HAL Id: hal-02566353
https://hal.archives-ouvertes.fr/hal-02566353
Submitted on 7 May 2020HAL is a multi-disciplinary open access
archive for the deposit and dissemination of sci-entific research documents, whether they are pub-lished or not. The documents may come from teaching and research institutions in France or abroad, or from public or private research centers.
L’archive ouverte pluridisciplinaire HAL, est destinée au dépôt et à la diffusion de documents scientifiques de niveau recherche, publiés ou non, émanant des établissements d’enseignement et de recherche français ou étrangers, des laboratoires publics ou privés.
Role of the Arginine 214 in the substrate specificity of
OXA-48
Saoussen Oueslati, Pascal Retailleau, Ludovic Marchini, Camille Berthault,
Laurent Dortet, Rémy A Bonnin, Bogdan Iorga, Thierry Naas
To cite this version:
Saoussen Oueslati, Pascal Retailleau, Ludovic Marchini, Camille Berthault, Laurent Dortet, et al.. Role of the Arginine 214 in the substrate specificity of OXA-48. Antimicrobial Agents and Chemother-apy, American Society for Microbiology, 2020, 64 (5), pp.e02329-19. �10.1128/aac.02329-19�. �hal-02566353�
Revised AAC02329-19 1
Role of the Arginine 214 in the substrate specificity of OXA-48 2
3
Saoussen OUESLATI1, Pascal RETAILLEAU2, Ludovic MARCHINI,2 Camille 4
BERTHAULT1, Laurent DORTET1,3,4, Rémy A. BONNIN1,3,Bogdan I. IORGA2, and 5
Thierry NAAS1,3,4* 6
7
1EA7361 “Structure, dynamic, function and expression of broad spectrum -lactamases”,
8
UMR1184, Faculty of Medicine, LabEx LERMIT, Université Paris-Saclay, Le
Kremlin-9
Bicêtre, France
10
2
Institut de Chimie des Substances Naturelles, CNRS UPR 2301, Labex LERMIT, Université
11
Paris-Saclay, Gif-sur-Yvette, France.
12
3
Associated French National Reference Center for Antibiotic Resistance:
Carbapenemase-13
producing Enterobacteriaceae, Le Kremlin-Bicêtre, France
14
4
Bacteriology-Hygiene unit, Assistance Publique/Hôpitaux de Paris, Bicêtre Hospital, Le
15
Kremlin-Bicêtre, France
16
Running title : Arginine 214 and substrate specificity of OXA-48 17
Keywords: oxacillinase, carbapenemase, OXA-232, antibiotic resistance, beta-lactamase, 18
OXA-48-like 19
Word count: Abstract 239; Text: 3996 20
Figures: 2; Tables: 4 21
22
*Corresponding author: Service de Bactériologie-Hygiène, Hôpital Bicêtre, 78 rue du Général 23
Leclerc, 94270 Le Kremlin-Bicêtre, France. Tel : +33 1 45 21 20 19. Fax : +33 1 45 21 63 40. 24
E-mail: thierry.naas@aphp.fr 25
Abstract 27
Increasing number of variants of the carbapenem-hydrolyzing class D -lactamase 28
OXA-48 are identified in Enterobacterales worldwide. Among them, OXA-181 and OXA-232 29
are of particular interest, as they differ from each other by a single amino-acid (AA) 30
substitution at position 214 (R in OXA-181, and S in OXA-232), that results in reduced 31
carbapenem-hydrolyzing activity for OXA-232. To investigate the role of the AA214, the X-32
ray structure of OXA-232 was determined and the AA214 of OXA-48 and of OXA-232 was 33
replaced by G, L, D, E, S, R, and K using site-directed mutagenesis. These mutants were 34
phenotypically characterized, and three mutants of OXA-232 were purified to study their 35
steady-state kinetic properties. X-ray structure of OXA-232 along with molecular modelling 36
studies showed that the interaction via a salt bridge between R214 and D159 in OXA-48 is 37
not possible with G214 or S214 mutations. In contrast, with K214 that is also positively 38
charged, the interaction with D159 is maintained. With the E214 mutant an alternative 39
binding conformation of imipenem was evidenced that is not compatible with a nucleophilic 40
attack by S70. Thus, imipenem has very poor apparent affinity for the E214 mutant because 41
of its non-productive binding mode. Similarly, we could explain the lack of temocillin 42
hydrolysis by OXA-232, which is due to the unfavorable interaction between the negatively 43
charged R1 substituent of temocillin with the S214 residue. 44
Overall, we demonstrate that the AA214 in OXA-48-like β-lactamases is critical for 45
the carbapenemase activity. 46
47 48
Introduction 49
The intensive use of antibiotics to treat infections led to the emergence of multidrug 50
resistant pathogens especially in Gram-negative bacteria (GNB). Among -lactams, 51
carbapenems are considered as last resort antibiotics to treat severe infections caused by GNB 52
(1). The major mechanisms of carbapenem resistance in Enterobacterales are (i) the 53
association of a ß-lactamase that very weakly hydrolyze carbapenems (ESBL or a 54
cephalosporinase) with decreased outer-membrane permeability or (ii) inactivation by specific 55
carbapenem-hydrolyzing -lactamases, carbapenemases (1). Genes encoding for these 56
enzymes are mostly harbored by plasmids explaining their rapid spread. -Lactamases are 57
classified into 4 groups (Amber’s classes A to D) based upon sequence homology (1,2). The 58
clinically-relevant carbapenemases in Enterobacterales belong to class A (KPC-type), B 59
(NDM, VIM, IMP) and D (OXA-48-like). OXA-48, initially identified in Turkey (3), has 60
since rapidly spread in the Mediterranean area, middle East, Europe, India and is now turning 61
into a major global threat (4). OXA-48 hydrolyzes penicillins including temocillin, narrow-62
spectrum cephalosporins, and also carbapenems at low rate, but spares expanded-spectrum 63
cephalosporins (ESC) e.g. ceftazidime and cefepime (5). Along with the spread of OXA-48, 64
several variants have been reported that differ from OXA-48 by amino-acid (AA) 65
substitutions or deletions (http://bldb.eu/BLDB.php?class=D#OXA) mostly located in the 5-66
6 loop (6). OXA-181 that differs from OXA-48 by 4 amino-acid substitutions (T103A,
67
N110D, E169Q and S171A) is the second most prevalent OXA-48 variant. OXA-232 that
68
differs from OXA-181 by an additional substitution R214S in the 5-6 loop, is particularly
69
interesting, as its carbapenem-hydrolyzing activity was significantly impaired, as compared to
70
OXA-181 or OXA-48 (7). These results suggested a pivotal role of R214 in the hydrolysis of
71
carbapenems. To further investigate the role of R214 in the hydrolytic profile of OXA-48-like
carbapenemases, the X-ray structure of OXA-232, and the steady-state kinetic parameters of
73
in-vitro generated OXA-232 mutants at AA position 214 were determined.
74 75
Results 76
Susceptibility testing. To determine the effect of the substitutions at the AA position
77
214 on the hydrolysis of imipenem and temocillin, MIC values of E. coli expressing OXA-48
78
and OXA-232 and their respective point mutant derivatives were determined.
79
The substitution R214S in OXA-48 led to a phenotype similar to that of OXA-232,
80
e.g. reduced MICs for temocillin and imipenem (Table 1). Conversely, OXA-232-S214R re-81
stored MICs for temocillin and imipenem similarly to OXA-48/181. When AA214 is
82
substituted with an uncharged AA, such as G and L, the MIC values were similar to those of
83
the OXA-232. The most interesting results were obtained with substitutions by negatively
84
charged AA at pH 7.0 such as aspartic acid and glutamic acid. Indeed, MIC values for
85
imipenem and temocillin were remarkably affected, similar to E. coli Top10 control isolate.
86
In a second step, we focused on the analysis of mutants of OXA-232:
OXA-232-87
S214K; OXA-232-S214E and OXA-232-S214G, which are representative for each amino
88
acid group: polar positively charged (K), polar negatively charged (E) and uncharged (G).
89
MIC values for other -lactams were performed and compared to those of 232,
OXA-90
181 (OXA-232-S214R) and OXA-48 (Table 2). Overall, MIC values for benzylpenicillin and
91
cephalothin were not affected and MIC values of the mutants for the other carbapenems
92
(meropenem and ertapenem) vary in the same way as for imipenem.
93 94
Biochemical properties determination. To further characterize the impact of the 95
nature of the residue at position 214 on the hydrolytic profile, steady-state kinetic parameters 96
of the 3 OXA-232 mutants (OXA-232-S214G; OXA-232-S214E and OXA-232-S214K) were 97
determined and compared to those of OXA-48, OXA-181 and OXA-232 (5,7). Overall, the 98
kinetic studies revealed three patterns (Table 3). OXA-232-S214G exhibited hydrolytic 99
activity towards all tested -lactams similarly to those of OXA-232. The mutant OXA-232-100
S214K possessed a higher catalytic efficiency (kcat / Km= 83 mM-1.s-1) for imipenem of 4-101
fold as compared to OXA-232 (kcat / Km= 23 mM-1.s-1), due to a weak increase of the turnover 102
of the enzyme (kcat). The most interesting result was observed with the OXA-232-S214E 103
mutant. Indeed, the substitution by a negatively charged amino acid led to a drastic increase 104
of the Km (>2000 µM) of at least 200-folds as compared to OXA-232 (Km= 9 µM), thus 105
decreasing the catalytic efficiency. Moreover, this mutant totally lost its hydrolytic properties 106
for temocillin. All these results were concordant with the observed MIC values. Taken 107
together, our results confirm that the hydrolysis of imipenem depends on the nature of the 108
residue at position 214. It appears in light of these results that positively charged amino-acids 109
favor hydrolysis of imipenem. 110
111
Crystal Structure of OXA-232. The crystals of OXA-232 were obtained at basic pH 112
in 30% w/v PEG3000 and in the presence of 0.2 M Li2SO4 (Figure 2 A). Glycerol was added 113
to the mother liquor to cryoprotect the crystals before flash-annealing for data collection. 114
They belonged to hexagonal space group, P 62, with 43.8 % solvent as calculated from the 115
Matthews coefficient of 2.19 Å3 Da-1 (8,5). At that time, the best available OXA-48 model in 116
the PDB was 3HBR (1.9Å) (5) obtained from crystals grown at lower pH (7.5 vs 8.5), in PEG 117
4000 then dipped into cryoprotectant ethyleneglycol, and which belonged to the monoclinic 118
space group P 21 with 48.3% solvent (Matthews coefficient, 2.38 Å3 Da-1). After deletion of 119
22 N-ter amino acids, one half of R214S-3HBR coordinates were used as quasi-homologous 120
model for phasing the hexagonal crystal to 2.2 Å resolution. The refined electron density 121
appeared neat throughout the entire backbone and most of side chains of the biological dimer 122
per asymmetric unit of the crystal. Alternate positions for Q41 and Q53 were observed as well 123
for D230, whose one position permitted a salt bridge with R107 at the dimer interface. The 124
average Real-space correlation factor calculated by Sfcheck software (10) is 0.932. Compared 125
to 3HBR, up to three additional C-ter amino acids could be displayed in chain B of OXA-232 126
(only one in chain A) but not the 9-His-tag chain, disordered in the surrounding solvent. 127
Additionally, some minor discontinuities observed in some parts of 3HBR main chains could 128
be fixed in 5HFO (N50D/K51D, T99C, S150A h-bonding D148A). The rmsd values with the 129
two 3HBR dimers range between 0.3452 and 0.4197 Å (Superpose, (11)). Obviously, the 130
OXA-232 tertiary structure of each monomer features the usual class D β-lactamase fold, with 131
an α-helical region (α3: 73–82; α4: 110-115; α5: 120–130; α6: 132–142; α9: 185–194) and a 132
mixed α-helix/β-sheet region (β1: 26–28; α1: 31–35; β2: 42–48; β3: 53–56; α2: 59–62; β4: 133
196–199; β5: 204–212; β6: 219–227; β7: 232–240). 98.2% of all residues were inside the 134
favored regions of the Ramachandran plot, and 1.8% in the allowed regions. The quaternary 135
structure of OXA-232 is dimeric as observed for OXA-48 around a non-crystallographic two-136
fold axis, in compliance with our data of size-exclusion chromatography. It was already 137
shown that less monomer surface (200Å2) was buried in that dimer formation compared to 138
OXA-10 and it was built upon an intermolecular β-sheet involving β4 from each subunit 139
linked by reciprocal H-bonds between the A199 NH and the A199 carbonyl O of one chain 140
with the corresponding one related by the ncs two-fold axis. Several other H-bonds and salt 141
bridges also participated in the dimeric interface stabilization, and among these interactions, 142
an anionic binding site lying on the ncs two-fold axis is tweezed by two facing R206 from ncs 143
related β5 sheets. This contrasts with the cation-binding site described at the interface of 144
OXA-10, involving other residues replaced in OXA-48. In the original 3HBR structure, a 145
water molecule was placed but on the faith of spherical electron density shape, low B-factor 146
and recurrent presence of halogens in other class D beta-lactamase structures (but not upon 147
anomalous signal), it was substituted by a residual chloride coming from previous purification 148
steps. The same misinterpretation was also done in in the structure 4S2K (12), where chlorine 149
was also reported in the crystallizing medium composition. If sulfate ions have also been seen 150
making that salt bridge in class D structures grown in the presence of large amount of sulfate 151
salts (e.g. structures 6NLW and 5FDH), here six such anions have been spotted essentially at 152
the molecule surface as well as five opportune glycerol molecules. Carbamylated lysines 153
(KCX 73) can be described in a similar environment in both structures. Regarding the 154
mutation of R214S (the R214 residue was shown in water-mediated interaction with the 155
avibactam drug in the active site in the structure 4S2K (12), this has not structurally disturbed 156
the β5-β6 hairpin and its direct environment, just adding few solvent molecules in the created 157
void (2-3 waters, and one glycerol molecule in chain A) and modifying the L158B and I215B 158
side chain conformers. Deeper into both active sites, respective carbamylated lysine (KCX 159
73) can be described in a similar environment in both structures OXA-232 and OXA-48. We 160
note however the presence of a sulfate ion in interaction with the same residues that recognize 161
the sulfate moiety of the avibactam (Figure 2B). More recently the structure of another 162
variant, OXA-181; which differs from OXA-232 by five substitutions T103A, N110D,
163
E169Q, S171A and R214S, has been released (PDB code 5OE0 (13)) at a similar resolution
164
(2.05 Å). The rmsd with OXA-232 is even lower than the former overlay (0.224 and 0.252Å)
165
in relation to the fact that structure crystallized in the same unit cell dimensions and space
166
group P62 than OXA-232. However, five residue mutations are on the protein surface that
167
could potentially modulate the contacts leading to the crystal growth and crystallization
168
conditions were slightly different with a neutral pH and PEG-mme 5000 instead of PEG 3000
169
or PEG 4000 for 3HBR. Like OXA-232, sulfate has been used in low quantity in the
170
crystallization drop and one anion was reported in the final 5HFO model bound to the amide
171
N of R186A as for OXA-232. Variability in β-lactamase crystal forms remains tenuous to
explain or even predict, since crystals of OXA-48 in complex with inhibitors grew in the
173
same unit cell dimensions (the choice of P32 space group instead of P62 could be interpreted
174
by the author’s intention to refine two more monomers and active sites independently) in
175
different crystallizing solution conditions (MPD was the major component) and in absence of
176
sulfate.
177 178
Molecular modelling. An in-silico study was performed to identify the structural
179
determinants that could explain the experimentally determined differences between the
180
hydrolytic profiles of OXA-232 and its variants in comparison with OXA-48. The OXA-232
181
structure was used as a starting point and the mutations S214G, S214K and S214E were
182
modeled based on predicted low energy conformations (14). The resulting models showed
183
that lateral chains of serine and lysine mutants are positioned in the same axis and direction as
184
the lateral chain of arginine, without any clashes. The glutamate mutant would be positioned
185
differently, oriented towards the active site, establishing hydrogen bonds with the backbone
186
nitrogen and with the sidechain hydroxyl of T213. This is the only conformation that avoids
187
clashes of this glutamate mutant with the neighboring residues I215 and D159, the latter being
188
involved in OXA-48 in a salt bridge with R214 that maintains a closed conformation of the
189
active site, which would stabilize the substrates (including carbapenems) in a conformation
190
compatible with an efficient hydrolysis, thus influencing the carbapenem turnover (5). For
191
OXA-232 and its variants the salt bridge R214-D159 cannot be formed, and this may explain
192
the decrease of kcat values compared to OXA-48.
193
Molecular docking calculations of imipenem and temocillin on the R214E mutant of
194
OXA-48 provided explanations for the different hydrolytic parameters that were observed
195
experimentally. Imipenem showed two alternative binding modes, one that was known, with a
196
single ionic interaction between the imipenem carboxylate and the sidechain of R250,
compatible with a nucleophilic attack by S70 (Figure 2C), and a second one, presumably
198
more stable, with two ionic interactions, one between the imipenem carboxylate and the
199
sidechain of R250 as in the previous binding mode, and a second between the positively
200
charged R2 substituent of imipenem and the mutated residue E214 (Figure 2C). This latter
201
binding mode is not productive, as it is not compatible with a nucleophilic attack by S70, thus
202
increasing the apparent Km value. To confirm this hypothesis, we have determined the IC50 for
203
imipenem with OXA-232 and with the mutant OXA-232 S214E, the only variant with an
204
increased Km value. Interestingly, the values obtained (2 µM and 0.059 µM, respectively)
205
showed that the mutant OXA-232 S214E has an IC50 ~34-fold lower as compared to that of
206
OXA-232. On the other hand, temocillin is the only β-lactam tested possessing a negative
207
charge on the R1 substituent. Our docking calculations showed that this negative charge
208
would establish strong unfavorable ionic interactions with the mutated residue E214, thus
209
precluding the binding in a conformation compatible with a nucleophilic attack by S70
210 (Figure 2D). 211 212 Discussion 213
OXA-48-producing Enterobacterales are now endemic in many countries, such as 214
Turkey, Middle East, North Africa, India and have widely spread across Europe. Since the 215
first description of OXA-48, several variants have been described (1,6). These variants can be 216
classified into 3 groups according to their hydrolysis profile. Most of them, including OXA-217
181 or OXA-162 (7), have an enzymatic activity similar to OXA-48 (5). The second group, 218
represented by OXA-163 (7), OXA-247 (15) and OXA-405 (16), have no carbapenemase 219
activity, but instead a marked hydrolytic activity against ESC, similarly to OXA-ESBLs (17). 220
Finally, the third group represented by OXA-244 (18) and OXA-232 (7) exhibit an overall 221
reduced activity towards all ß-lactams including carbapenems as compared to OXA-48. The 222
amino acid sequence comparison of OXA-48-variants suggested a link between the primary 223
structure and the function of these enzymes. Indeed, all 48-variants with an OXA-224
ESBLs phenotype (loss of carbapenem hydrolysis and the gain of activity towards ESC) have 225
amino-acid deletions in the 5-6 loop (Figure 1). This observation suggests that this loop 226
plays a role in the substrate specificity. The phenotypic study of OXA-244 (which differs 227
from OXA-48 by only one substitution R214G) (18) and the enzymatic study of OXA-232 228
(which differs from OXA-181 by a single substitution R214S) underline that the residue 214 229
is crucial for the carbapenem and temocillin hydrolysis. To confirm this hypothesis, we 230
generated mutants of OXA-48 and OXA-232 at the position 214, and analyzed their 231
hydrolytic profiles including OXA-232-S214R and OXA-48-R214S. Thus, the substitution of 232
S214R for OXA-232 re-stored hydrolytic properties toward temocillin and imipenem. 233
Similarly, the substitution of R214S for OXA-48, led to a drastic decrease of imipenem and 234
temocillin hydrolysis. In order to better understand this phenomenon, 5 mutants at position 235
214 of OXA-48 and OXA-232 were generated. These substitutions corresponded to amino 236
acid that were: polar positively charged (214K), polar negatively charged (214E and 214D), 237
non-polar (214L) and glycine (214G). Overall, the substitutions with D, E, L and G led to a 238
decreased MICs of imipenem and temocillin. The substitution S214K in OXA-232 led to 239
increased MIC values, but still lower than those of 48. The biochemical study of OXA-240
232-S214G and OXA-232-S214K showed that these 2 substitutions did not affect the 241
apparent affinity toward -lactams (Km values were similar to those of 232 and OXA-242
48) but have an effect on the acylation or deacylation steps of the substrate catalysis (kcat 243
values were similar to those of OXA-232 but smaller than those of OXA-48). The catalytic 244
efficiencies (kcat/Km) of these two mutants were in agreement with the MIC values. The most 245
significant differences were observed with OXA-232-S214E, with a drastic decrease in the 246
apparent affinity for imipenem (Km was at least 200 fold higher as compared to OXA-232 247
and OXA-48) and a loss of temocillin hydrolysis. The glutamate substitution seems to have a 248
direct effect on the apparent affinity of imipenem. Analysis of the 3D structure of OXA-48 249
showed that R214 interacts with D159 via a salt bridge (5), which maintains the shape and the 250
network of water molecules within the binding site. Our molecular modeling study revealed 251
that with the G214 or S214 mutations, this interaction with D159 is lacking, which 252
presumably increases the flexibility of this part of the binding site. In contrast, K is a 253
positively charged polar amino-acid, which appears to maintain the interaction with D159, 254
even with a lateral chain shorter than R. In the mutant E214, we evidenced an alternative 255
binding conformation of imipenem that is not compatible with a nucleophilic attack by S70. 256
In this case, imipenem acts as an inhibitor with an IC50 value of 0.059µM, which explains the 257
significantly higher Km values determined experimentally for this substrate. We also showed 258
that, in the same E214 mutant, the unfavorable interaction between the negatively charged R1 259
substituent of temocillin with the E214 residue precludes the binding of this antibiotic, thus 260
explaining the structural basis responsible for the lack of temocillin hydrolysis observed 261
experimentally. 262
Overall, we demonstrated that the AA position 214 in OXA-48-like β-lactamases is 263
critical for the carbapenemase activity but also for the temocillin hydrolysis. This point is of 264
outmost clinical importance and explains why detection of OXA-244 or OXA-232-producers 265
remains a challenge for clinical microbiology laboratories (18-20). Indeed, these isolates do 266
not grow on ChromID Carba Smart (bioMérieux, Marcy L’Etoile, France), one of the most 267
used type of medium for the screening of CPEs (19). The ChromID Carba Smart is a biplate 268
containing on one side a carbapenem and on the other temocillin, two substrates that are only 269
weakly hydrolyzed by OXA-244 and OXA-232 (19). 270
271
Materials and methods 272
Bacterial strains. The clinical strain Escherichia coli LIEU (20) expressing the OXA-232
-273
lactamase was used to clone the blaOXA-232 gene. E. coli TOP10 (Invitrogen, Saint-Aubin,
274
France) was used for cloning and mutagenesis experiments and E. coli BL21 DE3 was used
275
for overexpression experiments (Novagen, Fontenay-sous-Bois, France).
276 277
Antimicrobial agents, susceptibility testing and microbiological techniques. 278
Antimicrobial susceptibilities were determined by the disc diffusion technique on Mueller–
279
Hinton agar (Bio-Rad, Marnes-La-Coquette, France) and interpreted according to the
280
EUCAST breakpoints, updated in May 2018 (http://www.eucast.org). MICs were determined
281
using the Etest technique (bioMérieux). Antibiotics were purchased from Sigma (Saint-282
Quentin-Fallavier, France) except temocillin (Eumedica, Brussels, Belgium) 283
284
PCR, cloning, site-directed mutagenesis and DNA sequencing. The recombinant plasmids 285
pTOPO-blaOXA-232 and pTOPO-blaOXA-48, obtained from a previous study (7), were used as a
286
template for the site-directed mutagenesis assays and specific primers were designed for the
287
different mutations, using the program QuickChange Primer Design (Agilent Technologies).
288
QuikChange II Site-Directed Mutagenesis Kit (Agilent Technologies) was used, following the
289
manufacturer’s recommendations, in order to substitute the AA214 into a Glycine (G), a
290
Lysine (K), a Leucine (L), an Aspartic acid (D), a Glutamic acid (E), a Serine (S) and an
291
Arginine (R). Mutagenesis reaction products were transformed in E. coli TOP10 (Invitrogen,
292
Saint-Aubin, France) and selection was performed on TSA plate containing Kanamycin
293
(50g/ml). For the production of OXA-232, genes were amplified by PCR using the forward
294
primer OXA23-256NdeI (5’-aaaaaCATATGaaggaatggcaagaaaacaaa-3’) and the reverse primer
295
OXAXhoI-stop (5’-aaaaaCTCGAGggggaataatttttcctgtttgag-’3) to increase the purification yield.
296
For the mutants OXA-232 S214E, OXA-232 S214G and OXA-232 S214K the forward
primers OXANdeI (5′-aaaaaCATATGTTGGTGGCATCGATTATCGG-3′) and reverse primer
298
OXAXhoI-stop (5-aaaaaCTCGAGGAGCACTTCTTTTGTGATGGC-3’) were used.
299
Recombinant plasmids were cloned into pET41b (for OXA-232 wt) allowing the expression
300
of the enzyme with an His-tag and pET9a (for OXA-232 mutants) vector (Invitrogen, Life
301
Technologies, Cergy-Pontoise, France), then transformed into E. coli BL21 DE3 (Novagen).
302
All the recombinant plasmids were extracted using the Qiagen miniprep Kit and sequenced
303
using a T7 promoter and M13 reverse primers or T7 terminator (depending on the plasmid)
304
with an automated sequencer (ABI Prism3100; Applied Biosystems). The nucleotide
305
sequences were analyzed using software available at the National Center for Biotechnology
306
Information website (http://www.ncbi.nlm.nih.gov).
307 308
Protein purification. Overnight culture of E. coli BL21 DE3 harboring recombinant 309
plasmids pET41b-OXA-232 or pET9a-OXA-232-mut were used to inoculate 2 L of LB
310
medium broth containing 50 mg/L kanamycin. Bacteria were cultured at 37°C until an OD of
311
0.6 at 600 nm was reached. The -lactamase was induced overnight with 0.2 mM IPTG as
312
inducer and cultures were centrifuged at 6000 g for 15min. The pellets were resuspended with
313
the binding buffer 25mM phosphate sodium pH 7.4, 300 mM K2SO4, 10 mM imidazole for
314
OXA-232 wt and with the buffer 20 mM Bis-Tris H2SO4 (pH 7.2) for mutants of OXA-232.
315
Bacterial cells were disrupted by sonication and the bacterial pellet was removed by two
316
consecutive centrifugation steps at 10 000 g for 1h at 4°C; the supernatant was then
317
centrifuged at 96 000 g for 1h at 4°C. OXA-232 wt was purified with a NTA-Nickel column
318
(GE Healthcare, Freiburg, Germany) by using the elution buffer 25 mM phosphate sodium pH
319
7.4, 300 mM K2SO4, 500 mM imidazole. Mutants of OXA-232 were purified by using 2
320
anion-exchange chromatography HiTrapTM QHP GE Healthcare (20 mM Bis-Tris H2SO4 pH
321
7.2, then 20 mM piperazine H2SO4 pH 9.5) (7). Finally, a gel filtration step was performed for
the purifications of all -lactamases with 100mM sodium phosphate buffer pH 7 and 150 mM
323
NaCl with a Superdex 75 column (GE Healthcare, Freiburg, Germany). The protein purity
324
was estimated by SDS–PAGE and the pooled fractions were dialyzed against 10 mM
Tris-325
HCl pH 7 for the mutants and against 0.1 M Hepes (pH 7.5) for OXA-232 wt and
326
concentrated using vivaspin 20 (10 000 MWCOPES Sartorius) columns. Protein
327
concentration was determined according to the Bradford method using the Bio-Rad protein
328
assay standard II kit (Bio-Rad, Marnes-la-Coquette, France) with bovine serum albumin
329
(BSA) as a standard.
330 331
Kinetics assays. Steady-state kinetic parameters were determined using a spectrophotometer 332
ULTROSPEC 2000 (Amersham Pharmacia Biotech) and performed at 30°C in 100 mM
333
phosphate buffer (pH 7) for 10 minutes (7). The disappearance of substrate (-lactam
334
antibiotics) was monitored at the specific wavelengths and converted to initial velocities using
335
the specific extinction coefficients. The kcat and Km values were determined by analyzing
-336
lactam hydrolysis under initial-rate conditions using the Eadie-Hoffstee linearization of the
337
Michaelis-Menten equation with SFWIFT II software (7). Fifty percent inhibitory 338
concentrations (IC50) for imipenem were determined in 100 mM sodium phosphate buffer (pH 339
7) and 100 µM of benzylpenicillin as a reporter substrate. 340
341
Crystallization and data collection. Crystal conditions were screened with 10 mg/mL of 342
concentrated protein using a range of commercially available screens. OXA-232 crystals grew
343
in 0.2 M Lithium sulfate, 0.1M Tris pH 8.5 and 30% w/v PEG 3000. Crystals were dipped
344
briefly in cryoprotectant (20% v/v glycerol in the reservoir solution) and flash-frozen in liquid
345
nitrogen prior to be transferred in a stream of nitrogen at 100K (delivered by a Rigaku
X-346
tream cryosystem). A 2.21-Å resolution data set was collected on a Rigaku
HF rotating-anode generator with Cu Kα radiation, Varimax HF mirror focusing optics
348
(Rigaku) and MAR345 image-plate detector (MAR Research).
349 350
Structure determination and refinement. X-ray diffraction data sets were processed using 351
XDS (21) and AIMLESS (22); the crystals diffracted to 2.2 Å resolution (Table 4). Initial
352
phases were obtained by molecular replacement with MOLREP (23) using OXA-48 (PDB ID
353
3HBR) as a search probe. Refinement was performed by successive and alternate rounds of 354
refinement with BUSTER (24) and model improvement using Coot (25). The final model was 355
evaluated using MolProbity (26). Data collection and refinement statistics are provided in
356
Table 4.
357 358
Molecular modelling. Molecular modelling was performed to evaluate the effects of the 359
mutations on OXA-232 β-lactamase. Substitution of amino acid was performed by mutating
360
the OXA-232 structure (PDB code 5HFO) in silico using the Dunbrack rotamer library
361
(swapaa command) as a part of the UCSF Chimera software (14, 27). The Dunbrack rotamer
362
library predicts the conformation of the amino acid side-chain based on the global energy
363
minimum of the protein. The identification of interatomic clashes based on VDW radii (28)
364
was performed with UCSF Chimera software (14, 27). Three-dimensional structures of the
β-365
lactam ligands were generated using Corina 3.60 (Molecular Networks GmbH, Erlangen,
366
Germany). Molecular docking calculations were performed using Gold (Cambridge
367
Crystallographic Data Centre, Cambridge, UK) (29) and the GoldScore scoring function. The
368
binding site, defined as a 20 Å radius sphere, was centered on the OG oxygen atom of Ser70.
369
All other parameters had default values. The receptor-ligand complex images were produced
370
using UCSF Chimera (14, 27).
PDB accession codes. The OXA-232 structure atomic coordinates were deposited in the 372
Protein Data Bank with the accession code 5HFO.
373 374
Acknowledgments 375
All the authors are members of the Laboratory of Excellence in Research on 376
Medication and Innovative Therapeutics (LERMIT). This work was partially funded by the 377
University Paris-Saclay, and by grants from the French National Research Agency (ANR-10-378
LABX-33, and ANR-17-ASTR-0018). 379
380
Transparency declarations 381
We have no competing interests to declare. 382
REFERENCES 384
1. Nordmann P, Naas T, Poirel P. 2011. Global Spread of Carbapenemase-producing
385
Enterobacteriaceae. Emerg Infect Dis 17: 1791–1798.
386
2. Bush K. 2013. The ABCD’s of β-lactamase nomenclature. J Infect Chemother Off J
387
Jpn Soc Chemother 19:549–559.
388
3. Poirel L, Héritier C, Tolün V, Nordmann P. 2004. Emergence of
oxacillinase-389
mediated resistance to imipenem in Klebsiella pneumoniae. Antimicrob Agents
390
Chemother 48:15–22.
391
4. Poirel L, Potron A, Nordmann P. 2012. OXA-48-like carbapenemases: the phantom
392
menace. J Antimicrob Chemother 67:1597–1606.
393
5. Docquier J-D, Calderone V, De Luca F, Benvenuti M, Giuliani F, Bellucci L, Tafi A,
394
Nordmann P, Botta M, Rossolini GM, Mangani S. 2009. Crystal structure of the
395
OXA-48 beta-lactamase reveals mechanistic diversity among class D carbapenemases.
396
Chem Biol 16:540–547.
397
6. Naas T, Oueslati S, Bonnin RA, Dabos ML, Zavala A, Dortet L, Retailleau P, Iorga
398
BI. 2017. Beta-lactamase database (BLDB) – structure and function. J Enzyme Inhib
399
Med Chem 32:917–919.
400
7. Oueslati S, Nordmann P, Poirel L. 2015. Heterogeneous hydrolytic features for
OXA-401
48-like β-lactamases. J Antimicrob Chemother 70:1059–1063.
402
8. Matthews BW. 1968. Solvent content of protein crystals. J Mol Biol 33:491–497.
403
9. Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM,
404
Krissinel EB, Leslie AGW, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS,
405
Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS. 2011. Overview of the
406
CCP4 suite and current developments. Acta Crystallogr D Biol Crystallogr 67:235–
407
242.
10. Vaguine AA, Richelle J, Wodak SJ. 1999. SFCHECK: a unified set of procedures for
409
evaluating the quality of macromolecular structure-factor data and their agreement
410
with the atomic model. Acta Crystallogr D Biol Crystallogr 55:191–205.
411
11. Krissinel E, Henrick K. 2004. Secondary-structure matching (SSM), a new tool for
412
fast protein structure alignment in three dimensions. Acta Crystallogr D Biol
413
Crystallogr 60:2256–2268.
414
12. King DT, King AM, Lal SM, Wright GD, Strynadka NCJ. 2015. Molecular
415
Mechanism of Avibactam-Mediated β-Lactamase Inhibition. ACS Infect Dis 1:175–
416
184.
417
13. Lund BA, Thomassen AM, Carlsen TJO, Leiros H-KS. 2017. Structure, activity and
418
thermostability investigations of OXA-163, OXA-181 and OXA-245 using
419
biochemical analysis, crystal structures and differential scanning calorimetry analysis.
420
Acta Crystallogr Sect F Struct Biol Commun 73:579–587.
421
14. Dunbrack RL. 2002. Rotamer libraries in the 21st century. Curr Opin Struct Biol
422
12:431–440.
423
15. Gomez S, Pasteran F, Faccone D, Bettiol M, Veliz O, De Belder D, Rapoport M, Gatti
424
B, Petroni A, Corso A. 2013. Intrapatient emergence of OXA-247: a novel
425
carbapenemase found in a patient previously infected with OXA-163-producing
426
Klebsiella pneumoniae. Clin Microbiol Infect Off Publ Eur Soc Clin Microbiol Infect
427
Dis 19:E233-235.
428
16. Dortet L, Oueslati S, Jeannot K, Tandé D, Naas T, Nordmann P. 2015. Genetic and
429
Biochemical Characterization of OXA-405, an OXA-48-Type Extended-Spectrum
β-430
Lactamase without Significant Carbapenemase Activity. Antimicrob Agents
431
Chemother 59:3823–3828.
17. Poirel L, Naas T, Nordmann P. 2010. Diversity, epidemiology, and genetics of class D
433
beta-lactamases. Antimicrob Agents Chemother 54:24-38.
434
18. Potron A, Poirel L, Dortet L, Nordmann P. 2016. Characterisation of OXA-244, a
435
chromosomally-encoded OXA-48-like β-lactamase from Escherichia coli. Int J
436
Antimicrob Agents 47:102–103.
437
19. Hoyos-Mallecot Y, Naas T, Bonnin RA, Patino R, Glaser P, Fortineau N, Dortet L.
438
2017. OXA-244-Producing Escherichia coli Isolates, a Challenge for Clinical
439
Microbiology Laboratories. Antimicrob Agents Chemother. 61(9).
440
20. Potron A, Rondinaud E, Poirel L, Belmonte O, Boyer S, Camiade S, Nordmann P.
441
2013. Genetic and biochemical characterisation of OXA-232, a
carbapenem-442
hydrolysing class D β-lactamase from Enterobacteriaceae. Int J Antimicrob Agents
443
41:325–329.
444
21. Kabsch W. 2010. XDS. Acta Crystallogr D Biol Crystallogr 66:125–132.
445
22. Evans PR, Murshudov GN. 2013. How good are my data and what is the resolution?
446
Acta Crystallogr D Biol Crystallogr 69:1204–1214.
447
23. Vagin A, Teplyakov A. 2010. Molecular replacement with MOLREP. Acta
448
Crystallogr D Biol Crystallogr 66:22–25.
449
24. Bricogne, G., Blanc, E., Brandl, M., Flensburg, C., Keller, P., Paciorek, W., Roversi,
450
P., Sharff, A., Smart, O. S., Vonrhein, C. & Womack, T. O. BUSTER. Global Phasing
451
Ltd., Cambridge, England 2017.
452
25. Emsley P, Lohkamp B, Scott WG, Cowtan K. 2010. Features and development of
453
Coot. Acta Crystallogr D Biol Crystallogr 66:486–501.
454
26. Williams CJ, Headd JJ, Moriarty NW, Prisant MG, Videau LL, Deis LN, Verma V,
455
Keedy DA, Hintze BJ, Chen VB, Jain S, Lewis SM, Arendall WB, Snoeyink J, Adams
456
PD, Lovell SC, Richardson JS, Richardson DC. 2018. MolProbity: More and better
reference data for improved all-atom structure validation. Protein Sci Publ Protein Soc
458
27:293–315.
459
27. Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin
460
TE. 2004. UCSF Chimera--a visualization system for exploratory research and
461
analysis. J Comput Chem 25:1605–1612.
462
28. Tsai J, Taylor R, Chothia C, Gerstein M. 1999. The packing density in proteins:
463
standard radii and volumes. J Mol Biol 290:253–266.
464
29. Verdonk ML, Cole JC, Hartshorn MJ, Murray CW, Taylor RD. 2003. Improved
465
protein–ligand docking using GOLD. Proteins Struct Funct Bioinforma 52:609–623.
466 467
Table 1: Minimum inhibitory concentrations (MICs) of -lactams for E. coli Top 10 (pTOPO-OXA-48 and variants), E. coli Top 10
(pTOPO-468
OXA-232 and variants) and E. coli Top 10.
469 470
MIC (mg/L)
β-lactams E. coli Top 10 pTOPO-OXA-48 E. coli Top 10 pTOPO-OXA-232 E. coli
Top 10 wt R214S R214G R214L R214K R214D R214E wt S214R S214G S214L S214K S214D S214E Temocillin >256 32 32 64 128 8 8 32 256 32 64 128 8 8 8 Imipenem 0.75 0.25 0.25 0.25 0.38 0.25 0.25 0.38 0.75 0.25 0.25 0.5 0.25 0.25 0.25 471 472
473
Table 2: Minimum inhibitory concentrations (MICs) of -lactams for E. coli Top 10 (pTOPO-OXA-48), E. coli Top 10 (pTOPO-OXA-
474
181), E. coli Top 10 (pTOPO-OXA-232 and its variants), and E. coli Top10.
475
MIC (mg/L)
E. coli Top 10 (pTOPO-) E. coli Top 10
β-lactams OXA-232 S214G OXA-232 S214E OXA-232 S214K
OXA-232 OXA-181 OXA-48
Benzypenicillin >256 >256 >256 >256 >256 >256 64 Temocillin 32 8 128 32 >256 >256 8 Cephalothin 16 16 16 16 16 16 4 Imipenem 0.25 0.25 0.5 0.38 0.75 0.75 0.25 Meropenem 0.047 0.032 0.094 0.047 0.125 0.094 0.016 Ertapenem 0.012 0.012 0.094 0.012 0.19 0.094 0.004 476 477
478
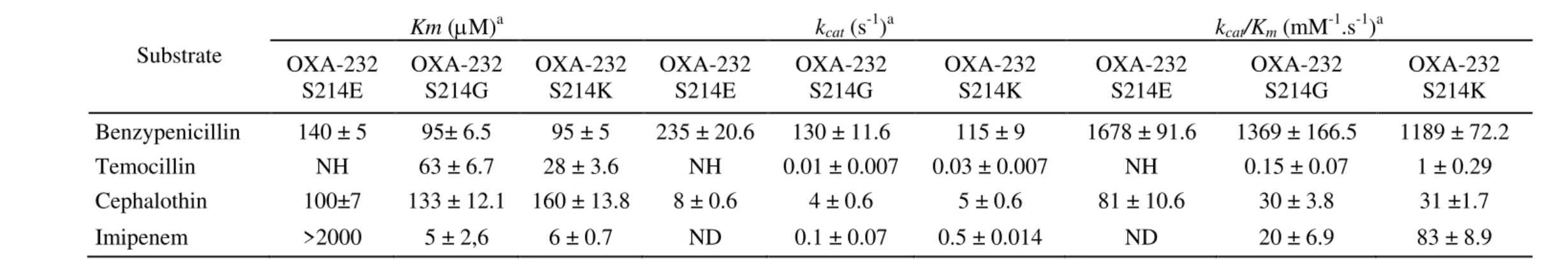
Table 3: Kinetic parameters of OXA-232 mutants
479 480
Substrate
Km (M)a kcat(s-1)a kcat/Km (mM-1.s-1)a
OXA-232
S214E OXA-232 S214G OXA-232 S214K OXA-232 S214E OXA-232 S214G OXA-232 S214K OXA-232 S214E OXA-232 S214G OXA-232 S214K Benzypenicillin 140 ± 5 95± 6.5 95 ± 5 235 ± 20.6 130 ± 11.6 115 ± 9 1678 ± 91.6 1369 ± 166.5 1189 ± 72.2
Temocillin NH 63 ± 6.7 28 ± 3.6 NH 0.01 ± 0.007 0.03 ± 0.007 NH 0.15 ± 0.07 1 ± 0.29
Cephalothin 100±7 133 ± 12.1 160 ± 13.8 8 ± 0.6 4 ± 0.6 5 ± 0.6 81 ± 10.6 30 ± 3.8 31 ±1.7
Imipenem >2000 5 ± 2,6 6 ± 0.7 ND 0.1 ± 0.07 0.5 ± 0.014 ND 20 ± 6.9 83 ± 8.9
NH, no detectable hydrolysis was observed with 1 μM of purified enzyme and 500 μM of substrate; ND, not determined. a Standard deviation 481
values are indicated after the kinetic parameter values. 482
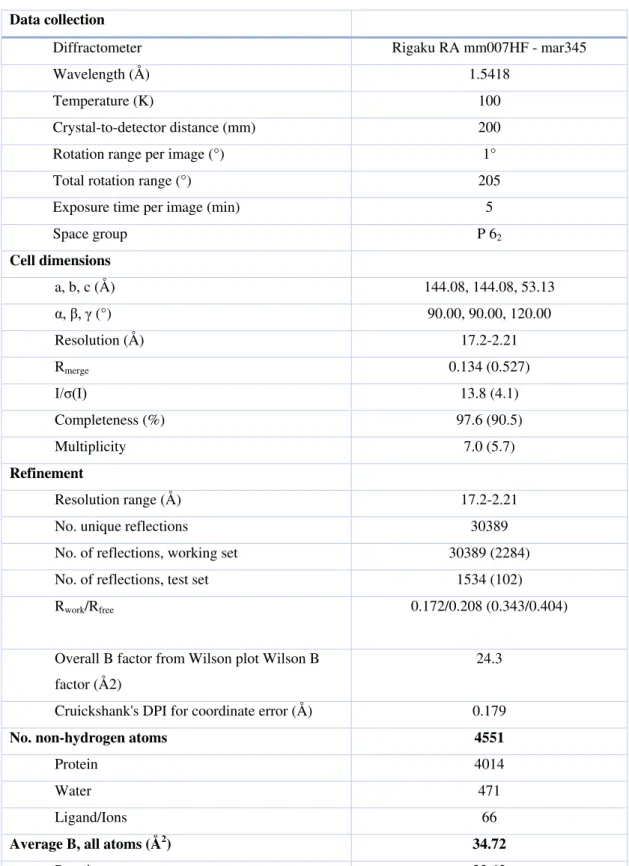
Table 4: X-ray crystallography and refinement statistics. 483
Data collection
Diffractometer Rigaku RA mm007HF - mar345
Wavelength (Å) 1.5418
Temperature (K) 100
Crystal-to-detector distance (mm) 200
Rotation range per image (°) 1°
Total rotation range (°) 205
Exposure time per image (min) 5
Space group P 62 Cell dimensions a, b, c (Å) 144.08, 144.08, 53.13 α, β, γ (°) 90.00, 90.00, 120.00 Resolution (Å) 17.2-2.21 Rmerge 0.134 (0.527) I/σ(I) 13.8 (4.1) Completeness (%) 97.6 (90.5) Multiplicity 7.0 (5.7) Refinement Resolution range (Å) 17.2-2.21
No. unique reflections 30389
No. of reflections, working set 30389 (2284)
No. of reflections, test set 1534 (102)
Rwork/Rfree 0.172/0.208 (0.343/0.404)
Overall B factor from Wilson plot Wilson B factor (Å2)
24.3
Cruickshank's DPI for coordinate error (Å) 0.179
No. non-hydrogen atoms 4551
Protein 4014
Water 471
Ligand/Ions 66
Average B, all atoms (Å2) 34.72
Protein 33.63
31.01 (m.c) / 36.11 (s.c)
Water 42.85
Ligand/Ions 46.37
Bond lengths (Å) 0.01
Bond angles (°) 1.03
Ramachandran plot
Most favoured (%) 98.2
Allowed (%) 1.8
Values in parentheses are for the outer shell (2.21-2.36Å) 484
Legends to figures 486
Figure 1. Amino acid sequence alignment of OXA-48 variants. Asterisks indicate identical 487
residues in all the three sequences, colon indicate a substitution with another amino acid but 488
with the same proprieties. Amino acid motif that are well conserved among class D 489
lactamases are indicated by grey boxes, and the black-outlined box corresponds to the 5-6 490
loop. Positions D159 and R/S214 are indicated by a thin and large arrow, respectively. 491
CLUSTAL O (1.2.4) multiple sequence alignment. 492
493
Figure 2. A Superposition of crystal structures and of 48 (yellow, PDB: 3HBR) and OXA-494
232 (grey; PDB: 5HFO). The 5-6 loop is delimited by the circle. B Partial active-site close-495
up of sulfate 302A-bound OXA-232. Atoms are colored by type (C, white;N, blue; O, red; S, 496
yellow). Hydrogen bonding and electrostatic interactions are depicted as blue dashes. C 497
Alternative docking conformations of imipenem (colored in magenta (A) and yellow (B), 498
closed form of the β-lactam ring) as non-covalent (Michaelis) complexes with the OXA-48 499
R214E mutant (colored in cyan, generated from structure 6P97). Hydrogen bonds and 500
favorable ionic interactions are shown as blue springs. Protein hydrogens are hidden for 501
clarity. D Docking conformation of temocillin (colored in green, closed form of the β-lactam 502
ring) as non-covalent (Michaelis) complex with OXA-48 (colored in pink, PDB 6P97), 503
superposed with the OXA-48 R214E mutant (colored in purple, generated from structure 504
6P97, only Gln214 residue shown). Hydrogen bonds and favorable ionic interactions are 505
represented as blue springs, and unfavorable ionic interactions are represented as yellow 506
thick lines. Protein hydrogens are hidden for clarity. 507
508 509