (Table 1-1). All known oligosaccharyltransferases contain a WWDXGX2
motif, and sequence variability in this motif may reflect different activities of the various versions of the protein in the same species.55 It would be highly noteworthy if a link could be established between the type of AgiB and the lipid-linked oligosaccharide substrates it uses.
Table 1-1: Features of archaeal N-linked glycosylation in select archaeal species, highlighting the WWDX1GX2 motif of AglB. Verified oligosaccharyltransferases are marked with (*).
Species AgIB WWDX1GX2 Observed LLO Crystal structure
H. salinarium OE2548F WWDYGH Dol-P-HexHexA"'
Dol-PP-tetrasaccharide56
H. volcanii HVO 1530 WWDYGH Dol-P-tetrasaccharide43 _
M. voltae Mvol_1038* WWDNGH
M. maripaludis MMP1424* WWDNGH Dol-P-trisaccharide4 9
S. acidocaldarius Sacil274 WWDYGY Dol-P-Hex"
P. furiosus PF0156* WWDYGY Dol-P-heptasaccharide PFO 156 C-term"7 PF0411 WWDWGH (Chapter 2)
P. horikoshii PH0242 WWDYGY PH0242 C-term8
PH1271 WWDWGH
A. fulgidus AF0380* WWDYGH AF0380 C-term, 5 ful5
AF0329 WWDYGN AF0329 C-term58
AF0040 WWDYGN AF0040 C-term58
Thesis objectives
The overall goal of this thesis is to gain a deeper understanding of the process of N-linked glycosylation in archaea. Chapter 2 details the identification and characterization of the activities of AglK and AgIC, the first two glycosyltransferases in the M voltae N-glycosylation pathway.
Chapter 3 discusses the biochemical characterization of AgIB activity using purified substrates, which represents the first biochemical demonstration of archaeal oligosaccharyltransferase activity. Chapter 4 describes our progress towards developing a new system to investigate the
conformational dynamics of AglB upon substrate binding using a lanthanide-based luminescence resonance energy transfer technique. Finally, Chapter 5 describes our efforts to express Alg5, a dolichyl-phosphate glucosyltransferase, and the stereochemical analysis of the product.
References
1. Walsh, C. T., Garneau-Tsodikova, S. & Gatto, G. J. Protein Posttranslational Modifications: The Chemistry of Proteome Diversifications. Angew. Chem. Int. Ed. 44, 7342-7372 (2005). 2. Varki, A. et al. Essentials of Glycobiology. (Cold Spring Harbor Laboratory Press, 2009). at
<http://www.ncbi.nlm.nih.gov/books/NBK 1908/>
3. Varki, A. Biological Roles of Oligosaccharides - All of the Theories Are Correct.
Glycobiology 3, 97-130 (1993).
4. Burda, P. & Aebi, M. The dolichol pathway of N-linked glycosylation. Biochim. Biophys.
Acta BBA -Gen. Subj. 1426, 239-257 (1999).
5. Jarrell, K. F. et al. N-Linked Glycosylation in Archaea: a Structural, Functional, and Genetic Analysis. Microbiol. Mol. Biol. Rev. 78, 304-341 (2014).
6. Weerapana, E. & Imperiali, B. Asparagine-linked protein glycosylation: from eukaryotic to prokaryotic systems. Glycobiology 16, 91 R-1 01 (2006).
7. Steen, P. V. den, Rudd, P. M., Dwek, R. A. & Opdenakker, G. Concepts and Principles of O-Linked Glycosylation. Crit. Rev. Biochem. Mol. Biol. 33, 151-208 (1998).
8. Paulick, M. G. & Bertozzi, C. R. The Glycosylphosphatidylinositol Anchor: A Complex Membrane-Anchoring Structure for Proteinst. Biochemistry 47, 6991-7000 (2008).
9. Haynes, P. A. Phosphoglycosylation: A new structural class of glycosylation? Glycobiology 8, 1-5 (1998).
10. Doucey, M.-A., Hess, D., Cacan, R. & Hofsteenge, J. Protein C-Mannosylation Is Enzyme-catalysed and Uses Dolichyl-Phosphate-Mannose as a Precursor. Mol. Biol. Cell 9, 291-300 (1998).
11. Furmanek, A. & Hofsteenge, J. Protein C-mannosylation: facts and questions. Acta Biochim.
Pol. 47, 781-789 (2000).
12. Apweiler, R., Hermjakob, H. & Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta BBA - Gen. Subj. 1473, 4-8 (1999).
13. Lehle, L., Strahl, S. & Tanner, W. Protein Glycosylation, Conserved from Yeast to Man: A Model Organism Helps Elucidate Congenital Human Diseases. Angew. Chem. nt. Ed. 45, 6802-6818 (2006).
14. Jones, M. B., Rosenberg, J. N., Betenbaugh, M. J. & Krag, S. S. Structure and synthesis of polyisoprenoids used in N-glycosylation across the three domains of life. Biochim. Biophys.
Acta BBA - Gen. Subj. 1790, 485-494 (2009).
15. Helenius, J. et al. Translocation of lipid-linked oligosaccharides across the ER membrane
requires Rftl protein. Nature 415, 447-450 (2002).
16. Jelk, J. et al. Glycoprotein Biosynthesis in a Eukaryote Lacking the Membrane Protein Rftl.
J. Biol. Chem. 288, 20616-20623 (2013).
17. Helenius, A. & Aebi, M. Roles of N-Linked Glycans in the Endoplasmic Reticulum. Annu.
Rev. Biochem. 73, 1019-1049 (2004).
18. Guha-Niyogi, A., Sullivan, D. R. & Turco, S. J. Glycoconjugate structures of parasitic protozoa. Glycobiology 11, 45R-59R (2001).
19. Larkin, A. & Imperiali, B. The Expanding Horizons of Asparagine-Linked Glycosylation.
Biochemistry 50, 4411-4426 (2011).
20. Szymanski, C. M., Yao, R., Ewing, C. P., Trust, T. J. & Guerry, P. Evidence for a system of general protein glycosylation in Campylobacter jejuni. Mol. Microbiol. 32, 1022-1030 (1999).
21. Young, N. M. et al. Structure of the N-Linked Glycan Present on Multiple Glycoproteins in the Gram-negative Bacterium, Campylobacter jejuni. J Biol. Chem. 277, 42530-42539 (2002).
22. Hartley, M. D. & Imperiali, B. At the membrane frontier: A prospectus on the remarkable evolutionary conservation of polyprenols and polyprenyl-phosphates. Arch. Biochem.
Biophys. 517, 83-97 (2012).
23. Alaimo, C. et al. Two distinct but interchangeable mechanisms for flipping of lipid-linked oligosaccharides. EMBO J. 25, 967-976 (2006).
24. Scott, N. E. et al. Simultaneous Glycan-Peptide Characterization Using Hydrophilic Interaction Chromatography and Parallel Fragmentation by CID, Higher Energy Collisional Dissociation, and Electron Transfer Dissociation MS Applied to the N-Linked Glycoproteome of Campylobacter jejuni. Mol. Cell. Proteomics 10, M00003 1-MCP201 (2011).
25. Szymanski, C. M., Burr, D. H. & Guerry, P. Campylobacter protein glycosylation affects host cell interactions. Infect Immun 70, 2242-4 (2002).
26. Hendrixson, D. R. & DiRita, V. J. Identification of Campylobacter jejuni genes involved in commensal colonization of the chick gastrointestinal tract. Mol. Microbiol. 52, 471-484 (2004).
27. Larsen, J. C., Szymanski, C. & Guerry, P. N-Linked Protein Glycosylation Is Required for Full Competence in Campylobacterjejuni 81-176. J. Bacteriol. 186, 6508-6514 (2004).
28. Nothaft, H., Liu, X., McNally, D. J., Li, J. & Szymanski, C. M. Study of free oligosaccharides derived from the bacterial N-glycosylation pathway. Proc. Natl. Acad Sci. 106, 15019-15024 (2009).
29. Woese, C. R. & Fox, G. E. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc. Natl. Acad Sci. U. S. A. 74, 5088-5090 (1977).
30. Calo, D., Kaminski, L. & Eichler, J. Protein glycosylation in Archaea: Sweet and extreme.
Glycobiology 20, 1065-1076 (2010).
31. Kaminski, L., Lurie-Weinberger, M. N., Allers, T., Gophna, U. & Eichler, J. Phylogenetic-and genome-derived insight into the evolution of N-glycosylation in Archaea. Mol.
32. Eichler, J. & Adams, M. W. W. Posttranslational Protein Modification in Archaea.
Microbiol. Mol. Biol. Rev. 69, 393-425 (2005).
33. Mescher, M. F., Hansen, U. & Strominger, J. L. Formation of lipid-linked sugar compounds in Halobacterium salinarium. Presumed intermediates in glycoprotein synthesis. J. Biol.
Chem. 251, 7289-7294 (1976).
34. Lechner, J. & Wieland, F. Structure and Biosynthesis of Prokaryotic Glycoproteins. Annu.
Rev. Biochem. 58, 173-194 (1989).
35. Fujinami, D., Matsumoto, M., Noguchi, T., Sonomoto, K. & Kohda, D. Structural
elucidation of an asparagine-linked oligosaccharide from the hyperthermophilic archaeon, Pyrococcus furiosus. Carbohydr. Res. 387, 30-36 (2014).
36. Leigh, J. A., Albers, S.-V., Atomi, H. & Allers, T. Model organisms for genetics in the domain Archaea: methanogens, halophiles, Thermococcales and Sulfolobales. FEMS
Microbiol. Rev. 35, 577-608 (2011).
37. Mescher, M. F. & Strominger, J. L. Purification and characterization of a prokaryotic glucoprotein from the cell envelope of Halobacterium salinarium. J. Biol. Chem. 251, 2005-2014 (1976).
38. Kuntz, C., Sonnenbichler, J., Sonnenbichler, I., Sumper, M. & Zeitler, R. Isolation and characterization of dolichol-linked oligosaccharides from Haloferax volcanii. Glycobiology 7, 897-904 (1997).
39. Abu-Qam, M. & Eichler, J. Protein N-glycosylation in Archaea: defining Haloferax volcanii genes involved in S-layer glycoprotein glycosylation. Mol. Microbiol. 61, 511-525 (2006). 40. Abu-Qam, M. et al. Haloferax volcanii AglB and AgID are Involved in N-glycosylation of
the S-layer Glycoprotein and Proper Assembly of the Surface Layer. J. Mol. Biol. 374, 1224-1236 (2007).
41. Guan, Z., Naparstek, S., Kaminski, L., Konrad, Z. & Eichler, J. Distinct glycan-charged phosphodolichol carriers are required for the assembly of the pentasaccharide N-linked to the Haloferax volcanii S-layer glycoprotein. Mol. Microbiol. 78, 1294-1303 (2010).
42. Kaminski, L. et al. AgliJ Adds the First Sugar of the N-Linked Pentasaccharide Decorating the Haloferax volcanii S-Layer Glycoprotein. J. Bacteriol. 192, 5572-5579 (2010).
43. Calo, D., Guan, Z., Naparstek, S. & Eichler, J. Different routes to the same ending: comparing the N-glycosylation processes of Haloferax volcanii and Haloarcula marismortui, two halophilic archaea from the Dead Sea. Mol. Microbiol. 81, 1166-1177 (2011).
44. Naparstek, S., Guan, Z. & Eichler, J. A predicted geranylgeranyl reductase reduces the W-position isoprene of dolichol phosphate in the halophilic archaeon, Haloferax volcanii.
Biochim. Biophys. Acta BBA -Mol. Cell Biol. Lipids 1821, 923-933 (2012).
45. Guan, Z., Naparstek, S., Calo, D. & Eichler, J. Protein glycosylation as an adaptive response
in Archaea: growth at different salt concentrations leads to alterations in Haloferax volcanii S-layer glycoprotein N-glycosylation. Environ. Microbiol. 14, 743-753 (2012).
46. Voisin, S. et al. Identification and Characterization of the Unique N-Linked Glycan Common to the Flagellins and S-layer Glycoprotein of Methanococcus voltae. J. Biol.
Chem. 280, 16586-16593 (2005).
47. Chaban, B., Voisin, S., Kelly, J., Logan, S. M. & Jarrell, K. F. Identification of genes involved in the biosynthesis and attachment of Methanococcus voltae N -linked glycans: insight into N -linked glycosylation pathways in Archaea: Archaeal glycosylation genes.
48. Chaban, B., Logan, S. M., Kelly, J. F. & Jarrell, K. F. AgIC and AgiK Are Involved in Biosynthesis and Attachment of Diacetylated Glucuronic Acid to the N-Glycan in Methanococcus voltae. J. Bacteriol. 191, 187-195 (2009).
49. Larkin, A., Chang, M. M., Whitworth, G. E. & Imperiali, B. Biochemical evidence for an alternate pathway in N-linked glycoprotein biosynthesis. Nat. Chem. Biol. 9, 367-373 (2013).
50. VanDyke, D. J. et al. Identification of genes involved in the assembly and attachment of a
novel flagellin N-linked tetrasaccharide important for motility in the archaeon
Methanococcus maripaludis. Mol. Microbiol. 72, 633-644 (2009).
51. Kelly, J., Logan, S. M., Jarrell, K. F., VanDyke, D. J. & Vinogradov, E. A novel N-linked
flagellar glycan from Methanococcus maripaludis. Carbohydr. Res. 344, 648-653 (2009).
52. Peyfoon, E. et al. The S-Layer Glycoprotein of the Crenarchaeote Sulfolobus acidocaldarius
Is Glycosylated at Multiple Sites with Chitobiose-Linked N-Glycans. Archaea 2010, e754101 (2010).
53. Guan, Z., Meyer, B. H., Albers, S.-V. & Eichler, J. The thermoacidophilic archaeon
Sulfolobus acidocaldarius contains an unsually short, highly reduced dolichyl phosphate.
Biochim. Biophys. Acta BBA -Mol. Cell Biol. Lipids 1811, 607-616 (2011).
54. Meyer, B. H. & Albers, S.-V. AgIB, catalyzing the oligosaccharyl transferase step of the
archaeal N-glycosylation process, is essential in the thermoacidophilic crenarchaeon Sulfolobus acidocaldarius. MicrobiologyOpen 3, 531-543 (2014).
55. Magidovich, H. & Eichler, J. Glycosyltransferases and oligosaccharyltransferases in
Archaea: putative components of the N-glycosylation pathway in the third domain of life.
FEMS Microbiol. Lett. 300, 122-130 (2009).
56. Cohen-Rosenzweig, C., Guan, Z., Shaanan, B. & Eichler, J. Substrate Promiscuity: AgIB,
the Archaeal Oligosaccharyltransferase, Can Process a Variety of Lipid-Linked Glycans.
AppL. Environ. Microbiol. 80, 486-496 (2014).
57. Igura, M. et al. Purification, crystallization and preliminary X-ray diffraction studies of the
soluble domain of the oligosaccharyltransferase STT3 subunit from the thermophilic
archaeon Pyrococcus furiosus. Acta Crystallograph. Sect. F Struct. Biol. Cryst. Commun.
63, 798-801 (2007).
58. Maita, N., Nyirenda, J., Igura, M., Kamishikiryo, J. & Kohda, D. Comparative Structural
Biology of Eubacterial and Archaeal Oligosaccharyltransferases. J. Biol. Chem. 285, 4941-4950 (2010).
59. Matsumoto, S. et al. Crystal structures of an archaeal oligosaccharyltransferase provide insights into the catalytic cycle of N-linked protein glycosylation. Proc. NatL. Acad Sci. 110, 17868-17873 (2013).
Chapter 2: Identification and characterization of archaeal glycosyltransferases
A portion of the work described in this chapter has been published in the following:
Larkin, A., Chang, M.M., Whitworth, G.E., Imperiali, B. Biochemical evidence for an alternate pathway in N-linked glycoprotein biosynthesis. Nat. Chem. Biol. 2013, 9, 367-373.
Introduction
Nature exhibits an extensive and complex variety of glycan structures in part due to the enzymatic action of glycosyltransferases (GTs), which catalyze glycan assembly through the transfer of individual monosaccharide units. Due to the diversity of monosaccharide units, multiplicity of glycosidic linkages, branching of oligosaccharides, and processing by glycosyltransferases and glycosyihydrolases, the structure of glycans on mature glycoproteins can be highly heterogeneous.' GTs most commonly use sugar nucleotide derivatives (e.g. UDP-Glc, GDP-Man, CMP-NeuAc) as glycosyl donors, but they can also use lipid-phosphate sugars (e.g. Dol-P-Man) and unsubstituted phosphate sugars as activated donors. Acceptor substrates can be other sugars, proteins, lipids, nucleic acids, antibiotics, or other small molecules.2
Currently, the Carbohydrate-Active EnZYmes (CAZy) database includes over 148,000 unique GTs that have been identified and classified into 96 families based on sequence similarities.3 These family classifications are continually updated as new sequences are added to the CAZy database and new structures are deposited into the Protein Data Bank (PDB). The first X-ray crystal structure of a GT was reported for the bacteriophage T4-glucosyltransferase in 1994.4 There are now X-ray crystal structures available for over 100 GTs in at least 38 GT families.! A comparison of these GT structures shows that while these enzymes have low sequence homology, they exhibit a surprisingly high structural homology, with two major structural folds, called GT-A and GT-B (Figure 2-1).2 Both GT folds are largely composed of ca/p/a sandwiches. GT-A folds contain a single Rossmann fold, in which the active site is located between two



![Figure 2-9: ESI-MS (negative ion mode) of purified (S)-Dol-P-GlcNAc [M+ = 1051]](https://thumb-eu.123doks.com/thumbv2/123doknet/14160464.473197/46.918.214.708.138.519/figure-esi-ms-negative-mode-purified-dol-glcnac.webp)