1
CH-π interactions play a central role in protein
recognition of carbohydrates
by
Roger Christopher Diehl B.S. Biochemistry
University of Wisconsin-Madison, 2012 M.S. Biochemistry
University of Wisconsin-Madison, 2017
SUBMITTED TO THE DEPARTMENT OF CHEMISTRY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF
DOCTOR OF PHILOSOPHY IN CHEMISTRY AT THE
MASSACHUSETTS INSTITUTE OF TECHNOLOGY February 2021
©2021 Massachusetts Institute of Technology. All rights reserved.
Signature of Author:____________________________________________________ Department of Chemistry January 15, 2021
Certified by:_________________________________________________________ Laura L. Kiessling Novartis Professor of Chemistry Thesis Supervisor
Accepted by:_________________________________________________________ Adam Willard Associate Professor Graduate Officer
2
This doctoral thesis has been examined by a committee of the Department of Chemistry as follows:
Professor Barbara Imperiali………. Thesis Committee Chair Class of 1922 Professor of Chemistry and Biology
Professor Laura L. Kiessling………. Thesis Supervisor Novartis Professor of Chemistry
Professor Matthew D. Shoulders……… Thesis Committee Member Assistant Professor of Chemistry
3
CH-π interactions play a central role in protein
recognition of carbohydrates
by
Roger Christopher Diehl
Submitted to the Department of Chemistry in partial fulfillment of the requirements for the degree of Doctor of Philosophy at the Massachusetts Institute of Technology
Abstract
Carbohydrate-protein interactions play a central role in biology, but knowledge of the forces underlying them is limited. Carbohydrates are generally hydrophilic and therefore present unique challenges in their recognition. One underappreciated force involved in
carbohydrate-protein interactions is the CH-π interaction, an attractive interaction between the aliphatic protons of a carbohydrate and the π system of an aromatic ring. In this thesis, I
examine the fundamental nature, strength, and biological significance of this interaction, largely in the context of a family of carbohydrate-binding proteins known as galectins.
In Chapter 1, I review previous knowledge of the forces underlying carbohydrate-binding proteins and the forces they utilize to bind their ligands. In particular, I focus on CH-π
interactions and galectins.
In Chapter 2, I examine the forces that contribute to CH-π interactions in the context of carbohydrates and aromatic compounds in aqueous solution. I find the CH-π interaction to be electronic in nature, and demonstrate its selectivity between different carbohydrates.
In Chapter 3, I determine the contribution of the CH-π interaction to the ligand binding of galectin-3, a human carbohydrate-binding protein of medical significance. The data
demonstrate that the CH-π interaction accounts for a majority of the binding energy.
In Chapter 4, I explore the biological implications of the CH-π interaction in galectin-3. I demonstrate that the CH-π interaction is critical for the biological activities of galectin-3.
In Chapter 5, I propose several directions future researchers could take to extend this work. For three of four directions, I present the progress I have made during my studies.
The work contained within this thesis demonstrates that CH-π interactions play a central role in protein-carbohydrate interactions at both a molecular level and a biological level.
Understanding the CH-π interaction is key to explaining and predicting the activity of carbohydrate-binding proteins.
Thesis Supervisor: Laura L. Kiessling Title: Novartis Professor of Chemistry
4 Acknowledgements
This work was only possible due to the many wonderful mentors I have had throughout my growth and development as a scientist. I cannot name all of them here, but I will name some. To my parents Tim and Sita Diehl, thank you for all your support in raising me to be the man I am today. Of relevance to this work, thank you for encouraging my curiosity as a child and enabling me to pursue my dreams. To Professor Nancy A Rice at Western Kentucky University, thank you for giving me my first experience in a modern biochemistry laboratory at VAMPY 2005. Science has changed since then but my passion for it has not. To Ms. Lucy Organ at Hillsboro High School, thank you for helping me realize that chemistry in particular is my field of interest. I am still a proud chemist, and still a proud Burro. Professor Lauren Buchanan, thank you for making my first chemistry class of college enjoyable, and for helping me handle the stress of my first first-author paper. Professor M Thomas Record, thank you for taking me into your lab as a freshman and teaching me techniques and principles that serve me well to this day. Chapters 2 and 3 in particular greatly benefitted from your perspective. Dr. Emily J. Guinn, thank you for being the best graduate student mentor an undergraduate researcher could ask for. You knew of my potential as a scientist when I doubted it, and helped me build the
confidence I needed to make it through graduate school. Your example is one I look to whenever I am called upon to teach a fellow scientist.
Last but definitely not least, thank you Professor Laura L. Kiessling. Thank you for seven excellent years in your lab, and in particular for giving me the opportunity to design the projects in this thesis. You have placed great trust in me over these years, and I sincerely hope you feel that I have rewarded that trust. Thank you also for ensuring a positive workplace environment in Kiessling Lab, even as students and postdocs have come and gone and the lab has moved from Wisconsin to MIT. Science is never easy and it is of the utmost importance that it takes place in a supportive context.
Finally, I would like to thank all of the members of Kiessling Group that I have worked with during my time as a graduate student. All of you have been wonderful colleagues, and have helped me learn a broad array of disciplines and techniques ranging from vaccine development to reverse phase high-pressure liquid chromatography. I would particularly like to thank two subsets of Kiessling Group members. First, I would like to thank Dr. Christine Isabella, Dr. Alex Justen, Dr. Cassie Jarvis, and Professor Caitlin McMahon for organizing the move to MIT. It was an unforgettable experience and you four were key in making sure it proceeded as smoothly as possible. Second, I would like to thank Stephen Early, Melanie Halim, Alan Carter, Dr. Robert Brown, Dr. Mohammad Murshid Alam, Dr. R. Lyle McPherson, and Dr. Amanda Dugan for their contributions to the work in this thesis. Again, you have all been wonderful to work with.
5
Table of Contents
Abstract………3
Acknowledgements………4
Table of Contents……….5
List of Tables and Schemes………...7
List of Figures………8
Chapter 1: Molecular Features of Lectin-Carbohydrate Interactions……….10
Introduction………11
Previously described features of carbohydrate-binding sites………..13
Previously described features of CH-π interactions………..15
Biophysical background regarding galectins……….17
Biological background regarding galectins………20
Conclusions and present work………..22
References…………..……….25
Chapter 2: Electronic Nature of CH-π Interactions……….36
Background and significance……….37
Experimental section……….38
1H NMR assay shows CH-π interactions in aqueous solution……….40
CH-π interactions between carbohydrates and aromatic groups are electronic in nature……….43
CH-π interactions display selectivity between monosaccharides and anomers……….45
Conclusions and future directions………..48
Acknowledgements……….50
6
Chapter 3: A CH-π interaction drives glycan-binding to human galectin-3…………52
Background and significance……….53
Galectin-3 bears a conserved tryptophan centrally located in the binding site……….54
Materials and methods……….55
Galectin-3 variants are stable at room temperature……….57
Galectin-3 variants have reduced binding affinity towards lactose……….60
CH-π interactions account for the majority of binding energy in galectin-3………..63
Acknowledgements……….64
References………64
Chapter 4: Biological functions of the CH-π interaction in galectin-3………67
Roles and structures of human galectins………68
Full-length galectin-3 variants have reduced agglutination activity towards mouse red blood cells……….69
Wild-type galectin-3C binds to secreted mucins, but variants at W181 do not………..71
Wild-type galectin-3 is exported from HEK-293 cells, while variants at W181 are retained in the cell……….73
Conclusions……….76
Acknowledgements………..77
References……….77
Chapter 5: Further Avenues for Investigation………80
Hammett series to determine electronic nature of CH-π interactions in a protein……….81
Design of glycomimetic ligands with enhanced CH-π interactions……….………86
Conferral of CH-π interactions on existing antiretroviral lectins……….90
Characterization of tandem repeat galectin binding profiles to mammalian and microbial glycans……….93
Acknowledgements……….98
7 List of Tables and Schemes
Chapter 2: Electronic Nature of CH-π Interactions
Scheme 1: Deprotection of methyl 3,4,6-tri-O-benzyl-β-D-mannopyranoside…………...39
Chapter 3: A CH-π interaction drives glycan-binding to human galectin-3
Table 1: Strength of CH-π interactions with lactose at position 181 by variant………62
Chapter 5: Further Avenues for Investigation
Scheme 1: Two-step synthesis of tryptophan analogs from indole precursors………..82 Scheme 2: One-step chemoenzymatic synthesis of substituted tryptophan analogs……..82
8 List of Figures
Chapter 1: Molecular Features of Lectin-Carbohydrate Interactions
Figure 1: Classes of carbohydrate-binding proteins……….11 Figure 2: Aromatic residues are overrepresented in carbohydrate binding sites………….15 Figure 3: Electronic and dispersive forces contribute to the strength of CH-π interactions between carbohydrates and aromatic groups………16 Figure 4: Galectin subfamilies are based on domain structure and
mode of oligomerization………17 Figure 5: Conserved residues of the galectin binding site subsite B……….18 Figure 6: The galectin-glycoprotein lattice……….21 Chapter 2: Electronic Nature of CH-π Interactions
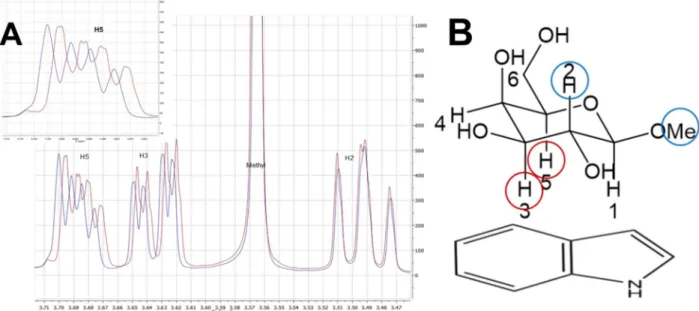
Figure 1: Methyl β-galactoside and indole form a geometrically defined complex
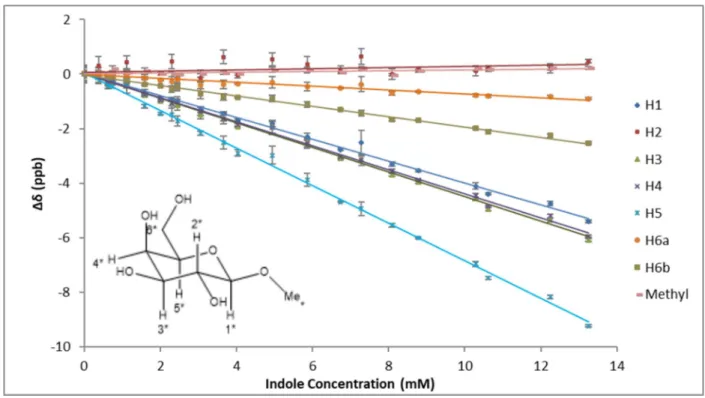
in aqueous solution……….42 Figure 2: Stacking geometry is consistent for all indole concentrations tested……….43 Figure 3: Carbohydrate-aromatic interactions in aqueous solution are
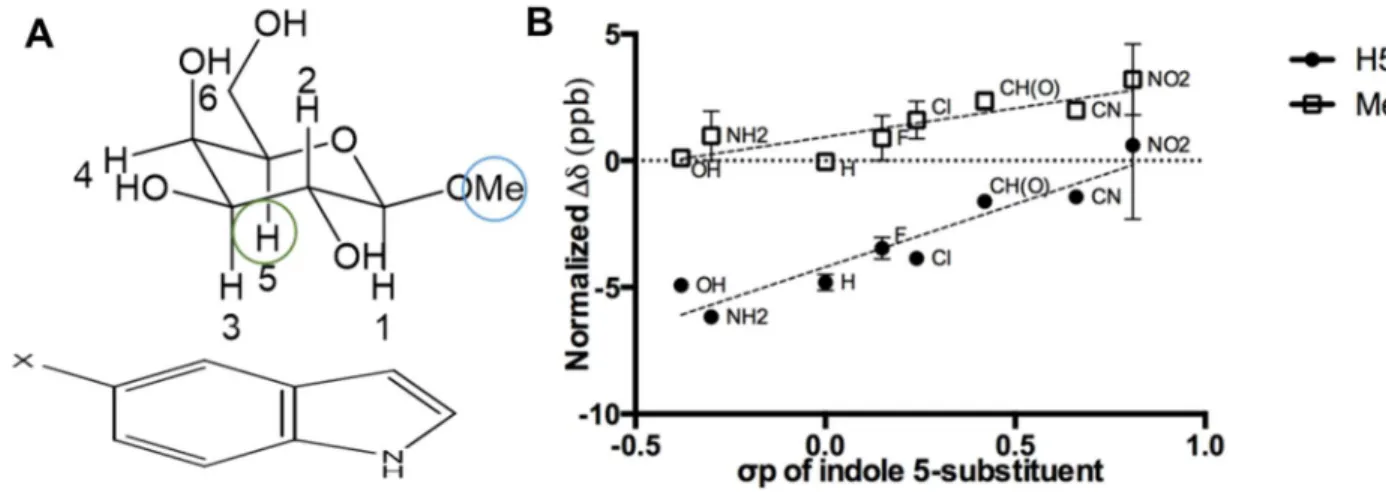
electronic in nature……….44 Figure 4: CH-π stacking geometries in solution resemble patterns found in
carbohydrate binding sites………..46 Figure 5: CH-pi interactions involving fucose and N-acetylglucosamine resemble those involving galactose and glucose respectively……….47 Chapter 3: Mutational tuning of the CH-π interaction in galectin-3
Figure 1: Structure of the galectin-3 binding site………57 Figure 2: Most galectin-3 variants at W181 are stable at room temperature………..59 Figure 3: Galectin-3C variants show decreased binding to lactose………..60 Chapter 4: Biological functions of the CH-π interaction in galectin-3
Figure 1: The W181 CH-π interaction is necessary for galectin-3 induced
hemagglutination……….70 Figure 2: Binding of galectin-3 to mucins is a carbohydrate-protein interaction
dependent on the W181 CH-π interaction………..73 Figure 3: Wild-type galectin-3 is robustly exported from HEK-293 cells, while variants are retained within the cell……….75
9 Chapter 5: Further Avenues for Investigation
Figure 1: Fluorinated analogs have similar lactose binding to wild-type galectin-3C……85 Figure 2: The CH3 bond of β-galactose is polarized by overlap with the
antiperiplanar CO4 hydroxy group………..87 Figure 3: Proposed monosaccharide analogs to test the relative importance of
contributors to CH-π interaction strength………....88 Figure 4: Algal and bacterial lectins recognize diverse motifs on HIV gp120
Man9 glycans………..91
Figure 5: Cyanovirin-N variants at T25 are stable at room temperature………..92 Figure 6: Tandem repeat N- and C-terminal domains are similar to each other…………..94 Figure 7: Tandem repeat galectin domains are stably folded at room temperature………96
10
Chapter 1: Molecular features of lectin-carbohydrate
interactions
This chapter is reproduced in part with permission from Hudson, K. L.; Bartlett, G. J.; Diehl, R. C.; Agirre, J.; Gallagher, T.; Kiessling, L. L.; Woolfson, D. N. Carbohydrate-Aromatic
Interactions in Proteins. J. Am. Chem. Soc. 2015, 137 (48), 15152–15160. Copyright 2015 American Chemical Society.
11 Introduction
Glycans are found in all cells1 and play crucial roles in many facets of biology. Some
primarily play structural roles, such as cellulose in wood, chitin in fungi and arthropods, and chondroitin in cartilage.2 Others, such as starch and glycogen, act as readily accessible energy
sources. However, one of the most diverse and necessary roles glycans play is as conveyors of information. Carbohydrates excel in this role because they not only vary in their linear sequence, but also the nature of the linkages between each monomer. Additionally, the multiple allowed linkages in a carbohydrate allows for branched structures, adding yet another element of diversification even in a small oligosaccharide.3 Oligosaccharides bear a narrower range of
functional groups than proteins and do not appear to form intricate secondary structures as RNA does, so most biological functions of carbohydrates require a protein to recognize the involved carbohydrate.4–6 Herein lies another challenge; carbohydrates are highly hydrophilic
and afford little binding energy through the hydrophobic effect, a source of binding energy that is essential to many other classes of ligand. Proteins that recognize carbohydrates must
therefore bring other forces to bear in order to bind their ligand. Carbohydrate-binding proteins
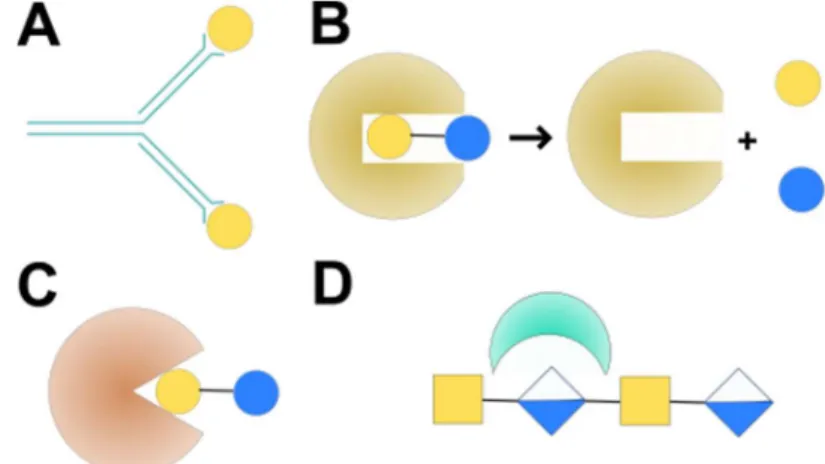
generally fall into four classes: antibodies, carbohydrate-modifying enzymes, GAG-binding proteins, and lectins (Figure 1.1). Given that
pathogens, like all cellular organisms, are coated in a carbohydrate-based glycocalyx,7 it is little surprise that the
human immune system generates antibodies against glycans.8,9 Due to
the difficulty of strongly binding a
Figure 1: Classes of carbohydrate-binding proteins. A) Antibodies generally bind carbohydrates loosely due to the hydrophilic nature of the ligand. B) Enzymes bind
carbohydrates tightly in order to modify them. C) Lectins have evolved for recognition of specific binding epitopes on glycans. D) GAG-binding proteins bind anionic
12
carbohydrate ligand, carbohydrates tend to be poorly immunogenic but carbohydrate-binding antibodies do occur, especially when the carbohydrate is presented on a peptide scaffold as would be the case with an enzymatically digested glycoprotein.10 Unlike other
carbohydrate-binding proteins, carbohydrate-carbohydrate-binding antibodies can often bind peptides in the same site, though generally by taking advantage of different features of the binding site than are used to bind carbohydrates.11 While relatively low in affinity,12 carbohydrate-binding antibodies display
many similar features to other carbohydrate-binding proteins, such as a heavy reliance on aromatic residues13 and extended binding sites.12 Due to the low monovalent affinity,
multivalency is crucial for antibody binding to carbohydrate targets.14
Another class of proteins that must recognize carbohydrates includes the enzymes that modify them. While carbohydrate-modifying enzymes have a range of specificities,15 they
generally have tighter, more encompassing binding sites in order to position the sugar residues properly for catalysis.16 In many cases, this binding is aided by the presence of a nucleotide
attached to the sugar, where the nucleobase and phosphate can afford substantial affinity.17
Covalent binding and transition metals can help to position the substrate for catalysis.18 Also,
the fine specificities of these enzymes are often conferred by a lectin-like carbohydrate-binding module that does not possess catalytic activity itself.19 Otherwise, many of the trends observed in
other carbohydrate-binding proteins also apply to enzymes.
Highly anionic carbohydrates known as glycosaminoglycans (GAGs) play key roles in mammalian development via cell-cell and cell-matrix interactions;20 These interactions are
mediated by a broad range of GAG-binding proteins.21 A common feature of such proteins is
their reliance on cationic amino acids for recognition, often in the pattern XBBXBX or
XBBBXXBX, where X is an uncharged residue and B a basic residue (cationic at neutral pH).22
These cationic motifs bear some similarity to those found in anti-microbial peptides, and there are a number of peptides with both GAG-binding and antimicrobial activity for this reason.23
13
These interactions are particularly strong — displaying a peptide consisting of the immediate binding site alone on a surface can allow adhesion of cells via their proteoglycans,24 whereas
with other classes of carbohydrate-binding protein, the entire protein is generally needed to yield a high-affinity interaction. The central principle involved is an electronic interaction between the positively charged GAG binding site and the negatively charged GAG.
Finally, lectins are a class of proteins evolved to bind carbohydrates, usually without any direct enzymatic activity. While the earliest work on lectins focused on the abundant and readily isolable plant lectins,25 lectins exist in all domains of life26 in a variety of roles. Some are toxic,
such as those in rattlesnake venom27 or cholera toxin,28 while others are essential for life, such as
XEEL, a lectin that forms a protective gel around Xenopus laevis eggs.29 To better understand
such a diverse category of proteins, lectins are classified into families based on their fold and generalizations regarding their binding specificity.30 Due to their relatively strong affinities and
relatively flexible specificities, I focused on a family of lectins known as the galectins in my studies of the mechanisms behind carbohydrate-protein interactions. That said, many of the conclusions reached here are likely applicable to enzymes and antibodies as well, and additional facets of GAG-binding, enzyme-carbohydrate, and antibody-carbohydrate interactions warrant further study.
Previously described features of carbohydrate binding sites
Given the challenges of glycan recognition, I analyzed data on how lectins typically bind carbohydrates. Weis and Drickamer published seminal work in this field in 1996,16 which
identified several forces that play a key role in binding carbohydrates, with a focus on the binding activities of lectins. The first force Weis identified, and perhaps the most obvious, is direct hydrogen bonding. Interestingly, this typically does not involve protein hydroxy groups such as those of serine, threonine, or tyrosine, likely due to the entropic cost of simultaneously fixing the rotamers of the protein and sugar hydroxy groups. A noted exception is the frequent
14
occurrence of serine hydroxy groups in binding sites for sialic acid, where the hydroxyl groups act as hydrogen-bond donors to the sialic acid carboxylate. In a number of cases where highly structured water molecules are present, water-mediated protein-carbohydrate hydrogen bonds can be found. In isolation, hydrogen bonds are strong, with a bond energy of 3 to 7 kcal/mol.31 A
central challenge for lectins is that hydrogen bonding is also present in bulk water, and lectins only function if they can generate a preference for the ligand binding over its retention in bulk water.3 Hydrogen bonding plays a far greater role when carbohydrates are bound in organic
solvents,32 or for generating selectivity between different carbohydrate ligands.33 Hydrogen
bonding can also be crucial for restricting the mobility of the ligand, as in the case of xylan carbohydrate-binding modules.34 A related effect, the displacement of water from a
preorganized binding site, can provide additional affinity for some lectins.35,36
A second group of forces available is those involving cations, especially bound divalent metals such as calcium and manganese. C-type lectins and intelectins use calcium ions, with the ion accepting coordinate bonds from lone pairs on two vicinal hydroxy groups on the sugar.29,37– 40 Intelectins use calcium ions in a similar role. Arginine residues can play a similar role, with
the electronic effects of proximity to partially negative hydroxyl oxygen atoms complementing the attractive strength of hydrogen bonds via the guanidinium protons.41 Cationic residues also,
as noted, play a large role in binding glycosaminoglycans42- interestingly divalent metals are not
as common in these cases.
Weis and Drickamer also examined the role of nonpolar residues in carbohydrate binding sites.16 Here, they noted that this form of recognition was most prevalent for galactose
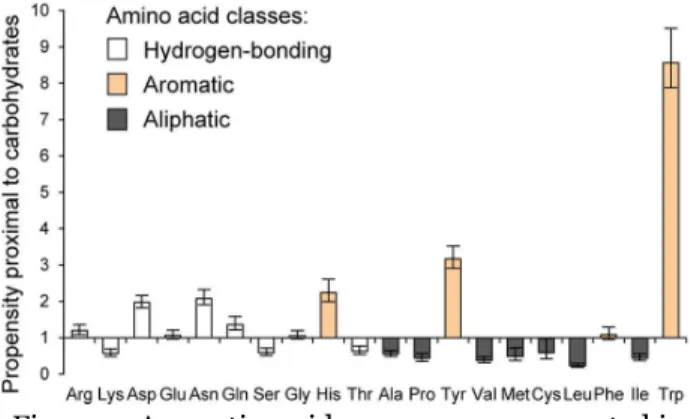
15 the sugar. On the face of it, this seems to be a surprising way to recognize such a hydrophilic ligand as galactose, and it bears mentioning that the residue involved is typically an aromatic residue (Figure 2).43 Similarly,
aromatic residues excel at binding N-acetylgalactosamine (GalNAc).44 These
interactions are also particularly prominent in cellulases, as the highly ordered β-glucosides can be sandwiched by tyrosine and tryptophan
residues to separate them from water or other polysaccharide chains.45 Likewise, human
α-galactosidase uses tyrosine and tryptophan residues to bind the α-face of α-galactose away from the anomeric position while the enzyme distorts the geometry of the sugar to allow for
cleavage.46
Previously described features of CH-π interactions
Based on the observed requirement for an aromatic residue, it is likely that the
interaction at work when tryptophan, phenylalanine, or tyrosine is in the binding site is a CH-π interaction. Thus, in order to comprehend the role of aromatic residues in carbohydrate
binding sites, one must understand the features of CH-π interactions. In the 1980s, it became apparent that under certain circumstances, aliphatic and aromatic CH protons could act as hydrogen bond donors to oxygen or nitrogen-based hydrogen bond acceptors.47 By the same
token, canonical hydrogen bond donors consisting of OH or NH protons were observed
interacting with π-systems, especially those of aromatic compounds, with the π system playing a role of a hydrogen bond acceptor.48 This led to the notion that a CH proton could bond with a
π system in an analogous manner, as Motohiro Nishio demonstrated computationally in 1995.49
Further computational work by the Chandrasekhar group showed that this interaction could
Figure 2: Aromatic residues are overrepresented in carbohydrate binding sites. The horizontal axis is ordered with increasing hydrophobicity to the right, while the vertical axis shows the ratio between the occurrence of a given amino acid within 4 Å of a noncovalently bound glycan compared to the frequency in the UniProt database. Figure adapted from Hudson et. al. 2015.
16
occur between two aromatic systems, and that CH-π interactions are somewhat stronger for heterocyclic aromatic systems than benzene, and that the orientation of a single hydrogen directly towards the aromatic ring is preferred over two hydrogens angled towards opposite edges of the ring.50
Earlier computational works generally cite dispersion forces as providing the majority of the interaction energy, aided by the high polarizability of the C-H bond.51 However, the CH bonds of an
aliphatic hydrocarbon such as methane grant only a modest interaction with an aromatic π system.50,52
Partial substitution of methane with chlorine and fluorine, by contrast, results in a considerably stronger interaction between the remaining
hydrogens and the benzene π system,52–54 suggesting
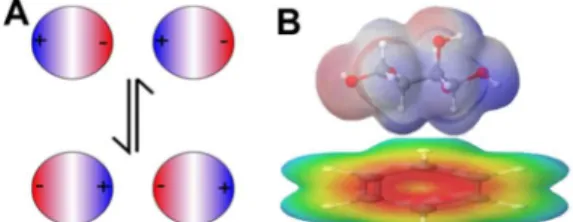
that a strong dipole moment leads to a large electronic contribution (Figure 3).55 That there
would be a strong electronic contribution was also investigated in the context of π-π
interactions mediated through aromatic CH bonds, in the case of substituted benzenes, where an electron-withdrawing substituent on the edge-on aromatic ring resulted in a stronger interaction.56
In recent years, the CH-π interaction has been explained in terms of hard/soft acid-base theory, where it represents a hydrogen bond between a soft acid (the CH group) and a soft base (the π system).57 The ability of CH bonds to act as soft acids is demonstrated by their preference
for softer chloride over harder oxyanions in synthetic anionophores.58 It is clear that both
electronic forces and the hydrophobic effect are important in mediating these interactions in water,59 and furthermore that the interaction is highly solvent-dependent60,61 and often
cooperative.62,63 CH-π interactions are useful in achieving fine control over organic synthesis
Figure 3: Electronic and dispersive forces contribute to the strength of CH-π interactions between carbohydrates and aromatic groups. A) Dispersive forces rely on the high polarizablility of C-H bonds to create transient partial charges in both molecules, which oscillate to maintain an attractive interaction. B) Electronic forces take the form of a dipole-quadrupole interaction between the partially positive face of the sugar and the partially negative face of the aromatic group.
17
reactions via shifting the balance between transition states.64 The balance of electronic and
dispersive effects has been an active area of debate. In the case of nonpolar CH groups63,65 and to
a lesser extent weakly polarized carbohydrates such as β-GlcNAc66 the dispersive component is
dominant, but strongly polarized carbohydrates such as β-Gal are more likely to be found in proximity to an aromatic group43 and in these cases the electronic component is likely large.67 In
the case of carbohydrates, the cooperativity of the interaction means that aromatic rings have some selectivity in favor of monosaccharides that can make three CH-π interactions.68 By
contrast, furanosides, which have flexible ring structures with poor complementarity with the aromatic systems, are unlikely to make CH-π interactions.69 Computational studies suggest that
the indole group of tryptophan, due to its greater electron richness,70 should be a better CH-π
acceptor than a benzene ring as found on phenylalanine,71 which is reflected in the
overrepresentation of tryptophan in carbohydrate binding sites.43 Histidine, by contrast, has an
especially electron-poor π system and generally serves as a hydrogen donor when it is found in a carbohydrate binding site.43,72
Biophysical background regarding galectins Galectins are a family of proteins
found across animal taxa, distinguished by their affinity for β-galactosides and the distinctive “jelly roll” fold of their
carbohydrate recognition domain.73 Fifteen
are described in mammals, of which eleven are found in humans.74,75 Galectins are found
in a broad range of tissue types, and in
humans galectins-1 and 3 are ubiquitously expressed.76 Mammalian galectins consist of three
subfamilies, based on their domain structure and oligomerization (Figure 4). Prototype
galectins, such as galectins-1, 2, and 7, contain a carbohydrate recognition domain and no other
Figure 4: Galectin subfamilies are based on domain structure and mode of oligomerization. A) Prototype galectins consist of a single carbohydrate-recognition domain and form noncovalent dimers. B) Chimeric galectins have a proline-rich N-terminal domain that forms flexible oligomers. C) Tandem repeat galectins consist of two carbohydrate recognition domains connected by a disordered linker.
18
domain.76 These galectins are capable of noncovalent dimerization with the two monomers
oriented in opposite directions, but generally exist in a monomer-dimer equilibrium in solution, favoring the dimer upon binding to ligand.77 Tandem-repeat galectins, such as galectins-4, 8,
and 9, possess two distinct carbohydrate recognition domains and a linker of variable length and composition.78 The domains have differing carbohydrate specificity, the implications of which
are an active area of study.79
Galectin-3, the focus of my studies, exists in a subfamily all to its own as a chimera-type galectin. It possesses a C-terminal carbohydrate recognition domain highly similar to other galectins in the active site, but its N-terminal domain is a loosely ordered proline-rich helix that engages in numerous protein-protein interactions inside and outside of the cell.80 These
interactions include a highly flexible form of oligomerization; galectin-3 exists as a monomer in solution81, but oligomerizes upon binding multivalent ligands82, creating a galectin-glycan lattice
that can force receptor clustering or mediate host-pathogen interactions.83 Prototype galectin
dimers and bivalent tandem repeat galectins can form similar lattices as well. Galectins are generally unable to
benefit from multivalent binding to a single ligand, as their carbohydrate binding domains are oriented away from each other. In order to maximize binding affinity for their hydrophilic and usually uncharged ligand, galectins
instead utilize extended binding sites, where as many as four monosaccharide units are
recognized by the binding site, in subsites A, C, and D situated on either side of the required β-galactose or β-GalNAc residue,74,84 which
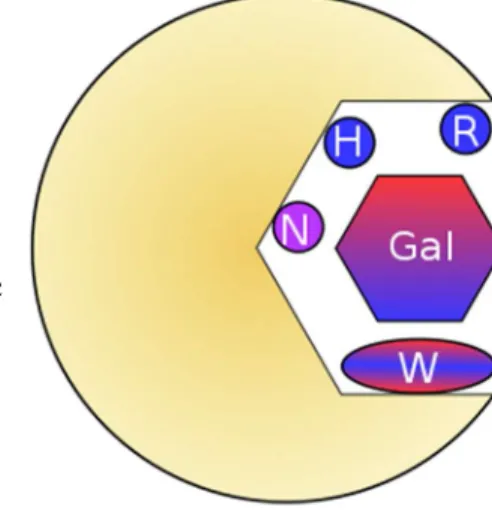
binds in subsite B. Like other cases of extended binding sites in lectins, the
Figure 5: Conserved residues of the galectin binding site subsite B. Bifurcated hydrogen bond donors and acceptors bind the hydroxy groups of β-galactose, while tryptophan makes a CH-π interaction with the α-face of the sugar.
19
peripheral monosaccharides (often Glc or GlcNAc, Fuc, or another Gal/GalNAc) need not be able to bind as monosaccharides, as their binding affinity is individually weaker than the
primary determinant β-Gal/β-GalNAc.16 In the case of galectin-3,β-galactose is able to bind as a
monomer, but its affinity is a meager 5-20 mM.84 The canonical ligand traditionally used to
block galectin-3 binding in biological assays is lactose (Gal-β1-4-Glc), for which galectin-3 has a binding affinity of approximately 100 μM.85 However, further elaboration of the glycan can
provide additional affinity approaching two orders of magnitude.86 In the case of an extended
LacNAc epitope, the stronger binding provided by a tetrasaccharide epitope is nearly as
beneficial as arranging for divalent binding via an alkyl linker.87 This additional affinity and the
widespread occurrence of lactose and lactosamine epitopes in mammalian glycans render galectins potent binders of carbohydrate self-epitopes, but in a far more selective manner than plant lectins such as wheat germ agglutinin.88
All vertebrate galectins possess a tryptophan residue in the β-galactose-binding subsite (Figure 5).89 In the case of galectin-3, this residue is Trp181, and notably is quite mobile in
apo-galectin-3 but is far more ordered in lactose-bound apo-galectin-3.81 A conserved arginine, R186 in
galectin-3, near the β-face of the sugar hydrogen bonds with O4 and O6 of the β-galactose residue and may help polarize the molecule to aid the CH-π interaction. Arginine and lysine are also heavily represented in the other subsites, especially subsite A in at the non-reducing end of the binding site,90 and these arginines are heavily targeted for inhibitor development via
cation-π interactions with aromatic substituents on the inhibitor.91 Due to the high conservation of
subsite B, efforts at mutational tuning of galectin-3 to alter its specificity have focused on the other subsites.92 As such, the precise importance of the subsite B residues remains
underexplored.
20
Galectins in general, and galectin-3 in particular, are heavily targeted for inhibitor development due to their myriad biological roles.93 First of all, galectin-3 tends to promote
tumor progression by a number of activities, most of which revolve around inducing clustering in N-glycosylated cell surface receptors.94 Through binding and clustering integrins, galectin-3
can promote cell mobility.95 Galectin-3 plays a key role in cell-matrix signaling,96,97 including
polymerizing hensin through its carbohydrate-binding activity.98 In tumors, strong galectin-3
ligands are upregulated, allowing galectin-3 to participate in the remodeling of the extracellular matrix that allows for invasion and metastasis.99 Galectin-3, while soluble, is able to act as a
receptor for transferrin possessing tri-antennary and tetra-antennary N-glycans, aiding the recycling of bound transferrin to the cell surface.100 Galectin-3 also promotes angiogenesis
through acting as a chemoattractant for endothelial cells101, and promotes metastasis through its
interaction with MUC1102 and via promotion of cytokine secretion.103 Overall, these activities link
upregulation of galectin-3 to increased tumor progression and worse prognoses.104
Secondly, galectin-3 is a potent immunomodulator.105 All immune cells express
galectin-3, and just as in cancer its ability to form a galectin-receptor lattice (Figure 6) is key to its function.106 Galectin-3 binding to T cell receptor can prevent the T cell receptor from clustering
around CD8, leading to anergy in cytotoxic T lymphocytes.107 Extracellular galectin-3 can also
induce apoptosis in T cells through binding to CD45 and clustering of CD71,108 even as
intracellular galectin-3 has the opposite effect, likely through protein-protein interactions
mediated by the N-terminal domain.109 Galectin-3 activates the JAK-STAT pathway, likely via its
N-terminal domain as this activity cannot be inhibited by lactose.110 Through recognition of
lactosamine-containing N-glycans, galectin-3 serves as a signal to the immune system that self-epitopes are present.111,112
21 Thirdly, galectin-3 can interact directly with pathogens and the microbiome.113 Through
its recognition of self-like lactosamine-containing epitopes, galectin-3 can bind bacteria that
express these epitopes as a mimicry
mechanism.114 This allows galectin-3 to bind a
variety of bacterial pathogens from K. pnuemoniae to H. pylori.115 In the case of H.
pylori, some studies have suggested a direct cytotoxic effect resulting from binding to Lewis antigens on the bacterial cell membrane.116 For
LPS-displaying bacteria, galectin-3 can bind the LPS and reduce resulting inflammation.117 Even
in cases where galectin-3 does not bind bacteria or pathogenic protists, as with Shigella and T. cruzi, it is associated with phagocytosis and vacuole lysis.118,119 Galectin-3 can also bind to some
commensal microbiota such as B. longum, and it does not appear to harm these bacteria.120 In
fact, as galectin-3 also binds strongly to intestinal mucins,121 it may serve to retain the
commensal microbiota.
One question these studies leave open is quality control. In canonical protein export, secreted proteins are N-glycosylated in the ER and Golgi,122 and structurally defective proteins
are routed to lectin chaperones for refolding.123 Like other galectins, human galectin-3 is
translated on free polysomes in the cytoplasm75 and is not trafficked via the Golgi,124 so it cannot
benefit from this quality control pathway. That said, export of misfolded galectin-3 variants could be highly deleterious considering both its biological role and that galectin-3 bears an unpaired cysteine that is buried if the protein is properly folded.97 For this reason, my work
Figure 6: The galectin-glycoprotein lattice. Galectin dimers or oligomers (brick) bind lactosamine-containing glycans (yellow/blue) on glycoproteins (grey) with multiple N- or O-glycosylation sites. This draws the glycoproteins and galectins into an ordered arrangement, inducing clustering of the glycoproteins.
22
explored the effect of W181 mutations that impair binding on galectin-3 export from HEK-293 cells.
Galectin-3 has six known polymorphisms in humans that occur with a frequency of 0.1% or greater.125 A225 is replaced with aspartate or valine 46% of the time, though no study has yet
identified any medical consequences of this variation. Likewise, no medical relevance has been found for the polymorphisms leading to the variations A53D, R183K, R212L, or R212Q.
Polymorphisms of medical relevance are found at two sites. First, T98 is replaced by proline in 43% of humans- this is associated with a modestly increased rate of gastric cancer.126 Second,
P64 is replaced with histidine 29% of the time- this polymorphism is associated with considerably worse cancer prognoses127. The only known variation of W181 is a somatic
mutation to arginine that was found in a gastric tumor128.
Conclusions and present work
To investigate the importance of CH-π interactions in carbohydrate binding sites of lectins, I first chose to examine CH-π interactions in the simplest relevant context available, namely the interaction between a single monosaccharide and indole, choosing the latter because it is the functional group of tryptophan, which in turn is heavily overrepresented in
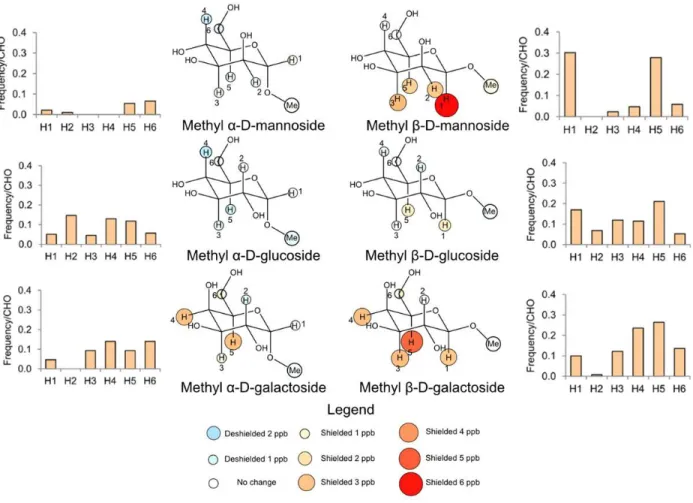
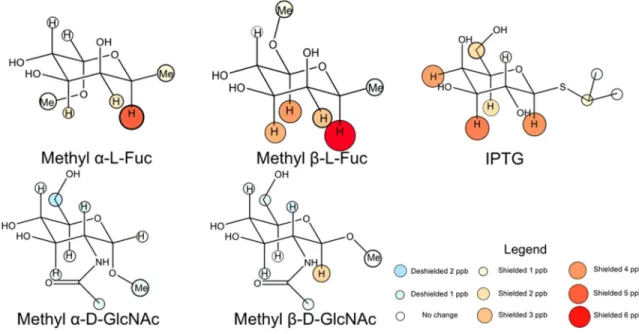
carbohydrate binding sites.43 In Chapter 2, I measured the strength of these interactions using a 1H NMR based assay, in which the change in chemical shift of glycoside protons involved in
CH-π interactions indicates the likelihood that they stack on the aromatic ring.129,130 I repeated this
process for α- and β- anomers of mannose, glucose, and galactose, and for a Hammett series of substituted indoles.43,131
As the results of these experiments showed a strong electronic contribution to CH-π interactions in a biologically relevant small molecule context,43 I sought in Chapter 3 to examine
the interaction in the context of a protein. After consideration of several β-galactose binding lectins, I chose to work with human galectin-3 due to its ease of bacterial expression,132 its
23
medical relevance,133–136 and the availability of sub-Ångstrom crystal structures showing three
β-galactose protons favorably oriented for a CH-π interaction with the invariant tryptophan residue W181.137,138 I generated a range of variants at W181 to test the importance of this CH-π
interaction, measuring affinity by isothermal titration calorimetry, with a procedure optimized for measuring low affinity interactions.139 I chose phenylalanine and tyrosine as they present a
smaller aromatic π system than tryptophan, and to test the reason behind why phenylalanine is less overrepresented than tyrosine in carbohydrate binding sites.43 I chose histidine as it bears a
particularly electron-poor π system that would have weaker electronic interactions with the β-galactose residue43, but it is still aromatic. I chose methionine as a model non-aromatic residue,
as modeling the mutation in PyMOL140 indicated better shape complementarity than any other
non-aromatic residue. I generated an alanine variant but shortly found that it was unfolded at room temperature. Finally, I cloned and expressed an arginine variant, as the W181R somatic mutation has been found in a gastric cancer,128 and I wished to understand the effect that said
mutation would have on the folding and activity of the resulting protein. In addition to calorimetric assays, I used differential scanning fluorimetry141 to determine the denaturation
temperature (Tm) of each variant as a measure of its stability. For better handling, I conducted
these studies with a C-terminal construct that is commonly used for crystallography91,137,138 and
is present biologically as an enzymatically produced competitive inhibitor of biological activities of galectin-3.111
With this biophysical data in hand, I then explored the importance of the galectin-3 CH-π interaction in a biological context in Chapter 4, in order to determine if the CH-CH-π interaction still plays a crucial role with a larger ligand and highlight its importance to health. The first activity I chose to investigate was the well-established ability of galectin-3 to agglutinate red blood cells through binding to their Lewis antigens142,143. I tested this activity for wild type and
W181 variant galectin-3 through both a microscopy-based aggregation assay and a plate-based hemagglutination titer assay. Additionally, I sought to determine the role of the CH-π
24
interaction in more newly described galectin-3 activities. To this end, I applied wild type and variant galectin-3 to a panel of mucins purified by the Ribbeck Group144 in a nitrocellulose
dot-blot assay to determine if CH-π interactions are essential for binding to the
lactosamine-containing O-glycans that mucins bear.121 Finally, I expressed wild type and variant galectin-3 in
HEK-293 cells to determine if the CH-π interaction is necessary for export of the protein from a human cell.
While these results answer many questions about the nature of CH-π interactions and their role in carbohydrate binding by proteins, they also open up a number of new avenues for further study. Among these are three projects that extend beyond the scope of my doctoral research, and one that I have envisioned but not initiated. These projects are outlined in Chapter 5. First, the logical synthesis of my small molecule Hammett series and my mutagenesis of galectin-3 would be to use non-natural amino acid incorporation within the context of a lectin. While this is a more challenging effort than I had originally anticipated, my preparatory research and experience bear lessons for those who would bring such a project to completion. Second, varying groups on the carbohydrate to make it a stronger CH-π donor could extend the small molecule portion of my project. This could be further elaborated into developing stronger galectin inhibitors, as current inhibitors mainly vary outlying areas of the binding site rather than the β-galactose residue itself.145 Third, a future researcher could demonstrate the potency
of CH-π interactions by conferring them upon a lectin that does not use them in nature. I have explored the possibility of doing so for cyanovirin-N, a lectin that can bind the high-mannose glycans of HIV protein gp120, but does so without recognizing the β-mannose residue at the core of the glycan that would be an excellent CH-π donor.146,147 Finally, the biological portion of
the project could be extended to the tandem repeat galectins, namely galectins-4, 8, and 9. These galectins have noted antimicrobial activity,148 so assaying their binding to pathogens or
isolated microbial glycan would shed light on the mechanism by which they operate. These projects will ensure many paths of investigation for future researchers.
25 References
(1) Varki, A. Evolutionary Forces Shaping the Golgi Glycosylation Machinery: Why Cell Surface Glycans Are Universal to Living Cells. Cold Spring Harb. Perspect. Biol. 2011, 3 (6), 1–14.
(2) Koch, B. E.; Stougaard, J.; Spaink, H. P. Keeping Track of the Growing Number of Biological Functions of Chitin and Its Interaction Partners in Biomedical Research. Glycobiology 2015, 25 (5), 469–482.
(3) Jayaraman, N. Multivalent Ligand Presentation as a Central Concept to Study Intricate Carbohydrate-Protein Interactions. Chem. Soc. Rev. 2009, 38 (12), 3463–3483. (4) Zhuo, L.; Kanamori, A.; Kannagi, R.; Itano, N.; Wu, J.; Hamaguchi, M.; Ishiguro, N.;
Kimata, K. SHAP Potentiates the CD44-Mediated Leukocyte Adhesion to the Hyaluronan Substratum. J. Biol. Chem. 2006, 281 (29), 20303–20314.
(5) Pipirou, Z.; Powlesland, A. S.; Steffen, I.; Pöhlmann, S.; Taylor, M. E.; Drickamer, K. Mouse LSECtin as a Model for a Human Ebola Virus Receptor. Glycobiology 2011, 21 (6), 806–812.
(6) Wang, S.-F.; Tsao, C.-H.; Lin, Y.-T.; Hsu, D. K.; Chiang, M.-L.; Lo, C.-H.; Chien, F.-C.; Chen, P.; Arthur Chen, Y.-M.; Chen, H.-Y.; et al. Galectin-3 Promotes HIV-1 Budding via Association with Alix and Gag P6. Glycobiology 2014, 24 (11), 1022–1035.
(7) Varki, A. Biological Roles of Glycans. Glycobiology 2017, 27 (1), 3–49.
(8) Muthana, S. M.; Xia, L.; Campbell, C. T.; Zhang, Y.; Gildersleeve, J. C. Competition between Serum IgG, IgM, and IgA Anti-Glycan Antibodies. PLoS One 2015, 10 (3), 1–17. (9) Shilova, N.; Navakouski, M.; Khasbiullina, N.; Blixt, O.; Bovin, N. Printed Glycan Array:
Antibodies as Probed in Undiluted Serum and Effects of Dilution. Glycoconj. J. 2012, 29 (2–3), 87–91.
(10) Bennett, N. R.; Jarvis, C. M.; Alam, M. M.; Zwick, D. B.; Olson, J. M.; Nguyen, H. V. T.; Johnson, J. A.; Cook, M. E.; Kiessling, L. L. Modular Polymer Antigens to Optimize Immunity. Biomacromolecules 2019, 20 (12), 4370–4379.
(11) Harris, S. L.; Craig, L.; Mehroke, J. S.; Rashed, M.; Zwick, M. B.; Kenar, K.; Toone, E. J.; Greenspan, N.; Auzanneau, F. I.; Marino-Albernas, J. R.; et al. Exploring the Basis of Peptide-Carbohydrate Crossreactivity: Evidence for Discrimination by Peptides between Closely Related Anti-Carbohydrate Antibodies. Proc. Natl. Acad. Sci. U. S. A. 1997, 94 (6), 2454–2459.
(12) Deng, S. J.; MacKenzie, C. R.; Sadowska, J.; Michniewicz, J.; Young, N. M.; Bundle, D. R.; Narang, S. A. Selection of Antibody Single-Chain Variable Fragments with Improved Carbohydrate Binding by Phage Display. J. Biol. Chem. 1994, 269 (13), 9533–9538. (13) Cygler, M.; Rose, D. R.; Bundle, D. R. Recognition of a Cell-Surface Oligosaccharide of
Pathogenic Salmonella by an Antibody Fab Fragment. Science (80-. ). 1991, 253 (5018), 442–445.
(14) Zhang, Y.; Campbell, C.; Li, Q.; Gildersleeve, J. C. Multidimensional Glycan Arrays for Enhanced Antibody Profiling. Mol. Biosyst. 2010, 6 (9), 1583–1591.
26
(15) Franceus, J.; Desmet, T. Sucrose Phosphorylase and Related Enzymes in Glycoside Hydrolase Family 13: Discovery, Application and Engineering. Int. J. Mol. Sci. 2020, 21 (7).
(16) Weis, W. I.; Drickamer, K. Structural Basis of Lectin-Carbohydrate Recognition. Annu. Rev. Biochem. 1996, 65, 441–473.
(17) Lairson, L. L.; Withers, S. G. Mechanistic Analogies amongst Carbohydrate Modifying Enzymes. Chem. Commun. 2004, No. 20, 2243–2248.
(18) Frandsen, K. E. H.; Simmons, T. J.; Dupree, P.; Poulsen, J. C. N.; Hemsworth, G. R.; Ciano, L.; Johnston, E. M.; Tovborg, M.; Johansen, K. S.; Von Freiesleben, P.; et al. The Molecular Basis of Polysaccharide Cleavage by Lytic Polysaccharide Monooxygenases. Nat. Chem. Biol. 2016, 12 (4), 298–303.
(19) Hettle, A.; Fillo, A.; Abe, K.; Massel, P.; Pluvinage, B.; Langelaan, D. N.; Smith, S. P.; Boraston, A. B. Properties of a Family 56 Carbohydrate-Binding Module and Its Role in the Recognition and Hydrolysis of β-1,3-Glucan. J. Biol. Chem. 2017, 292 (41), 16955– 16968.
(20) Carey, D. J. Syndecans: Multifunctional Cell-Surface Co-Receptors. Biochem. J. 1997, 327 (1), 1–16.
(21) Esko, J. D.; Prestegard, J. H.; Linhardt, R. J. Proteins That Bind Sulfated
Glycosaminoglycans. In Essentials of Glycobiology; Varki, A., Cummings, R. D., Esko, J. D., Stanley, P., Hart, G. W., Aebi, M., Darvill, A. G., Kinoshita, T., Packer, N. H.,
Prestegard, J. H., et al., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, 2015; pp 493–502.
(22) Capila, I.; Linhardt, R. J. Heparin - Protein Interactions. Angew. Chemie - Int. Ed. 2002, 41, 390–412.
(23) Andersson, E.; Rydengård, V.; Sonesson, A.; Mörgelin, M.; Björck, L.; Schmidtchen, A. Antimicrobial Activities of Heparin-Binding Peptides. Eur. J. Biochem. 2004, 271 (6), 1219–1226.
(24) Klim, J. R.; Li, L.; Wrighton, P. J.; Piekarczyk, M. S.; Kiessling, L. L. A Defined Glycosaminoglycan-Binding Substratum for Human Pluripotent Stem Cells. Nat. Methods 2010, 7 (12), 989–994.
(25) Showalter, A. M. Structure and Function of Plant Cell Wall Proteins. Plant Cell 1993, 5 (1), 9–23.
(26) Lis, H.; Sharon, N. Lectins: Carbohydrate-Specific Proteins That Mediate Cellular Recognition. Chem. Rev. 1998, 98 (2), 637–674.
(27) Polgár, J.; Clemetson, J. M.; Kehrel, B. E.; Wiedemann, M.; Magnenat, E. M.; Wells, T. N. C.; Clemetson, K. J. Platelet Activation and Signal Transduction by Convulxin, a C-Type Lectin from Crotalus Durissus Terrificus (Tropical Rattlesnake) Venom via the P62/GPVI Collagen Receptor. J. Biol. Chem. 1997, 272 (21), 13576–13583.
(28) MacKenzie, C. R.; Hirama, T.; Lee, K. K. Quantitative Analysis of Bacterial Toxin Affinity and Specificity for Glycolipid Receptors by Surface Plasmon Resonance. J. Biol. Chem. 1997, 272 (9), 5533–5538.
27
Xenopus Embryonic Epidermal Lectin Reveal a Conserved Mechanism of Microbial Glycan Recognition. J. Biol. Chem. 2016, 291 (11), 5596–5610.
(30) Taylor, M. E.; Drickamer, K.; Schnaar, R. L.; Etzler, M. E.; Varki, A. Discovery and Classification of Glycan-Binding Proteins. In Essentials of Glycobiology; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, 2017.
(31) Biedermann, F.; Schneider, H. J. Experimental Binding Energies in Supramolecular Complexes. Chem. Rev. 2016, 116 (9), 5216–5300.
(32) Davis, A. P.; Wareham, R. S. Carbohydrate Recognition through Noncovalent
Interactions: A Challenge for Biomimetic and Supramolecular Chemistry. Angew. Chemie - Int. Ed. 1999, 38 (20), 2978–2996.
(33) Boraston, A. B.; Nurizzo, D.; Notenboom, V.; Ducros, V.; Rose, D. R.; Kilburn, D. G.; Davies, G. J. Differential Oligosaccharide Recognition by Evolutionarily-Related β-1,4 and β-1,3 Glucan-Binding Modules. J. Mol. Biol. 2002, 319 (5), 1143–1156.
(34) Xie, H.; Bolam, D. N.; Nagy, T.; Szabó, L.; Cooper, A.; Simpson, P. J.; Lakey, J. H.; Williamson, M. P.; Gilbert, H. J. Role of Hydrogen Bonding in the Interaction between a Xylan Binding Module and Xylan. Biochemistry 2001, 40 (19), 5700–5707.
(35) Schwefel, D.; Maierhofer, C.; Beck, J. G.; Seeberger, S.; Diederichs, K.; Möller, H. M.; Welte, W.; Wittmann, V. Structural Basis of Multivalent Binding to Wheat Germ Agglutinin. J. Am. Chem. Soc. 2010, 132 (25), 8704–8719.
(36) Li, Z.; Lazaridis, T. The Effect of Water Displacement on Binding Thermodynamics: Concanavalin A. J. Phys. Chem. B 2005, 109 (1), 662–670.
(37) Botos, I.; Wlodawer, A. Proteins That Bind High-Mannose Sugars of the HIV Envelope; 2005; Vol. 88.
(38) Lee, R. T.; Hsu, T. L.; Huang, S. K.; Hsieh, S. L.; Wong, C. H.; Lee, Y. C. Survey of Immune-Related, Mannose/Fucose-Binding C-Type Lectin Receptors Reveals Widely Divergent Sugar-Binding Specificities. Glycobiology 2011, 21 (4), 512–520.
(39) Holla, A.; Skerra, A. Comparative Analysis Reveals Selective Recognition of Glycans by the Dendritic Cell Receptors DC-SIGN and Langerin. Protein Eng. Des. Sel. 2011, 24 (9), 659–669.
(40) Wesener, D. A.; Wangkanont, K.; McBride, R.; Song, X.; Kraft, M. B.; Hodges, H. L.; Zarling, L. C.; Splain, R. A.; Smith, D. F.; Cummings, R. D.; et al. Recognition of Microbial Glycans by Human Intelectin-1. Nat. Struct. Mol. Biol. 2015, 22 (8), 603–610.
(41) Sun, G.; Zhao, H.; Kalyanaraman, B.; Dahms, N. M. Identification of Residues Essential for Carbohydrate Recognition and Cation Dependence of the 4KDa Mannose 6-Phosphate Receptor. Glycobiology 2005, 15 (11), 1136–1149.
(42) Esquivies, L.; Blackler, A.; Peran, M.; Rodriguez-Esteban, C.; Izpisua Belmonte, J. C.; Booker, E.; Gray, P. C.; Ahn, C.; Kwiatkowski, W.; Choe, S. Designer Nodal/BMP2 Chimeras Mimic Nodal Signaling, Promote Chondrogenesis, and Reveal a BMP2-like Structure. J. Biol. Chem. 2014, 289 (3), 1788–1797.
(43) Hudson, K. L.; Bartlett, G. J.; Diehl, R. C.; Agirre, J.; Gallagher, T.; Kiessling, L. L.; Woolfson, D. N. Carbohydrate-Aromatic Interactions in Proteins. J. Am. Chem. Soc. 2015, 137 (48), 15152–15160.
28
(44) Bernardi, A.; Arosio, D.; Potenza, D.; Sánchez-Medina, I.; Mari, S.; Cañada, F. J.; Jiménez-Barbero, J. Intramolecular Carbohydrate-Aromatic Interaction and
Intermolecular van Der Waals Interactions Enhance the Molecular Recognition Ability of GM1 Glycomimetics for Cholera Toxin. Chem. - A Eur. J. 2004, 10 (18), 4395–4406. (45) Payne, C. M.; Bomble, Y. J.; Taylor, C. B.; McCabe, C.; Himmel, M. E.; Crowley, M. F.;
Beckham, G. T. Multiple Functions of Aromatic-Carbohydrate Interactions in a Processive Cellulase Examined with Molecular Simulation. J. Biol. Chem. 2011, 286 (47), 41028– 41035.
(46) Guce, A. I.; Clark, N. E.; Salgado, E. N.; Ivanen, D. R.; Kulminskaya, A. A.; Brumer, H.; Garman, S. C. Catalytic Mechanism of Human α-Galactosidase. J. Biol. Chem. 2010, 285 (6), 3625–3632.
(47) Taylor, R.; Kennard, O. Crystallographic Evidence for the Existence of C-H⋯O, C-H⋯N, and C-H⋯Cl Hydrogen Bonds. J. Am. Chem. Soc. 1982, 104 (19), 5063–5070.
(48) Pereira Silva, P. S.; Cardoso, C.; Ramos Silva, M.; Paixão, J. a.; Matos Beja, a.; Nogueira, F. Density Functional and X-Ray Diffraction Studies of Two Polymorphs of N,N′,N″-Triphenylguanidine. J. Mol. Struct. 2008, 888 (1–3), 92–98.
(49) Nishio, M.; Umezawa, Y.; Hirota, M.; Takeuchi, Y. The CH/π Interaction: Significance in Molecular Recognition. Tetrahedron 1995, 51 (32), 8665–8701.
(50) Samanta, U. .; Chakrabarti, P. .; Chandrasekhar, J. . Ab Initio Study of Energetics of X-H ‚‚‚ π ( X ) N , O , and C ) Interactions Involving a Heteroaromatic Ring. J. Phys. Chem. A 1998, 8964–8969.
(51) Oki, M.; Takano, S.; Toyota, S. Benzene-Ethene Interactions as Studied by Ab Initio Calculations. Bull. Chem. Soc. Jpn. 2000, 73, 2221–2230.
(52) Tsuzuki, S.; Honda, K.; Uchimaru, T.; Mikami, M.; Tanabe, K. The Interaction of Benzene with Chloro- and Fluoromethanes: Effects of Halogenation on CH/π Interaction. J. Phys. Chem. A 2002, 106 (17), 4423–4428.
(53) Ams, M. R.; Fields, M.; Grabnic, T.; Janesko, B. G.; Zeller, M.; Sheridan, R.; Shay, A. Unraveling the Role of Alkyl F on CH-π Interactions and Uncovering the Tipping Point for Fluorophobicity. J. Org. Chem. 2015, 80 (15), 7764–7769.
(54) Dey, R. C.; Seal, P.; Chakrabarti, S. CH/π Interaction in Benzene and Substituted Derivatives with Halomethane: A Combined Density Functional and Dispersion-Corrected Density Functional Study. J. Phys. Chem. A 2009, 113 (37), 10113–10118. (55) Ugozzoli, F.; Arduini, A.; Massera, C.; Pochini, A.; Secchi, A. CH/π Interaction between
Benzene and Model Neutral Organic Molecules Bearing Acid CH Groups. New J. Chem. 2002, 26 (12), 1718–1723.
(56) Sinnokrot, M. O.; Sherrill, C. D. Substituent Effects in π-π Interactions: Sandwich and t-Shaped Configurations. J. Am. Chem. Soc. 2004, 126 (24), 7690–7697.
(57) Nishio, M. The CH/π Hydrogen Bond in Chemistry. Conformation, Supramolecules, Optical Resolution and Interactions Involving Carbohydrates. Phys. Chem. Chem. Phys. 2011, 13 (31), 13873–13900.
(58) Lisbjerg, M.; Valkenier, H.; Jessen, B. M.; Al-Kerdi, H.; Davis, A. P.; Pittelkow, M. Biotin[6]Uril Esters: Chloride-Selective Transmembrane Anion Carriers Employing
C-29
H···anion Interactions. J. Am. Chem. Soc. 2015, 137 (15), 4948–4951.
(59) Jiménez-Moreno, E.; Jiménez-Osés, G.; Gómez, A. M.; Santana, A. G.; Corzana, F.; Bastida, A.; Jiménez-Barbero, J.; Asensio, J. L. A Thorough Experimental Study of CH/π Interactions in Water: Quantitative Structure-Stability Relationships for
Carbohydrate/Aromatic Complexes. Chem. Sci. 2015, 6 (11), 6076–6085. (60) Stanca-Kaposta, E. C.; Çarçabal, P.; Cocinero, E. J.; Hurtado, P.; Simons, J. P.
Carbohydrate-Aromatic Interactions: Vibrational Spectroscopy and Structural Assignment of Isolated Monosaccharide Complexes with p-Hydroxy Toluene and N-Acetyl L-Tyrosine Methylamide. J. Phys. Chem. B 2013, 117 (27), 8135–8142.
(61) Salonen, L. M.; Ellermann, M.; Diederich, F. Aromatic Rings in Chemical and Biological Recognition: Energetics and Structures. Angew. Chemie - Int. Ed. 2011, 50 (21), 4808– 4842.
(62) Zhao, C.; Li, P.; Smith, M. D.; Pellechia, P. J.; Shimizu, K. D. Experimental Study of the Cooperativity of CH-π Interactions. Org. Lett. 2014, 16 (13), 3520–3523.
(63) Baggioli, A.; Meille, S. V.; Raos, G.; Po, R.; Brinkmann, M.; Famulari, A. Intramolecular CH/π Interactions in Alkylaromatics: Monomer Conformations for
Poly(3-Alkylthiophene) Atomistic Models. Int. J. Quantum Chem. 2013, 113 (18), 2154–2162. (64) Krenske, E. H.; Houk, K. N. Aromatic Interactions as Control Elements in Stereoselective
Organic Reactions. Acc. Chem. Res. 2013, 46 (4), 979–989.
(65) Ninković, D. B.; Vojislavljević-Vasilev, D. Z.; Medaković, V. B.; Hall, M. B.; Brothers, E. N.; Zarić, S. D. Aliphatic-Aromatic Stacking Interactions in Cyclohexane-Benzene Are Stronger than Aromatic-Aromatic Interaction in the Benzene Dimer. Phys. Chem. Chem. Phys. 2016, 18 (37), 25791–25795.
(66) Chen, W.; Enck, S.; Price, J. L.; Powers, D. L.; Powers, E. T.; Wong, C.; Dyson, H. J.; Kelly, W. Structural and Energetic Basis of Carbohydrate−Aromatic Packing Interactions in Proteins. JACS 2013, 135, 9877–9884.
(67) Hsu, C. H.; Park, S.; Mortenson, D. E.; Foley, B. L.; Wang, X.; Woods, R. J.; Case, D. A.; Powers, E. T.; Wong, C. H.; Dyson, H. J.; et al. The Dependence of
Carbohydrate-Aromatic Interaction Strengths on the Structure of the Carbohydrate. J. Am. Chem. Soc. 2016, 138 (24), 7636–7648.
(68) Bautista-Ibañez, L.; Ramírez-Gualito, K.; Quiroz-García, B.; Rojas-Aguilar, A.; Cuevas, G. Calorimetric Measurement of the CH/Pi Interaction Involved in the Molecular
Recognition of Saccharides by Aromatic Compounds. J. Org. Chem. 2008, 73 (3), 849– 857.
(69) Asensio, J. L.; Arda, A.; Canada, F. J.; Jiménez-Barbero, J. Carbohydrate-Aromatic Interactions. Acc. Chem. Res. 2013, 46 (4), 946–954.
(70) Nishio, M.; Umezawa, Y.; Fantini, J.; Weiss, M. S.; Chakrabarti, P. CH-π Hydrogen Bonds in Biological Macromolecules. Phys. Chem. Chem. Phys. 2014, 16 (25), 12648–12683. (71) Tsuzuki, S.; Uchimaru, T.; Mikami, M. Magnitude and Nature of Carbohydrate-Aromatic
Interactions in Fucose-Phenol and Fucose-Indole Complexes: CCSD(T) Level Interaction Energy Calculations. J. Phys. Chem. A 2011, 115 (41), 11256–11262.
30
Interaction in Protein–Carbohydrate Binding: Bioinformatics and In Vitro Quantification. Chem. - A Eur. J. 2020, 26 (47), 10769–10780.
(73) Barondes, S. H. Stumbling on Galectins. In Galectins; 2008; pp 1–8.
(74) Johannes, L.; Jacob, R.; Leffler, H. Galectins at a Glance. J. Cell Sci. 2018, 131 (9), 1–9. (75) Cummings, R. D.; Liu, F.-T.; Vasta, G. R. Galectins. In Essentials of Glycobiology; Varki,
A., Cummings, R. D., Esko, J. D., Stanley, P., Hart, G. W., Aebi, M., Darvill, A. G.,
Kinoshita, T., Packer, N. H., Prestegard, J. H., et al., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, 2015; pp 469–480.
(76) Klyosov, A. a. Galectins and Their Functions in Plain Language. In Galectins; 2008; pp 9–31.
(77) Nesmelova, I. V.; Dings, R. P. M.; Mayo, K. H. Understanding Galectin Structure-Function Relationships to Design Effective Antagonists. In Galectins; Klyosov, A. a., Witczak, Z. J., Platt, D., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, 2008; pp 33–69. (78) Troncoso, M. F.; Ferragut, F.; Bacigalupo, M. L.; Cardenas Delgado, V. M.; Nugnes, L. G.;
Gentilini, L.; Laderach, D.; Wolfenstein-Todel, C.; Compagno, D.; Rabinovich, G. a.; et al. Galectin-8: A Matricellular Lectin with Key Roles in Angiogenesis. Glycobiology 2014, 24 (10), 907–914.
(79) Cagnoni, A. J.; Troncoso, M. F.; Rabinovich, G. A.; Mariño, K. V.; Elola, M. T. Full-Length Galectin-8 and Separate Carbohydrate Recognition Domains: The Whole Is Greater than the Sum of Its Parts? Biochem. Soc. Trans. 2020, 48 (3), 1255–1268.
(80) Pugliese, G.; Iacobini, C.; Pesce, C. M.; Menini, S. Galectin-3: An Emerging All-out Player in Metabolic Disorders and Their Complications. Glycobiology 2015, 25 (2), 136–150. (81) Diehl, C.; Genheden, S.; Modig, K.; Ryde, U.; Akke, M. Conformational Entropy Changes
upon Lactose Binding to the Carbohydrate Recognition Domain of Galectin-3. J. Biomol. NMR 2009, 45 (1–2), 157–169.
(82) Lepur, A.; Salomonsson, E.; Nilsson, U. J.; Leffler, H. Ligand Induced Galectin-3 Protein Self-Association. J. Biol. Chem. 2012, 287 (26), 21751–21756.
(83) Sato, S.; Rabinovich, G. a. Galectins As Danger Signals in Pathogen and Host-Tumor Interactions: New Members of the Growing Group of “Alarmins”? In Galectins; 2008; pp 115–146.
(84) Salomonsson, E.; Carlsson, M. C.; Osla, V.; Hendus-Altenburger, R.; Kahl-Knutson, B.; Oberg, C. T.; Sundin, A.; Nilsson, R.; Nordberg-Karlsson, E.; Nilsson, U. J.; et al.
Mutational Tuning of Galectin-3 Specificity and Biological Function. J. Biol. Chem. 2010, 285 (45), 35079–35091.
(85) Cumpstey, I.; Salomonsson, E.; Sundin, A.; Leffler, H.; Nilsson, U. J. Double Affinity Amplification of Galectin-Ligand Interactions through Arginine-Arene Interactions: Synthetic, Thermodynamic, and Computational Studies with Aromatic Diamido Thiodigalactosides. Chemistry 2008, 14 (14), 4233–4245.
(86) Bachhawat-Sikder, K.; Thomas, C. J.; Surolia, A. Thermodynamic Analysis of the Binding of Galactose and Poly-N-Acetyllactosamine Derivatives to Human Galectin-3. FEBS Lett. 2001, 500 (1–2), 75–79.
31
Elling, L.; Křen, V.; Bojarová, P. Chemo-Enzymatic Synthesis of LacdiNAc Dimers of Varying Length as Novel Galectin Ligands. J. Mol. Catal. B Enzym. 2014, 101, 47–55. (88) Gabius, H.-J.; Wu, A. M. Galectins As Regulators of Tumor Growth and Invasion by
Targeting Distinct Cell Surface Glycans and Implications for Drug Design. In Galectins; 2008; pp 71–85.
(89) Nesmelova, I. V.; Dings, R. P. M.; Mayo, K. H. Understanding Galectin Structure-Function Relationships to Design Effective Antagonists. In Galectins; 2008; pp 33–69. (90) Sörme, P.; Qian, Y.; Nyholm, P. G.; Leffler, H.; Nilsson, U. J. Low Micromolar Inhibitors
of Galectin-3 Based on 3′-Derivatization of N-Acetyllactosamine. ChemBioChem 2002, 3 (2–3), 183–189.
(91) Bum-Erdene, K.; Gagarinov, I. a; Collins, P. M.; Winger, M.; Pearson, A. G.; Wilson, J. C.; Leffler, H.; Nilsson, U. J.; Grice, I. D.; Blanchard, H. Investigation into the Feasibility of Thioditaloside as a Novel Scaffold for Galectin-3-Specific Inhibitors. Chembiochem 2013, 14 (11), 1331–1342.
(92) Lundquist, J. J.; Kiburz, B. M.; Wu, J. K.; Gibbs, K. D.; Toone, E. J. Towards High Affinity Carbohydrate-Binding Proteins: Directed Evolution of Murine Galectin-3. Can. J. Chem. 2002, 80 (8), 999–1009.
(93) Salameh, B. a.; Cumpstey, I.; Sundin, A.; Leffler, H.; Nilsson, U. J. 1H-1,2,3-Triazol-1-Yl Thiodigalactoside Derivatives as High Affinity Galectin-3 Inhibitors. Bioorganic Med. Chem. 2010, 18 (14), 5367–5378.
(94) Funasaka, T.; Raz, a.; Nangia-Makker, P. Galectin-3 in Angiogenesis and Metastasis. Glycobiology 2014, 24 (10), 886–891.
(95) Saravanan, C.; Liu, F. T.; Gipson, I. K.; Panjwani, N. Galectin-3 Promotes Lamellipodia Formation in Epithelial Cells by Interacting with Complex N-Glycans on Α3β1 Integrin. J. Cell Sci. 2009, 122 (20), 3684–3693.
(96) Friedrichs, J.; Torkko, J. M.; Helenius, J.; Teräväinen, T. P.; Füllekrug, J.; Muller, D. J.; Simons, K.; Manninen, A. Contributions of Galectin-3 and -9 to Epithelial Cell Adhesion Analyzed by Single Cell Force Spectroscopy. J. Biol. Chem. 2007, 282 (40), 29375– 29383.
(97) Woo, H. J.; Lotz, M. M.; Jung, J. U.; Mercurio, A. M. Carbohydrate-Binding Protein 35 (Mac-2), a Laminin-Binding Lectin, Forms Functional Dimers Using Cysteine 186. J. Biol. Chem. 1991, 266 (28), 18419–18422.
(98) Hikita, C.; Vijayakumar, S.; Takito, J.; Erdjument-Bromage, H.; Tempst, P.; Al-Awqati, Q. Induction of Terminal Differentiation in Epithelial Cells Requires Polymerization of Hensin by Galectin 3. J. Cell Biol. 2000, 151 (6), 1235–1246.
(99) Lagana, A.; Goetz, J. G.; Cheung, P.; Raz, A.; Dennis, J. W.; Nabi, I. R. Galectin Binding to Mgat5-Modified N-Glycans Regulates Fibronectin Matrix Remodeling in Tumor Cells. Mol. Cell. Biol. 2006, 26 (8), 3181–3193.
(100) Carlsson, M. C.; Bengtson, P.; Cucak, H.; Leffler, H. Galectin-3 Guides Intracellular Trafficking of Some Human Serotransferrin Glycoforms. J. Biol. Chem. 2013, 288 (39), 28398–28408.
32
Galectin-3 Induces Endothelial Cell Morphogenesis and Angiogenesis. Am. J. Pathol. 2000, 156 (3), 899–909.
(102) Yu, L. G.; Andrews, N.; Zhao, Q.; McKean, D.; Williams, J. F.; Connor, L. J.;
Gerasimenko, O. V.; Hilkens, J.; Hirabayashi, J.; Kasai, K.; et al. Galectin-3 Interaction with Thomsen-Friedenreich Disaccharide on Cancer-Associated MUC1 Causes Increased Cancer Cell Endothelial Adhesion. J. Biol. Chem. 2007, 282 (1), 773–781.
(103) Chen, C.; Duckworth, C. A.; Zhao, Q.; Pritchard, D. M.; Rhodes, J. M.; Yu, L. G. Increased Circulation of Galectin-3 in Cancer Induces Secretion of Metastasis-Promoting Cytokines from Blood Vascular Endothelium. Clin. Cancer Res. 2013, 19 (7), 1693–1704.
(104) Barrow, H.; Guo, X.; Wandall, H. H.; Pedersen, J. W.; Fu, B.; Zhao, Q.; Chen, C.; Rhodes, J. M.; Yu, L. G. Serum Galectin-2, -4, and -8 Are Greatly Increased in Colon and Breast Cancer Patients and Promote Cancer Cell Adhesion to Blood Vascular Endothelium. Clin. Cancer Res. 2011, 17 (22), 7035–7046.
(105) Thiemann, S.; Baum, L. G. Galectins and Immune Responses-Just How Do They Do Those Things They Do? Annu. Rev. Immunol. 2016, 34, 243–264.
(106) Rabinovich, G. A.; Toscano, M. A. Turning “sweet” on Immunity: Galectin-Glycan Interactions in Immune Tolerance and Inflammation. Nat. Rev. Immunol. 2009, 9 (5), 338–352.
(107) Demotte, N.; Stroobant, V.; Courtoy, P. J.; Van Der Smissen, P.; Colau, D.; Luescher, I. F.; Hivroz, C.; Nicaise, J.; Squifflet, J. L.; Mourad, M.; et al. Restoring the Association of the T Cell Receptor with CD8 Reverses Anergy in Human Tumor-Infiltrating Lymphocytes. Immunity 2008, 28 (3), 414–424.
(108) Stillman, B. N.; Hsu, D. K.; Pang, M.; Brewer, C. F.; Johnson, P.; Liu, F.-T.; Baum, L. G. Galectin-3 and Galectin-1 Bind Distinct Cell Surface Glycoprotein Receptors to Induce T Cell Death. J. Immunol. 2006, 176 (2), 778–789.
(109) Matarrese, P.; Tinari, N.; Semeraro, M. L.; Natoli, C.; Iacobelli, S.; Malorni, W. Galectin-3 Overexpression Protects from Cell Damage and Death by Influencing Mitochondrial Homeostasis. FEBS Lett. 2000, 473 (3), 311–315.
(110) Jeon, S.-B.; Yoon, H. J.; Chang, C. Y.; Koh, H. S.; Jeon, S.-H.; Park, E. J. Galectin-3 Exerts Cytokine-Like Regulatory Actions through the JAK–STAT Pathway. J. Immunol. 2010, 185 (11), 7037–7046.
(111) Sato, S.; Nieminen, J. Seeing Strangers or Announcing “Danger”: Galectin-3 in Two Models of Innate Immunity. Glycoconj. J. 2004, 19 (7–9), 583–591.
(112) Varki, A. Letter to the Glyco-Forum: Since There Are PAMPs and DAMPs, There Must Be SAMPs? Glycan “Self-Associated Molecular Patterns” Dampen Innate Immunity, but Pathogens Can Mimic Them. Glycobiology 2011, 21 (9), 1121–1124.
(113) Baum, L. G.; Garner, O. B.; Schaefer, K.; Lee, B. Microbe-Host Interactions Are Positively and Negatively Regulated by Galectin-Glycan Interactions. Front. Immunol. 2014, 5 (JUN), 1–8.
(114) Mandrell, R. N.; Apicella, M. A.; Lindstedt, R.; Leffler, H. Possible Interaction between Animal Lectins and Bacterial Carbohydrates. In Methods in enzymology; 1994; Vol. 236, pp 231–254.