Publisher’s version / Version de l'éditeur:
Vous avez des questions? Nous pouvons vous aider. Pour communiquer directement avec un auteur, consultez la première page de la revue dans laquelle son article a été publié afin de trouver ses coordonnées. Si vous n’arrivez pas à les repérer, communiquez avec nous à PublicationsArchive-ArchivesPublications@nrc-cnrc.gc.ca.
Questions? Contact the NRC Publications Archive team at
PublicationsArchive-ArchivesPublications@nrc-cnrc.gc.ca. If you wish to email the authors directly, please see the first page of the publication for their contact information.
https://publications-cnrc.canada.ca/fra/droits
L’accès à ce site Web et l’utilisation de son contenu sont assujettis aux conditions présentées dans le site LISEZ CES CONDITIONS ATTENTIVEMENT AVANT D’UTILISER CE SITE WEB.
Biochemistry, 31, 21, pp. 4996-5004, 1992-06-02
READ THESE TERMS AND CONDITIONS CAREFULLY BEFORE USING THIS WEBSITE.
https://nrc-publications.canada.ca/eng/copyright
NRC Publications Archive Record / Notice des Archives des publications du CNRC :
https://nrc-publications.canada.ca/eng/view/object/?id=88a42b18-d5ea-4595-898c-93f9b254c4cd
https://publications-cnrc.canada.ca/fra/voir/objet/?id=88a42b18-d5ea-4595-898c-93f9b254c4cd
NRC Publications Archive
Archives des publications du CNRC
This publication could be one of several versions: author’s original, accepted manuscript or the publisher’s version. / La version de cette publication peut être l’une des suivantes : la version prépublication de l’auteur, la version acceptée du manuscrit ou la version de l’éditeur.
For the publisher’s version, please access the DOI link below./ Pour consulter la version de l’éditeur, utilisez le lien DOI ci-dessous.
https://doi.org/10.1021/bi00136a012
Access and use of this website and the material on it are subject to the Terms and Conditions set forth at
Helical epitope of the group B meningococcal α(2-8)-linked sialic acid
polysaccharide
Brisson, Jean-Robert; Baumann, Herbert; Imberty, Anne; Pérez, Serge;
Jennings, Harold J.
Biochemistry
Helical
Epitope
of
the Group
B
Meningococcal
a(2-8)-Linked
Sialic
Acid
Polysaccharide*
Jean-Robert Brisson,1Herbert Baumann,1 Anne Imberty,****8
SergePérez,8 and
Harold
J. Jennings*’1 Institutefor
BiologicalSciences, NationalResearch Councilof
Canada,Ottawa, CanadaK1A OR6, andLaboratoire dePhysicochimiedesMacromolécules, Instituí Nationaldela Recherche Agronomique, BP527, 44026NantesCédex, France Received December28, 1991; RevisedManuscriptReceivedMarch 11, 1992
abstract: The immunologicalproperties
of
the group Bmeningococcala(2-8)-linked
sialicacid poly-saccharide have been rationalized intermsof
a model where the randomcoil natureof
the polymer canbedescribed by thepresence
of
local helices. Theconformational versatilityof
theaNeuAc(2-8)aNeuAc
linkagehasbeenexploredby
NMR
studiesat 600MHz
in conjunctionwith
potentialenergycalculationsfor colominicacid,an
a(2-8)NeuAc
polymer,andthetrisaccharideaNeuAc(2-8)aNeuAc(2-8)j8NeuAc.
Potentialenergy calculationswere usedto estimate theenergeticallyfavorable conformers andtodescribe thewide range
of
heliceswhichthe polymercan adopt. Nounique conformerwas found to satisfyallNMR
constraints, andonly ensemble averaged nuclear Overhauser enhancements could correctly simulate the
experimentaldata. Conformationaldifferences between thepolymerandthe trisaccharide couldbebest
explainedinterms
of
slightchangesintherelativedistribution
of
conformersin solution.Similar
helical parametersforthea(2-8)NeuAc
polymerandpoly(A)
were proposedasthebasisfor their cross-reactivitytoa monoclonalantibodyIgMNOV. The unusual length dependencyfor binding
of
oligosaccharide to group Bspecific antibodieswas postulated toarisefromthe recognitionof
ahigh-order local helixwithan extendedconformation whichwas not highly populatedin solution.
(jTroup
BNeisseriameningitidisandEscherichiacoliK1are majorcauses ofmeningitis. Both bacteriahaveidentical capsular polysaccharides composedofa(2-8)-linkedsialic acid residues. Althoughvaccine developmenthasbeenhampered
by thepoorimmunogenicity ofthegroup B meningococcal polysaccharide inhumans,byN-propionylation ofits sialic acid residues, antibodies are produced which are both
bac-tericidal and protective [for reviews, see Jennings (1989, 1990)].
Thea(2-8)-linkedsialic acid homopolymerhasmany in-terestingimmunological propertieswhichhave beenextensively
studied. Thepresenceofa conformationalepitopewas
hy-pothesizedby Jennings et al. (1984) to explain theinability ofoligosaccharidesofup tofiveresiduestoinhibitthebinding
ofthegroup B polysaccharide toahomologoushorseantibody. Later,
it
was established that an unusually large deca-saccharide was required to bind or inhibit group B poly-saccharide specific antibodies (Jennings et al., 1985;Finne&
Makela, 1985;Hayrinenet al., 1989). Thisisincontrastwith
aconventionalsequentialoligomerwhereonlyamaximum6 or 7 residues are necessary for binding (Rabat, 1960). Cross-reactivityofahumanmonoclonal antibody(IgMNOV)
withthe a(2-8)NeuAc‘ polymer, poly(A), and other
poly-nucleotideshas also been observed (Rabat et al., 1988). Various attemptshavebeenmade to relatethese immuno-logical properties to the three-dimensional structure ofthe antigen. Lindon et al. (1984), from
NMR
studies at 360MHz,determined differencesintheorientationofthe exocyclic
chainsofthea(2-8)NeuAcanda(2-9)NeuAcpolysaccharides andinthe segmentalmotionofthe polysaccharides. Michon etal.(1987) determinedfrom
NMR
studiesat 500MHzthat the conformation ofthe linkagesofthe polysaccharideand fThisisNationalResearchCouncilofCanadaPublicationNo.32011.Part ofthisworkwas made possiblethrough the France-Canada ex-changeprogram grantedtoJ.R.B.andfinancial support from INRA.
1NationalResearchCouncilofCanada. 8InstituíNationalde laRecherche
Agronomique.
shortoligosaccharideswere different. Yamasakiand Bacon (1991),fromaquantitativeanalysisof2DNOE’sat 500MHz
usingrigidmodels, suggestedthatthepolysaccharide adopts orderedhelical structures in solution. Inthe present paper, theconformational propertiesofcolominicacid,an
a(2-8)-NeuAc polymer, and the trisaccharide
aNeuAc(2-8)-aNeuAc(2-8)/3NeuAcwere evaluated by
NMR
studiesat 600MHz,in conjunctionwithpotentialenergy calculations. The interpretationoftheNOE’swas based onamodelwhichallows
fortheinherent
flexibility
about thelinkages. The confor-mational versatilityofthemoleculewas estimated bypotentialenergy calculations. The biological implications were
ra-tionalized in termsofamodel wheretherandom coil nature
ofapolysaccharidecan bedescribedbythe presenceoflocal helices (Perez
&
Vergelati, 1985).Experimental
ProceduresNuclear MagneticResonanceExperiments. Thecolominic acid polysaccharideand the trisaccharidewere preparedas
describedbefore(Michonetal., 1987),at concentrations
of
10-20mgin0.5mLof95%D20, 5% acetone-1/6,pH7,Na+ counterion. Samples were purgedwith helium gas. Upon storage atacoldtemperature,samplesinsolutionwere found to bestableformonths, withno signs ofdegradation. The
H3a and H3e resonances of the reducing terminus were
partiallydeuterated(>50%) afterseveralmonthsinsolution.
All experiments were performed on a Bruker AMX 600 spectrometer at 288 R. Foradditional temperature stability, thedeuteriumlockwas establishedon internalacetone-</6. No
significantchangesin chemicalshiftwere observed upon the
addition
of
acetone-d6. Forcolominicacid,pureabsorptionphasesensitive2DNOEexperimentswere doneinTPPImode
(Bodenhausen et al., 1984) withsix mixing timesof25, 50, 75, 100, 150, and 200 ms. The relaxation delay between
1
Abbreviations: COSY, correlation spectroscopy; NeuAc,
N-acetylneuraminicacid;NOE,nuclear Overhauser enhancement;NMR,
nuclear magneticresonance;TPPI, time-proportionalphase increments.
ConformationofMeningococcalB Polysaccharide Biochemistry, Vol. 31, No. 21, 1992 4997
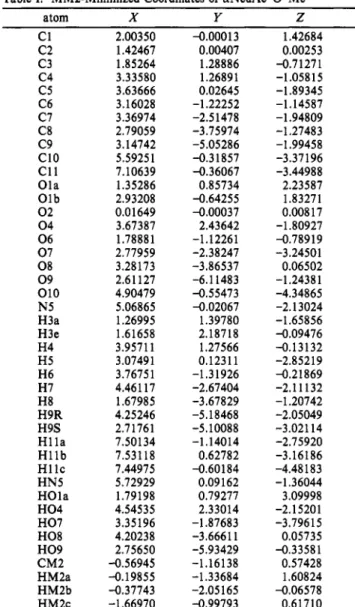
TableI: MM2-MinimizedCoordinatesofaNeuAc-O-Me
atom X Y Z Cl 2.00350 -0.00013 1.42684 C2 1.42467 0.00407 0.00253 C3 1.85264 1.28886 -0.71271 C4 3.33580 1.26891 -1.05815 C5 3.63666 0.02645 -1.89345 C6 3.16028 -1.22252 -1.14587 C7 3.36974 -2.51478 -1.94809 C8 2.79059 -3.75974 -1.27483 C9 3.14742 -5.05286 -1.99458 CIO 5.59251 -0.31857 -3.37196 Cll 7.10639 -0.36067 -3.44988 Ola 1.35286 0.85734 2.23587 Olb 2.93208 -0.64255 1.83271 02 0.01649 -0.00037 0.00817 04 3.67387 2.43642 -1.80927 06 1.78881 -1.12261 -0.78919 07 2.77959 -2.38247 -3.24501 08 3.28173 -3.86537 0.06502 09 2.61127 -6.11483 -1.24381 010 4.90479 -0.55473 -4.34865 N5 5.06865 -0.02067 -2.13024 H3a 1.26995 1.39780 -1.65856 H3e 1.61658 2.18718 -0.09476 H4 3.95711 1.27566 -0.13132 H5 3.07491 0.12311 -2.85219 H6 3.76751 -1.31926 -0.21869 H7 4.46117 -2.67404 -2.11132 H8 1.67985 -3.67829 -1.20742 H9R 4.25246 -5.18468 -2.05049 H9S 2.71761 -5.10088 -3.02114 Hila 7.50134 -1.14014 -2.75920 Hllb 7.53118 0.62782 -3.16186 HI lc 7.44975 -0.60184 -4.48183 HN5 5.72929 0.09162 -1.36044 HOla 1.79198 0.79277 3.09998 H04 4.54535 2.33014 -2.15201 H07 3.35196 -1.87683 -3.79615 H08 4.20238 -3.66611 0.05735 H09 2.75650 -5.93429 -0.33581 CM2 -0.56945 -1.16138 0.57428 HM2a -0.19855 -1.33684 1.60824 HM2b -0.37743 -2.05165 -0.06578 HM2c -1.66970 -0.99793 0.61710
acquisitions was 6 s, and the sweep width was 2400 Hz. Matricesof512by 2048 pointswere recordedwitheightscans
pertxincrement. Transformationwasdoneusinga /2-shifted
sine-squaredbell
filter
in both dimensionand zero-fillingtoa square matrix
of
1024 X 1024 points for a final digitalresolutionof2.4Hz/point. Afirst-order polynomialbaseline correlationwas applied in the
fl
dimensionprior to volume integrationofthe cross-peaks. Theaverageoftheequivalent volumes aboveandbelowthe diagonalwas usedforanalysis.All
data processingwas done usinguxnmr software providedby Bruker. Steady-state ID NOE experimentson the
tri-saccharidewere doneasdescribed before(Michonet al., 1987). The spectrum
of
afewmilligrams ofcolominic acidwastakenat various temperatures up to343K,inorder to observe the proton coupling constants. As described previously
(Michonet al., 1987),protonassignmentswere obtainedfrom
a COSY experiment. Spin simulations were done using
LAOCN5(QCPE) withftnmr (HareResearchInc.). Theerror on the coupling constantswas ± 0.3 Hz.
Calculations. The starting geometryof aNeuAcwas built
fromthecrystal structureoftheßsialicacidmethylglycoside (Flippen, 1973) afterproper settingofthe anomeric config-uration.
It
was thensubmitted to acomplete refinementof
bond lengths,valence, and torsion angles byusing the
mo-lecular mechanics program MM2(87) (QCPE) (Burket
&
Allinger, 1982).
All
calculationswere done usingthemini-mized coordinatesforthemethylglycosideofaNeuAc (Table
I).
Foragivendisaccharide, the conformationalanalysiswas
evaluated usingthePFOSapproachwhich included the par-titioned contributions arisingfromvan derWaals interactions andtorsionaland exo-anomericpotential (Tvaroska
&
Perez, 1986). Rotational barriersof1.0and0.5kcal/molwere used for and,
respectively. Noelectrostaticcontributionwasconsidered. The ring geometrywas treatedasinvariant,and
hydroxylichydrogen atoms were not takenintoaccount. Four angles determine the linkage conformation for
aNeuAc(2-8)aNeuAc(h-a): =
(¿>06-¿>C2-¿>02-aC8),
= (¿>C2-¿02-tiC8-tiC7),
a>8 =
(07-C7-C8-08),
and 7 =(06-C6-C7-07).
Theorientationofthe pendant groups forCl
andC9was definedby =(Q6-C2-C1
-01
A) ando>9 =(08-C8-C9-09). All
anglesare expressedindegrees. Theangles<vb 7, 8, and 9 were alwaysset the same forboth
residues. Thecv7anglewas setat60°,and separate mapswere
generatedfor 8 = 60°,
-60°,
and 180°(g+,g",t). The
( , )
mapswere calculated in thefollowingway. Foragivenset of 7, w8values, theglycosidic torsionalangles and werestepped in increments of 5° over the angular range which containedlowenergyminima,-120°to+240°for and40° to 220° for
.
At
eachpointofthisgrid search, the wj and9 angleswere stepped in increments
of
20° over the whole angular range, andtheminimumenergy conformerwasse-lected. Suchaprocedureallowedsevere steric constraintsto
berelievedbetweenthehydroxymethyl group atC9 and the carboxyl group at
Cl
across the glycosidic linkage.All
the low energy conformers derivedfromthe analysisofsuch“rigidpotentialenergysurfaces”were refined using theMM2(87)
software. From alltheseminimizedconformers, theaverages
ofthe and<v8anglesforthethree staggeredtu8conformers
were takenand therigidmapswere calculated againusing
thesemean values(Table
II).
TheaverageglycosidicbridgeangleforbC2-b02-aCS of 117° was also used.
Helicesare customarilydescribedintermsofasetofhelical parameters, (n,h),withnbeing the numberofresiduesperturn
of
the helix and h being the translationof
correspondingresidues along thehelixaxis. The parameternisderivedfrom the rotationangle (µ) about the helix axis per repeatunit,
wheren =
2 /µ.
Withsetvaluesofglycosidic valence angles andresidue geometry, these parameterscan becalculatedforTableII: MinimumEnergy Conformersfor<*NeuAc(2-8)a:NeuAc
E“ ^ OJg (Ug "1 <oi>6 n hc h/L
Gl+ 3.2 -45 130 55 70 138 -40 9.1 2.3 4.8 0.8 G2+ 0.0 75 100 55 70 138 -20 6.5 3.6 -3.2 -0.5 G3+ 2.9 -180 105 55 70 138 -40 6.7 11.3 5.7 0.9 Gl- 3.6 -50 120 60 -62 -162 -40 6.3 2.5 2.9 0.7 G2- 0.3 60 115 60 -62 78 -20 7.4 2.2 -4.2 -1.0 G3- 2.3 170 135 60 -62 -162 -40 6.9 2.8 -1.9 -0.5 T1 5.2 15 155 67 179 138 180 3.8 2.9 1.2 0.2 T2 1.0 80 160 67 179 98 180 4.7 3.0 '4.0 0.8 °Relative
Biochemistry, 31,
figure 1: Structureof aNeuAc(2-8)aNeuAcshowingthefour torsion
angleswhich determine thelinkage conformation with
(06-C6-C7-07) = 7 = 60° and(07-C7-C8-Q8) =
8 = 60°. The
ori-entation ofthe pendant groups at Cl and C9 isdefined by =
(06-C2-C1-01A)and 9 = (08-C8-C9-09),respectively.
any given value of the set of torsional angles (Sugeta
&
Miyazama,1967). Thelengthofthevirtualbond(L) between two adjacentlinking atoms,namely,02
and08,
represents themaximum ofthe possible extension. The quotienth/L
yields the percentageoftotalextension. The chiralityisdefined by the signofh,positivefor right-handedhelices and negativeforleft-handedhelices.
The NOE was calculated using the complete relaxation
matrix, under the assumption of isotropic tumbling. The
steady-stateNOEwascalculatedasdescribedbefore (Brisson
&
Carver, 1983). The 2DNOEwas calculated accordingto Borgias and James(1988). Determinationofdistancesfrom 2D NOE data was done using the distance extrapolation method (Balejaet al., 1990). The ensemble averagedNOE
was calculatedasdescribed byGummingandCarver (1987). For betterBoltzman weightingoftheNOE,apotentialenergymapwith Io increments in
( , )
was generatedfromthe 5° interval map using cubic spline interpolation. Only (r™6)motional averaging was used. Averagingwas onlydonefor
( , ),
theo>7, 8, and 9 anglesbeingfixed. Foragiveno>7and 8value, the
( , )
potentialenergy surfaceforwhich , and (ti9adopted minimumenergyconformers was used (see above) andtheaveragedNOEwas calculated separatelyfor(ti9 = 60°,
-60°,
and 180°. This couldbe done becausenomajor interglycosidic NOE was observed on the H9
reso-nances. Therelativeenergyofconformerswithinawellwere
modifiedbyfirstdefiningacontourlevelwhichencompassed
alocalminimaandthen changing the energyoftheconformers at thegridpointswithinthewellbythesame amount. The relative population within a well was calculated using the Boltzmanndistributionon a 10 X 10 gridinterpolatedfrom a 5o X 5° grid.
Inorder to calculate theNOE,therotationalcorrelation timeofthe molecule mustbedetermined. It was estimated fromabest
fit
ofthecalculatedNOEtoH3a-H5andH3e-H5 forvarious rigid conformers. Correlation timesof8 ns
and0.5nswere usedfor simulationoftheNOEfor colominic acid and the trisaccharide, respectively. The “standard deviation”,Sdev,fortheNOEwas calculatedusing7V(Sdev)2
=
( „
)2,
whereN
isthenumberof
points. The n - hplots for poly(A) were generated ina similar manner as described before (Yathindra & Sundaralingam, 1976) usingX-raycoordinates (Saenger etal., 1975). Ball andstickmodelswere generatedwiththeschakal program.
Calculationswere performedon Micro VAX 3200 and 3500.
Results
In order to understand the conformational behavior
of
«NeuAc(2-8)«NeuAc(Figure 1), thelinkage conformation
,
,
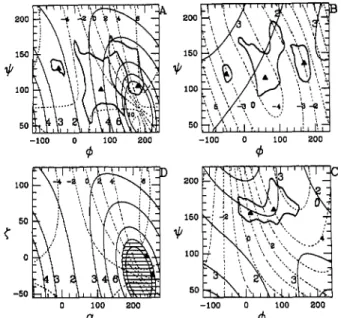
7, and 8, andtheorientation ofthe pendant groups mustbedetermined. Thepyranoseringadoptsawell-definedfigure 2: Potentialenergy contourmapfor aNeuAc(2-8)«NeuAc with( 7, 8)= (55°,70°)(A), (60°,-62°) (B),and(67°,179°) (C)
and(tijand 9 adoptingminimumenergy conformers. Withrespect
tothe globalminimumofeach section,the8,6,and10kcal/mollevels are drawnfor(A),(B),and(C),respectively. For(A),fromleftto right,three seperate wells,WG1+, WG2+,andWG3+,arethusdefined.
Similarly for (B),three wellsWG1",WG2™,andWG3'are defined. Localenergyminima (TableII)areindicated. Thehelical parameters
n (solid lines) andh(dotted line) are alsodisplayed. For poly(A)
(D),then- h
map and the energetically favoredhelices (dashed area)
are shown. The heliceswith (n,h) = (9,2.8Á) and (9,5.5Á)are
marked.
2C5conformation,andthe
NAc
groupat C5 adoptsaplanarconfiguration withthe
H5-C5-N5-HN5
inthe trans con-figuration (Christian&
Schultz, 1987).PotentialEnergy Calculations. Potentialenergy calcula-tionswere performed in ordertoassess the
( , )
conforma-tional spaceforthestaggeredrotamers
of
8, with 7 fixed to theg+rotamer (Figure2). Thehelical parameters, ob-tained upon propagation of the molecule, are alsoplotted. Three( , )
mapswere represented wherethe , and 9ro-tamers were allowed to adoptminimum energyconformers
foreachsettingofthelinkage conformation. The 7 and 8 angleswere optimizedbymolecularmechanicscalculations. For the
( 7, 8)
=(55°,70°) map(Figure2A), three major wells,WG1+, WG2+,andWG3+, whichencompassminima
Gl+,G2+, and G3+(Table
II)
are definedbythe8 kcal/mollevelwithrespecttotheglobal minimum. Forthe ( 7, >8) =
(60°,-62°) map (Figure 2B), three separate wells, WG1", WG2~, andWG3“, couldbedefinedby the6 kcal/mollevel abovetheG2~conformer (Table
II).
The( , )
energy map for((ti7,(ti8) = (67°,179°) mapswas more restricted,andonly the 10kcal/mollevelwithrespect tominimum T2 (TableII)
isshownin Figure2C. Althoughother localminima could befoundwithinthese wells,onlytheareas defined bythese wellswere useful in interpreting theNOEresults. Relevant parameters forvarious low-energy conformers are givenin
TableII. Molecularmechanics calculationswere performed
forvariousconformerswithinthesewells (data not shown).
It was foundthattheseconformerswere stable,the linkage
anglesnot changingsignificantlyuponcompleteminimization.
Multipleorientationsofo>9were alsofavorable. The ,
ori-entationwas mostly restrictedto conformers around -170°
and-40°.
In orderto estimate the role
of
the charge, the average distance betweenthefouroxygen atomsforthecarboxylgroupConformationofMeningococcalB Polysaccharide Biochemistry, Vol. 31, No. 21, 1992 4999
Table III: Experimental 2DNOEVolumes‘ with75msof MixingTimeforColominicAcidandCalculated Relative NOE6
protons exp average' minima <G+> <G-> <T> Gl+ G2+ G3+ H3a-H3e 100 100 100 100 100 100 100 H3a-H8 9 8-11 8 2 32-41 2 28 H3e-H8 5 5-6 5 2 17-23 2 47 H3a-H7 3 3 2 2 4-6 2 74 H3e-H7 3 3 1 1 3 3 60 H5-H8 4 2-4 3 2 12-16 2 5 H3a-H5 24 22 22 22 24 22 30 H3e-H5 12 12 11 11 12-14 11 16 Sdev1' 1-2 2-3 4 10-15 4 34 H8-H6 28 26-33 5-6 7 30-42 26-30 40 H8-H7 30 20-24 27 9 22-31 19-22 31 H3a-H9“ 2 2-19 1-6 1 7-80 1 3 H3e-H9' 1 1-11 1-3 1 4-57 1 4 H7-H9' 11 3-4 6-18 4-19 5 3 4
“Estimatederror = ±25%. 6TheH3a-H3eNOEissetto 100. ( 7, 8) forG+conformers = (55°,70°), for(G
> = (60°,-62°),andfor<T) = (67°,179°).
( , )
forGl+= (-45°,130°), forG2+= (75°,100°),forG3+= (180°, 105°). Significantchangesfor 9 = 60°,-60°,and 180° are shown. '
( , )
ensemble averagedNOE. Relativeenergies(in kilocaloriespermole)forwells WG1+, WG2+,andWG3+are 1.2:0:3andforwells
WG1", WG2“ andWG3"are 0.5:0:2. ‘'Standard deviationofthe fitfor sixmixing timesandforthefirst8volumesonly. The calculatedNOE
rangeforH9RandH9Sisalsoincluded.
5
figure 3: Partial NMRspectraof colominicacid at 600MHz and343K: (a)spinsimulated spectrum and (b) resolutionenhanced
spectrum.
less favored in solution due to the close proximity of the charged groups betweenadjacentresidues,whileforall the gauche conformers the carboxyl groups are further apart.
Colominic Acid. Colominic acid, which has a proton
spectrum identical tothat
of
thegroupBmeningococcal po-lysaccharide (data not shown), was used sinceit
was morereadily available. Thechoiceofcounterion, thepH,andthe molecular weightfractionofthe polymerare importantfactors whichwere consideredin order togeta sampleas
homoge-neous aspossible anda stable samplewhich did not degrade
with time. The high molecular fraction of commercially available colominic acid (Na+ salt) was selected by gel
fil-tration. ThepH was adjusted to 7. Differentsampleswerepreparedwhichalways gave thesame spectra(Michonet al., 1987). Especially, the H8andH9resonances were resolved which permittedthe detectionof
NMR
parameterswhichwerecrucialinassessingthesolutionconformationofcolominic acid. In order to determine more accurately the vicinal proton
couplingconstants forcolominicacid, the spectrumwas
obtained at343
K
(Figure3).All
multipletscouldberesolved andcorrectlyspinsimulatedto obtain accurate parameters. The proton coupling constants did not changewithtemperature (Lindonet al., 1984). Proton couplingconstantswere found to bethesame asthoseforresidue bof
thetrisaccharideat 300K(Michonet al., 1987), exceptfor/(H8,H9')
whichwas5.5 Hzinstead of6.9 Hz.
The 2DNOEexperimentwithamixing timeof75ms, at
600MHz,and 288K, forcolominic acidisshownin Figure
4. Thevolumesthatcouldbeaccuratelymeasuredare given
7
figure 4: 2DNOEspectrum at 600MHzand 288K ofcolominic
acid. Themixing timewas 75ms. The IDspectrumisshownon
top along the
fl
axis.in Table
III,
and some are plotted in Figure 5.All
NOEcalculations were done using the coordinates for the di-saccharide(b-a). Adimercan be usedtosimulate theNOE’s
becausethe mostfavorable conformers formextended helices
whenpropagated (Figure2)with pitches(nh) greater than
5 Á. Hence, long-range interactions between nonadjacent residues
will
notbeimportant. For colominicacid, theNOE betweentwo protonswas taken asthesumof
theinter-and intraresidue NOE ofthe equivalent pair. Forexample, theH3a-H7 NOEwas thesum ofthe ¿H3a-¿>H7NOEandthe
¿>H3a-aH7 NOE. Forsome conformers, theinter- and in-traresiduecontributions were comparable.
TheNOE’s for
H3a-H5
and H3e-H5are dominated bytheintraresidueNOEonly. Fromthe2DNOEbuilduprates, using the distance extrapolation approximation (Table
IV),
theH3a-H5 andH3e-H5distanceswere foundto be2.5and
Biochemistry,
figure 5: 2DNOEbuildup rate for colominicacid. The experimental
volumesfor(H3a-H3e)/2
(·),
H3a-H5( ),
H3e-H5( ),
H3a-H8(A), H3e-H8
( ),
H5-H8 (O),H3a-H7 (X),andH3e-H7(+)are shown. The( , )
ensembleaveraged theoreticalNOE builduprates (solid lines) using thepotentialenergymapfor 7 =55°, 8 = 70°
(Figure 2A) withanaltered relative energyforwells WG1+, WG2+, andWG3+of1.2:0:3kcal/moland the 9valuewhichgavethebest
fitare alsoplotted.
TableIV: DistancesforaNeuAcObtained from Minimized
Coordinates, from Distance Extrapolation0ofCalculated 2DNOE
Using ConformerGl+,andfromtheExperimental 2D NOE6 buildup
RateforColominicAcid
NOE
protons coordinates caled exptl
H3a-H3e 1.79
H3a-H5 2.51 2.5 2.5
H3e-H5 3.74 3.3 3.3
°a0 fromfit of y =
a0+ axx+ a2x2,where (y/r0)6 =
V/VQ, V0 =
H3a-H3evolume,r0= 1.79A, V=
cross-peakvolume, andx = mix-ing timesof25, 50, 75, 100, 150, and 200ms. ‘Estimatederror = 0.2
inagreementwiththe distanceof2.51
Á
calculatedfromthe coordinates ofTable I,adistanceof3.74Á
isexpected for H3e-H5. However,if
thebuildupratesare calculatedusing thecoordinatesofconformerGl+
andtheH3e-H5distance extrapolated from thecalculated volumes, avalueof
3.3Á
isobtained. Thereason for this discrepancy is thathigherorder effectsare neglectedinthe distance extrapolation method (Landy
&
Rao, 1989). Hence,inorder to accountforcorrect summation of the inter- and intraresidue NOE’s and the complete spin geometry,afullmatrixanalysiswas alwaysused.TheNOEalso depends on allpairson interprotondistances
and thuson
,
,
7, 8,and 9. For motionalaveraging, theNOEmustbeaveragedinacertain way dependingontherates ofinternal motion (Kessler etal., 1988). Inthiscaseonly
(r6)
averagingwas considered, since the modeofaveraging
will
notaffect anyoftheconclusions reachedin this study. The r"6 terms for a conformer were weighted according to theBoltzmann distribution function.
TheNOE’swere analyzed usingamodel wherethe molecule
was allowed tosampleenergetically favored conformersover
( , )
with the 8 and 9 rotamers adopting staggeredcon-formersonly. The 7 anglewas kept fixedtotheg+rotamer. Since,ingeneral,NOE’s thatwere highlydependentof and
were not greatly affected by thevalueof 9, thepotential energy map where 9 isallowed tofreelyrotate couldbeused
to calculatethe
( , )
ensemble averagedNOE for differentvalues
of
<u9. TheNOE’s for H3a-H8, H3e-H8, H3a-H7,H3e-H7,andH5-H8 satisfied this conditionandthuscould
be usedtoassessthe
( , )
linkage conformation. The NOE’sbetween
H3a-H9
andH3e-H9were notused becausetheywere highlydependenton 9, and theH9assignment toeither
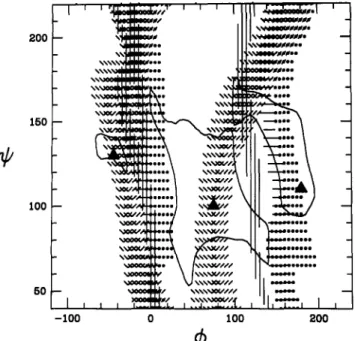
figure 6: NOEconstraints mapofcolominic acidforamixing time of75ms. Thecalculatedrelative NOE’susing thefullrelaxation matrix, whichsatisfy theexperimental NOE’s (Table
III)
withan error boundof±25%are plottedas afunctionof( , ),
with 7 =55°, 8 = 70°. TheNOEconstraintsforH3a-H8 (-),H3e-H8
(|), H3a-H7(/),H3e-H7(\),andH5-H8(O) relative toH3a-H3eare
displayed. The mapsfor 9 = 60°,-60°,and 180°areoverlaid. The
8kcal/mollevelswhichdefine the threemajorenergy wellsWG1+, WG2 ,and WG3+(lefttoright) around theGl+,G2+, and G3+ confomers(Table
II)
are also shown.H9Ror H9Swas ambiguous. Still,thesecalculatedNOE’s for differentvalues of<o9 were within the range of the
ex-perimentalones. Conversely, theNOE’sfor H8-H6, H8-H7, and
H7-H9
couldbeusedto determinetheorientationofthe exocyclic chainsincetheywere nothighlydependenton( , ).
Theconformationofthe exocyclic chainwas determined fromthe protonvicinalcoupling constantswhicharedependent on thetorsionalangle
H-C-C-H
andfromNOE’s. Fromaspinsimulationofthe spectrum at343 K,
/(H6,H7)
and/(H7,H8)
were found to be 1.0 and 2.3 Hz, respectively(Figure 3). Also, the strongNOE for
H8-H6
andH5-H7
(Figure4), indicatedthatthe 7 = 60°and
8 = 60° rotamers
were predominant in solution. Forthehydroxymethyl group at C9,
/(H8,H9)
and/(H8,H9')
were 4.5and 5.5Hz,indi-catingthatthestaggeredrotamersfor 9 were allaccessible. These resultsare similarto theones obtainedforthe
corre-spondinglinkage inthetrisaccharide (Michonetal., 1987). However, an NOE between the H7 and the H9 resonance
indicated thatthe other rotamers for 8 were also present.
Most probably,theg" rotamer was present dueto the small
/(H8,H7)
= 2.3Hzandthe largeH8-H7
NOE,whichcanarisefrom bothgauche rotamers (Table
III).
FiveNOE’s couldbe used to evaluate the
( , )
linkage conformation,namely thoseforH3a-H8, H3e-H8, H3a-H7,H3e-H7, and
H5-H8.
AnNOEconstraints mapwascal-culated to determine
if
allsetsofNOE’scan satisfyauniqueconformer. TheNOEwas calculatedforeachgridpointand
if
itsvalue fallswithinerror boundsofthe experimentalNOE, it was indicatedon themap. Forg+rotamers of 7 and 8, threemapswere calculatedforstaggered rotamers of 9 and overlaid. Nounique energetically favorable conformer satisfiedthefive experimentalNOEconstraints (Figure6). Similar resultswere obtainedfortheotherstaggered rotamer of 8. The experimental NOE’swere successfullysimulatedusing motionalaveragingover
( , )
(TableIII,
Figure5). FortheConformation ofMeningococcal B Polysaccharide Biochemistry, Vol. 31, No. 21, 1992 5001
Table V: Experimental IDNOE" for aNeuAc(2-8)«NeuAc(2-8)/3NeuAc (c-b-a)andCalculated Relative NOE4
protonssat.obsd exptl
average' minima (G+> <G> <T> Gl+ G2+ G3+ cH3a-cH3e 100 100 100 100 100 100 100 cH3a-6H8 16 10-17 14 3 39-44 1 19-21 cH3a-6H6 3 2 1 4 6 1 12 cH3a-cH4 31 28 27 28 28 27 33 cH3a-cH5 53 55 57 59 51-54 56 78 cH3a-Z>H7 _d 3 2 2 5 2 80 cH3a-cH6 6 6 4-7 8 6 6 8 cH3a-cH7 3 4 3 3 4 4 6 cH3e-cH3a 100 100 100 100 100 100 100 cH3e-6H8 7 4-7 8 5 13-16 1 58-65 cH3e-6H6 4 1 1 6 2 1 22 cH3e-cH4 46 55-64 54 52 63-102 51 66 cH3e-cH5 21 22 21 22 20-26 21 23 cH3e-6H7 ud 4 1 2 3 5 55 cH3e-cH6 7 6-8 4-8 8 7-12 6 8 cH3e-cH7 -e 2 2 2 4 2 3 SdeV 4-6 4 5 9-19 6 40 6H8-6H7 100 100 100 100 100 100 100 6H8-6H6 114 102-106 12-23 71-87 108-122 104 240 6H8-6H3a 21 16-19 12-14 6 33-64 2 43 ¿>H8-6H3e 14 7 6-8 8 12-21 2 130 6H8-6H5 12 8 5 37-48 3 13
“Estimatederror = ±25%. 4ThereferenceNOEissetto 100. ( 7, 8) forG+conformers = (55°,70°), for<G) =
(60°,-62°),andfor (T) =
(67°,179°).
( , )
forGl+= (-45°,130ß), forG2+= (75°,100°),andforG3+= (180°,105°). Significantchangesfor , = 60°,-60°,and 180»
areshown. '
( , )
ensemble averagedNOE. Relativeenergies(in kilocaloriespermole) forwellsWG1+, WG2+,andWG3+are 0.4:0:3andforwells
WG1", WG2",are 0.5:0:2. ‘'ResonancescH5and6H7overlap. ‘AbsoluteNOE<1 ± 1. ^ForcH3aandcH3eNOE’s only.
7 = 55°, 8 = 70° energy surface, threemajorwells,WG1+,
WG2+,andWG3+, whichencompassconformers
Gl+,
G2+, and G3+can bedefined by the8kcal/mollevelwithrespect totheglobalminimumG2+ (Figure2A).With
therelative energy for minimaGl+,
G2+, and G3+ of 3:0:3 kcal/mol obtainedfromthe potential energy calculations, the relativepopulationofwellWG1+was 99%,whiletheotherwells had
lessthan 1%occupancy. The
( , )
ensemble averagedNOE’s calculated using this distribution were the same as those calculated using the singleminimum conformerG2+, sincefor allconformerswithinwell WG2+, interglycosidic NOE’sweresmall, compared tothoseforconformer
Gl+
or G3+(TableIII).
Hence, poor agreementwas obtainedwiththeexperi-mental values. However,
it
was observed that the NOE’sshowedvery
little
dependenceon andthatinterglycosidicNOE’s had maxima near conformers
Gl+
and G3+. Forconformer
Gl+,
H3a interglycosidicNOE’sare bigger thanthose
of
H3e, whilefor conformerG3+ the reserve is true. Hence,aslightincreaseinthe populationofwellWG1+,would leadtogoodagreement betweentheaveragedNOE’sand the experimental data. The occupancyofwellWG3+mustremain low,ortherelative magnitudeoftheH3a-H3eNOE’s would change.A
relativeenergy between wellsWG1+,WG2+,and WG3+ of 1.2:0:3 kcal/mol, equivalent to a population of13:86:1, was found togive the best
fit.
Only the ratio forWG1+andWG2+was altered, theone for WG2+andWG3+ being kept fixed as that found from the potential energy
calculations. The
( , )
ensemble averaged NOE’s werecalculatedforthe threestaggered 9 rotamersandexhibited
no majordependencyon 9, except thoseinvolvingH9(Table
III).
Sinceother rotamersfor 8 can bepresent,
( , )
ensemble averagedNOEcalculationswere alsodonefor allstaggered 8 rotamers. Forthe g"rotamer,energy wellssimilar tothose fortheg+rotamerwere observed, andagoodfit
ofall NOE’s, exceptforH8-H6,
couldbeobtained (TableIII).
Distinctwells WG1-, WG2-, and WG3“ could be defined by the 6
kcal/mollevelwithrespecttoconformerG2" (Figure2B). The relativeenergyofwellWG1"hadtobemodifiedfora good
figure 7: ID NOEdifference spectraof
aNeuAc(2-8)aNeuAc(2-8)/3NeuAc(c-b-a)for saturationofH3aofresidueaandc(a),of
H3eofresiduec,andofH8 andH9ofresidueb(c)are shown along
withthe ID spectrum(d).
fit.
The relative energyof
well WG3" was not altered. Relative energiesforwellsWG1",WG2“,andWG3™of0.5:0:2kcal/molequivalent toa populationof8:90:2 gavethe best agreement between the averagedNOE’sandtheexperimental data. Poor agreementwas foundforthe transrotamer (Figure 2C),sinceno energeticallyfavored conformersnear = -50°
were present where stronginterglycosidic NOE’swiththetwo
H3 geminalresonances occurred.
Trisaccharide.
NMR
experimentswere performedon the trisaccharide in order to compare the linkage conformationofthe polymer to theoneofthe linkagec-bofthetrisaccharide
aNeuAc(2-8) aNeuAc(2-8)0NeuAc,(c-b-a). The
ID
NOEspectra for the trisaccharide at 600 MHz and 288 K are
plotted inFigure7,and theNOEdataare givenin Table V. Byperforming the experimentsat alower temperature, the correlation time
of
the molecule was decreased, leading to bigger negative NOE’s, which couldbemore accuratelyin-tegrated. The chemicalshiftassignments andcoupling
con-stants were similartothosereported before (Michonet al., 1987).
Asfor colominicacid,astrongNOEbetween¿>H8andZ>H6,
Biochemistry,
the 7 = 60° and 8 = 60° rotamerswere predominant in
solution. Due topartialoverlapof6H9and6H8resonances,
the6H9resonance could notbeselectivelysaturatedand the
smallNOEbetween H9 andH7, which might indicate the presence of other 8 rotamers, could not be evaluated, as observed forcolominic acid.
Four sets of NOE’s could be used to evaluate the
( , )
linkage conformation distribution, namely interglycosidicNOE’s between cH3a-6H8, cH3e-6H8,
cH3a-iH6,
andcH3e-6H6. AlthoughthecH3a-6H7andcH3e-6H7 NOE’s couldbe observed,the overlapofthe 6H7resonance withthe
6H5resonance preventedtheir quantification. The¿>H8-cH5
NOEwas detected, but its magnitude
(<1)
and the noisy baseline prevented its accurate determination.All
NOEcalculationswere done usingthe coordinatesof
the disaccharideo¡NeuAc(2-8)aNeuAc, whichcorrespondto residuesc-b ofthetrisaccharide (TableV). TheNOE
con-straintsmapswere similar totheones for colominicacid, and they did not showthepresenceofapreferred conformer in
solution (data notshown). However, the
( , )
ensembleav-eragedNOE’s,usingmodifiedenergy surfacesforthe gauche rotamers of 8, were able to reproduce adequately the
ex-perimental NOE’s. Relativeenergiesfor WG1+, WG2+, and WG3+ of 0.4:0:3 kcal/mol, equivalent to a population of
20:79:1, were found toincreasethe magnitudeofthe
inter-glycosidicNOE’scomparable tothose
of
the observedones.The
( , )
ensemble averagedNOEcalculationsforg'
andthe transrotamersof 8were performedusing thesame potentialenergy surfaces asforcolominicacid.
Discussion
The conformation
of
colominic acid has been previously discussed by a numberof
authors, and all have relied oninterpretation
of
NMR
results. Lindonet al. (1984) deter-mined,fromNMR
at 360MHz,thatH7andH8weregaucheandmolecularmechanicscalculationswere usedtoassessthe
rotational barriers about the C7-C8 bond.
All
staggered rotamerswere found tobeaccessiblewithbarriersofabout10kcal/mol. The13C
NMR
spin-lattice relaxation times,wereinterpreted as suggesting that the
a(2-8)NeuAc
poly-saccharidetumbled isotropically in solutionasarigidspecies.Michonetal.(1987) studied differencesin NOE’sat 500MHz between the polysaccharide and trisaccharide and from a
qualitative interpretationoftheNOE’sonly,
it
was proposedthatthe polysaccharide adoptedadifferent conformation from the trisaccharide. Recently, Yamasaki and Bacon (1991), fromaquantitativeanalysisof2DNOE’sat 500
MHz
usingrigid conformers, suggested that the polysaccharide adopts orderedhelical structures in solution.
Inour study,boththe results andtheir interpretationdiffer
fromthoseofYamasaki and Bacon (1991). Differentchemical
shiftsforthepolymerwere observed,whichcould possiblybe
attributed to sample preparation. While the H8 and the
downfieldH9resonance overlappedfor theirsample,theywere
resolvedinthis study. Thus, the overlapoftheH8 andH9
resonances prevented theseauthorsfromdetermining theC8 stereochemistry. Usingrigidmodels and steric constraints, they proposed ordered helical structuresforthefollowing
( , )
ranges:(-60°
to 0°, 115° to 175°) for 8 = 60°and (90°to 120°, 55° to 175°) for 8 =
-60°.
Their( , )
ranges correspond to thetwoareasintheNOEmap(Figure6), where
some NOEconstraints intersect the most. Frommodels,they estimated the numberofturns perhelix,n, tobe from 3 to
4 and the pitch (nh) to be from 9 to 11 Á. Their helical parameters were overestimatedby a factor of2, since it is difficultfrommolecular models alone to determine therotation
angle(µ) along thehelixaxis betweentworesidues,from which
=
2 /µ.
Valuesofngreater than2for 8 = 60° and alsongreater than2for 8 = -60° correspond totheir
( , )
ranges(Figure 2).
Inthis study,NOEdataat600
MHz
forthe polysaccharide andthe trisaccharidewere interpretedbyinvokingflexibilityabout the exocyclic linkages. Thebest
fit
oftheNOEdataforcolominic acid (Table
III,
Figure 5)was obtainedfroma
( , )
ensemble average over the whole energy surface. Proton couplingconstants andNOEdataindicatedthattheg+ rotamers for o>7 and 8 were predominant in solution.
However,flexibilityhad tobeinvoked about theC7-C8bond due toNOE’s which could onlyarisefrom other rotamers, most probably the g"one. From the determination of the proton couplingconstantsfortheH9resonances,allstaggered conformersforthehydroxymethylgroup at C9werepostulated
tobe accessible. Hence, dueto the widerange
of
conforma-tionswhichcan besampled,the polysaccharideissuggestedto exist predominantly as a random coilin solution. As Perez andVergelati (1985)notedintheirstudyof po-lysaccharidehelicalstructures, the spatialorganizationofthe polymerchaincan bedefined by consecutivefragments which
havewell-definedhelicalparameters. Theselocalhelicescan
alsobereferredtoapseudohelices. Compared to other hom-opolysaccharides,a(2-8)-linkedsialicacid ranksfirstinterms
of
displaying awide rangeof
helical structures (Figure 2). Obviously, thepresenceoffourbondsinthe linkage junction amplifiesthenumberofallowedconformationalstates,aswellasthe numberofhelicaltypeswhichcan begenerated. The helicalparametersforafew low-energy conformersare given
inTable
II.
Threehelicesfortheg+rotamersof
a>7and 8are shownin Figure8. Wheneverthevalues h = 0 or =
2 are crossed,thechirality ofthehelixchanges. The = 2
helixcorrespondstoazigzagplanarconfiguration, whileh =
0 correspondsto cyclic oligomers. Conformers withsmall values
of
h would foldbackon themselves after nresidues. For the 8 =g+ rotamer, most energetically favored
con-formerscouldformlocalhelices, since the h= 0linecrosses
a regionwhere values
of
nwere greater than4 (Figure2A).Also, high-orderhelicescan form with > 6,while forthe other 8 rotamersnisfrom2to3 only. For theg+rotamer
ofo>8inthe high-order helical region,asmall changeinlinkage
conformationleadstoalargechangein helicalparameters. Hence, the topological featuresofthe polymercanchangewith
little
changein energy. Suchproperties wouldbefavorable to topological rearrangements fromdisordered randomcoils tomore orderedconformations which promotetheformationofthehelicalepitope.
Forapolysaccharide,theNOE’s
will
beaveragedover allthe conformerssampledin solution.
If
a predominantcon-formeror apseudohelix existsin highenoughproportion, this
shouldbereflectedintheNOE’s,andaNOEconstraints map
could detectthe presence
of
apreferred conformer.If
morethanone conformer satisfied the constraints,or nopreferred
conformerwas found, then thepresenceofmolecularflexibility mustbeinvokedto explainthe observedNOE’s. Even
if
the NOE’s indicatethepresenceofapreferredconformerandthe motionalaveragingcalculationsstillsatisfytheexperimental conditions, motionalflexibility
cannot beruled out.For thecase ofcolominic acidandthe trisaccharide, the
NOE’sare bestexplained by motional averaging. A small proportionofconformerswas found togreatly influence the averaged NOE’s, since their NOE’swere ofmuch greater
magnitude than themajority
of
conformers,as can beseencon-Conformation ofMeningococcalB Polysaccharide Biochemistry, Vol. 31,No. 21, 1992 5003
figure 8: Side and top viewwithrespect to thehelixaxisforhelicesofa(2-8)NeuAc10 which
G2+, and G3+. The linkage conformationhavebeenslightlyaltered to giveintegralhelicesofr
for poly(A)isshownon top. Thecharged groupsinthe moleculesare representedas spheres.
formersGl+and G2+ (Tables
III
andV). Therelativeenergyof
the potential energy wells had to be altered in order to reproducethe observedNOE’s. Empiricalpotentials, which neglect molecularflexibility,electronic interactions, and solvent effect, are used only tocrudely estimatetheallowablecon-formationalspace. Therigid potential wouldalsooverestimate barrierstointerconversion. TheNOEdatacan thusbeused
totestdifferent population distributions calculatedfrom
em-pirical potentialenergy functions(Gumming
&
Carver, 1987). For colominic acid a relative population between wells WG1+ and WG2+ of 15:100 was found to reproduce the NOE’s correctly. Forthetrisaccharide,a ratioof25:100forthesame wellswas needed. Therelative population
of
well WG3+ to well WG2+ must remainlow(1-2%) in bothcasesor theHe3-H8wouldbecomelarger thantheH3a-H8 NOE.
Hence, differenceinlinkage conformation between thepolymer andshort oligosaccharidescan beexplainedasarisingfrom
slightchangesintherelative populationdistributionof
con-formersinsolution,asopposedto different rigidconformations
(Michonet al., 1987).
Theabovemodelfortheconformational behaviorof
colo-minicacid in solutionhasimportant biological implications. The cross-reactivityofIgMNOVwithboth thea(2-8)NeuAc
polymerand poly(A)can now berationalized(Kabatet al., 1988), sincethey bothcan adoptsimilarhelices (Figure2). From the crystal structure of trinucleoside diphosphate of
adenosine,amodelforthepoly(A) helixwas proposedwith
= 9andh = 2.82
Á
(Saenger et al., 1975). Energetically preferredconformers due tofavorablebasestackingcan form
helicesfrom = 8to12(Yathindra &Sundaralingam, 1976).
Therefore, helicesforthea(2-8)NeuAcpolymerandpoly(A)
can easilybegeneratedwhichhavesimilar helicalparameters. The = 9helixforboth polymersisshowin Figure8. There
isnot onlya similarityin the spatialdistributionofcharges betweenthecarboxylgroupsof a(2-8)-linkedsialicacid and the phosphate groupof poly(A) butalsoastrikingresemblance
betweenthehelical parameters ofboth polymers (Figure2).
Thisobservationwouldtendtosuggestthattheconformational
epitopeoccurs withinone turn
of
thehelixof
a(2-8)-linkedsialic acidorpoly(A). Sincethe energetically preferred helices
of poly(A) have more than 8 residues perhelicalturn,
it
isproposedthattheconformationalepitopeforthe<x(2-8)NeuAc
polymerprobablyoccurs withinahigh-order helix which has an extendedconformation (Figure8). However, there isso
farno indicationoftheexactnumberofresidueswithinthis helix that actually bind to group B polysaccharide specific antibodies.
Theunusuallength requirementof a(2-8)-linkedsialicacid oligosaccharides to bind to group B meningococcal poly-saccharidespecific antibodiescan alsoberationalized. Jen-nings et al. (1985) and Finne andMakela(1985)observedthat at leastadecasaccharidewas neededtoinhibitor bind tothese
antibodies. Also,forlarge oligomers
of
up to 17sialic acidresidues,theinhibitorypropertiesdidnotmaximizebutsteadily
progressedwithincreasingsize. Hayrinenetal.(1989) showed
thatthecriticalchain lengthforbindingwas 10units which contributed90%ofthebindingenergy. Thebinding forlonger oligomersalsosteadily progressedwithincreasingsizebutwith minor contributions tothebindingenergyfromtheadditional
residues. Becauseofthelimitationsofthesizeofan antibody
site(maximum6 or 7residues) (Kabat, 1960), these
obser-vations donot imply thattheantibodywas bindingtoasmany as 10 or more consecutive residues, rather that at least 10
residuesare requiredforthe recognitionsite toform.
If
the conformationalepitope corresponds toahigh-order helix with an extendedhelical conformation, as suggested fromcross-reactivityofIgMNOVtopoly(A),large oligosaccharideswould
be required in order for the epitope to form. Since poly-saccharides haveto undergotopologicalrearrangementsfrom disordered randomcoils tomore ordered conformations fa-vorable to theformation
of
thehelical epitopeand becauseBiochemistry,
unlikely that the epitope would occur in any short
oligo-saccharidesequence. Inadditionforthe decasaccharide, the
conformational behaviors
of
the reducingandnonreducing residuesare expected tobedifferent from that ofthe inner residues,asobservedforthetrisaccharideandcolominicacid (Michonet al., 1987). Thecontinued increasein bindingabove 10units couldbeattributed tothelength stabilizationofthe epitope, i.e.,theincreasedprobabilityof
the presenceofthe epitope.The poorimmunogenicityofthe groupBpolysaccharide probably arises from the recognition ofa helical structure
whichisalsoanintegralcomponentoftheneural cell adhesion molecule
N-CAM
(Finneet al., 1983). Despitestructuralmimicry withmammaliantissue,antibodies specificforthe
a(2-8)-linkedsialic acid homopolymercan beproducedand fromthe examples wherebindingstudies havebeenreported
(Jennings et al., 1985; Finne
&
Makela, 1985;Hayrinen et· al., 1989;Kabatet al., 1988) allare specificforthe extended helicalepitope. Becauseour conformationalstudiesindicate thatthis epitopeisonlyaminor contributorto thetotalnumberof epitopes available, we must therefore presume that the dominanceofthis epitopeintheimmuneresponsemustbethe resultofimmunologicalselection. Thefailureoftheimmune systemtoproduceantibodies specificforthemore populous
lowerorder helices presentin thepolysacchariderandomcoil probably occurs because these shorthelices are
conforma-tionallysimilarto theonesfound in shorter sialyloligomers.
Theseshorta(2-8)NeuAcoligosaccharidesare also present
in humantissue(Finneet al., 1983), andthe production of
specificantibodytothem appears tobeeven more stringently
avoided.
Acknowledgments
We thank FrancisMichonforcriticalreview ofthe
man-uscriptandRobertPonforhelp insamplepreparations. We
are grateful to Andy Byrd and
Bill
Eagan for access to apreliminarymanuscripton NOE analysis. References
Baleja,J.D.,Moult,J.,
&
Sykes, B. D. (1990) J.Magn.Reson 87, 375-384.Bodenhausen,G.,Kogler, H.,
&
Ernst, R. R. (1984) J.Magn. Reson. 58, 370-388.Borgias, B.A.,
&
James,T.L. (1988)J.Magn. Reson. 79, 493-512.Brisson, J. R.,
&
Carver, J. P. (1983) Biochemistry 22, 1362-1368.Burket,U.,
&
Allinger,N.
L. (1982)MolecularMechanics,ACSMonographSeries 177,American ChemicalSociety, Washington, DC.
Christian,R.,
&
Schulz, G. (1987) Carbohydr.Res. 162,1-11.Gumming, D. A.,
&
Carver,J. P. (1987) Biochemistry26, 6664-6676.Finne,
J„ &
Makela, P. H. (1985) J. Biol. Chem. 260, 1265-1270.Finne, J., Finne, V., Deagostini-Bazin, H.,
&
Goridis, C. (1983) Biochem. Biophys.Res. Commun. 112, 482-487. Flippen, J. L. (1973)Acta Crystallogr.B29, 1881-1886. Hayrinen, J., Bitter-Suerman, D.,&
Finne, J. (1989) Mol.Immunol. 26, 523-529.
Jennings, H. J. (1989) Microbiol. Immunol. 10, 151-165. Jennings,H.J. (1990) Curr. Top.Microbiol. Immunol.150,
97-127.
Jennings,H.J., Katzenellenbogen, E.,Lugowski,C.,Michon, F., Roy, R.,
&
Kasper,D. L. (1984) Pure Appl. Chem. 56, 893-905.Jennings,H.J.,Roy, R.,
&
Michon,F. (1985)J.Immunol.134, 2651-2657.
Kabat,E. A. (1960) J.Immunol. 84, 82-85.
Kabat,E.A., Liao,J.,Osserman, E.F., Gandan,A., Michon, F.,
&
Jennings,H.J. (1988)J.Exp.Med. 168,699-711. Kessler, H.,Griesinger,C.,Lautz,J.,Muller, A.,van Gun-steren,W.F.,&
Berendsen,H. J.C.(1988)J.Am. Chem. Soc. 110, 3393-3396.Landy,S. B.,
&
Rao, B.D. N. (1989)J.Magn.Reson. 83, 29-43.Lindon, J. C., Vinter, J. G., Lifely, M. R.,
&
Moreno, C. (1984) Carbohydr.Res. 133, 59-74.Michon,F.,Brisson,J.R.,
&
Jennings,H.J.(1987) Biochem.26, 8399-8405.
Perez, S.,
&
Vergelati,C. (1985)Biopolymers 24,1809-1822. Saenger,W.,Riecke, J.,&
Suck, D.(1975)J.Mol. Biol. 93,529-534.
Sugeta,H.,
&
Miyazawa, T. (1967)Biopolymers 5,763-679. Tvaroska, I.,&
Perez, S. (1986) Carbohydr. Res. 149,389-410.
Yamasaki, R.,
&
Bacon, B. (1991) Biochem. 30, 851-857.Yathindra,N.,