Design framework of the MuA remodeling signal that
confers preferential complex disassembly by the AAA+
unfoldase ClpX
by
Lorraine Ling
B.A. Molecular and Cell Biology,
University of California, Berkeley (2007)
Submitted to the Department of Biology
in partial fulfillment of the requirements for the degree of
Doctor of Philosophy
at the
MASSACHUSETTS INSTITUTE OF TECHNOLOGY
September 2014
© 2014 Lorraine Ling. All rights reserved.
The author hereby grants to MIT permission to reproduce and to
distribute publicly paper and electronic copies of this thesis document in
whole or in part in any medium now known or hereafter created.
Signature of Author . . . .
Department of Biology
July 22, 2014
Certified by. . . .
Tania A. Baker
E. C. Whitehead Professor of Biology
Thesis Supervisor
Accepted by . . . .
Michael Hemann
Associate Professor of Biology
Co-Chair, Biology Graduate Committee
Design framework of the MuA remodeling signal that confers
preferential complex disassembly by the AAA+ unfoldase ClpX
by
Lorraine Ling
Submitted to the Department of Biology on July 22, 2014, in partial fulfillment of the
requirements for the degree of Doctor of Philosophy
Abstract
The cell employs many classes of molecular chaperones to facillitate proteins in adopt-ing the proper structure and preventadopt-ing non-functional and potentially toxic non-native states. The Clp/Hsp100 family of ATPases are unfolding chaperones that remodel macro-molecular complexes and facilitate ATP-dependent protein degradation. They are mem-bers of the superfamily of AAA+ enzymes (ATPases Associated with various cellular Activities), which is conserved across all kingdoms of life. Efficient selection of multi-meric protein complexes over constituent subunits is key to successful remodeling and disassembly reactions. Using E.coli ClpX as a model for AAA+ ATPases, I characterized the mechanism by which ClpX discriminates between two oligomeric states of one of its natural multimeric substrates, phage MuA tranposase.
I elucidated many strategies for ClpX’s preference for the assembled Mu trans-pososome (MuA complex) over unassembled subunits. First, the target substrate makes multiple weak interactions with the AAA+ ATPase via the pore in the conserved AT-Pase domain and a class-specific auxiliary domain. Second, recognition tags should be at the weaker end of the affinity spectrum to allow effective synergy of multiple tags in the assembled complex. Third, multimeric complexes can "divide the labor" of making these interactions among their subunits. Thus the holistic complex-specific targeting sig-nal is accessible only in the assembled complex. The work of this thesis has provided a framework to understand the design of recognition signals that specify and target macro-molecular complexes to unfolding chaperones and remodelers of the AAA+ superfamily. Thesis Supervisor: Tania A. Baker
Acknowledgments
I am a confidant scientist today thanks in large part my advisor Tania Baker. She is my mentor and role model. Even as an established professor many years away from the bench, Tania answered my nitty-gritty questions and would offer technical help in my MuA assays. Her encyclopedic memory saved me many times from repeating experiments with negative/ uninterpretable results because hardly anyone publishes inconclusive data. Her guidance and enthusiasm help build my confidence as a researcher. She has helped me become a better scientific writer. As a role model, Tania has shown me how to handle stress and life’s complications with grace and perseverance. She is like the CEO of the Tania Baker company. I thank Bob Sauer, my co-advisor, for great advice at our monthly SJ meetings and reminding me to look at the bigger picture in my research.
Thank you to my thesis committee members, Amy Keating and Mike Laub, who have been with me since the start of my thesis research. They provided wonderful constructive criticism at all my thesis meetings, helped me pivot my project at a critical time, and ensured that I graduated in a timely manner. Thank you to Jodi Camberg for sitting on my thesis defense committee and suggesting improvements to this thesis.
As an undergraduate who majored in Genetics, biochemistry at the graduate level seemed intimidating. I thank all my biochemistry teachers at MIT who taught the subject in an engaging, thought-provoking, and accessible manner. Bob and Frank Solomon taught Graduate Biochemistry. Amy and Bob taught Special Topics in Biochemistry. They all helped me unlock my "biochemistry" power.
Many thanks to all past and present members of the Baker Lab. I am so lucky to be colleagues with this group of smart, witty, and compassionate people. Special thanks to fellow graduate student Ben Stein, with whom I shared the lab room. I will miss our ’Party-time’ music-science mash ups. Although I didn’t have the pleasure of overlapping with Aliaa whose research my thesis work has built upon, Aliaa was so generous answering my emails. I thank Anne Meyer who was my rotation mentor. She was uncertain, maybe even skeptical, that I would join the Baker lab due to my wildly different rotations but I did!
Lastly, thank you to my family and friends who provide a wonderful counter-balance to graduate school.
Contents
1 Introduction 11
1.1 Protein homeostasis in the cell . . . 12
1.2 Clp/Hsp100 ATPases . . . 17
1.3 Structural features of Clp/Hsp100 family . . . 20
1.3.1 ATPase domain . . . 20
1.3.2 Auxiliary domain . . . 22
1.4 Substrate selection by Clp/Hsp100 ATPases . . . 22
1.4.1 Direct recognition . . . 23
1.4.2 Assisted recognition . . . 25
1.5 Remodeling enzymes in AAA+ superfamily . . . 27
1.6 The virus, Bacteriophage Mu . . . 28
1.6.1 MuA transposase . . . 30
1.6.2 Transposition pathway . . . 30
1.6.3 Transpososome remodeling by ClpX . . . 33
1.7 Motivation for thesis research . . . 34
2 Design logic of a multivalent recognition signal confers preferential com-plex disassembly by the AAA+ unfoldase ClpX 37 2.1 Abstract . . . 38
2.2 Introduction . . . 39
2.3 Results . . . 43
2.3.1 Identification of a region critical for enhanced recognition of trans-pososomes by ClpX . . . 43
2.3.2 A peptide encompassing the critical region interacts with the
N-terminal zinc-binding domain of ClpX . . . 47
2.3.3 Step-wise loss of the Enhancement tag trends with ClpX’s weaker affinity for complexes . . . 49
2.3.4 Mu pore-binding tag is an intrinsically poor ClpX signal without adaptor-like contacts . . . 54
2.3.5 Transpososomes with a strong pore-binding tag do not require En-hancement tags . . . 54
2.4 Discussion . . . 58
2.5 Methods . . . 64
2.6 Appendix: Geometry experiments on MuA monomer variants . . . 67
2.6.1 Introduction . . . 67
2.6.2 Results . . . 68
2.6.3 Discussion . . . 69
3 Conclusion & Future Directions 71 3.1 Conclusion . . . 72
List of Figures
1-1 A simplified protein life cycle . . . 13
1-2 Prokaryotic Heat Shock Protein chaperones . . . 16
1-3 Model of AAA+ ATPase unfolding and translocation cycles . . . 18
1-4 Domain structure of bacterial Clp/Hsp100s . . . 19
1-5 ClpX structure . . . 20
1-6 Mechanism of replicative transposition . . . 29
1-7 Domain structure of MuA transposase . . . 30
1-8 Phage Mu in vivo replicative transposition . . . 33
1-9 Structure of type1 and type 2 Mu transpososomes . . . 34
2-1 In vitro assays for Mu complex assembly and recognition by ClpX . . . . 41
2-2 Mutation of a sequence region R622-S624 reduces disassembly and degra-dation rates . . . 44
2-3 Comparison of all reaction rates for Mu aspartate variants . . . 45
2-4 Residues P623 S624 form a critical interaction between MuA complex and ClpX . . . 46
2-5 The N-terminal zinc-binding domain of ClpX binds to the enhancement peptide . . . 48
2-6 Mu transpososome is an asymmetric complex . . . 50
2-7 Tools for making homogeneous mixed mutant complexes . . . 51
2-8 Chimeric complexes disassembly controls . . . 52
2-9 All four subunits can provide the Enhancement tag in MuA complexes . 53 2-10 Mu pore-binding tag is a weak ClpX recognition signal . . . 55
2-11 Mu complexes with a strong pore-binding tag are recognized as well as
native Mu complexes . . . 56
2-12 Mu∆8ssrA monomer degradation by ClpXP . . . 57
2-13 Permutations of tag engagement in Mu transpososome by ClpX . . . 61
Chapter 1
Introduction
Overview
The foundational theme of this thesis is understanding the mechanism and reg-ulation of substrate selection by the Hsp100/Clp chaperones. In the cell, chaperones are molecular machines that facilitate a change in a protein’s functions by altering the structure of proteins. Depending at which stage of the protein life cycle that the target protein is in, chaperones aid in both folding and unfolding of the target protein to bring about changes in structure.
This introduction will focus on the chaperones found in prokaryotes. The first section situates the many roles of chaperones within the protein life cycle. The next few sections introduce the Hsp100/Clp family of chaperones and chaperone-linked pro-teases found in E.coli, their structure, some interacting partners which aid in substrate recognition, and a sampling of biological targets. The last section introduces phage MuA transposase, one of many substrates of ClpX, an E.coli Hsp100/Clp unfolding chaper-one. The ClpX-MuA transposase interaction is my model system to study mechanisms of specificity and design of recognition signals.
1.1
Protein homeostasis in the cell
Inside the densely packed environment of a cell, proteins perform many roles; as structural macromolecules providing support and shape, as enzymes catalyzing chemical reactions, and as signaling molecules and receptors communicating information between the extra-cellular environment and the cell. For these diverse roles, proteins must adopt the correct structure leading to the biochemists’ axiom “structure equals function.” For ideal cellu-lar function, proteins must be properly folded into their native three-dimensional forms, associated with appropriate partners or oligomerize if required, and disposed of when no longer needed or damaged. Nature has evolved machinery, referred to as molecular chaperones, to guide proteins through these major milestones in the protein life cycle.
The life cycle of a protein begins at its birth by the protein-making factory called a ribosome, itself a protein-RNA complex. The newly synthesized polypeptide contains all the information needed within its primary amino acid sequence to define its final folded
Folded protein Protein Complex Unfolded protein Aggregated protein Degraded protein
Holdases
Clp/Hsp100
proteases
Clp/Hsp100
proteases
Hsp90
Clp/Hsp100
Hsp60
Hsp70
Hsp90
Clp/Hsp100
Hsp70
Clp/Hsp100
pr
ot
eases
Non-native stateClp/Hsp100
Hsp70
Figure 1-1: In this simplified protein life cycle, the upper-left represents on-pathway fold-ing and associations for a functional protein, which in this example forms a multiprotein complex. The lower-right represents off-pathway states due to stress, represented by yel-low lightning bolts. Many arrows were omitted for clarity; stress on protein complexes can lead directly to unfolded and non-native states. Actions of various HSP chaperones are indicated by their respective arrows to facilitate proteins adopting proper structure and to prevent or rescue proteins from the aggregated state. The Holdases belong to a diverse group of sHSP, small heat shock proteins, which bind to misfolded proteins and maintain them in a refolding competent state.
(native) structure (Anfinsen, 1973). However, in the crowded intracellular environment, nascent polypeptides need the help of molecular chaperones to properly fold and to avoid non-functional non-native and aggregated states (Figure 1-1). Chaperones do not mold proteins into their native structure but simply facilitate the self-directed folding process (Hartl, 2011; Mayer, 2013).
In many cases, proteins combine with other proteins to form large macromolecular complexes which are the biologically active states. Individual proteins within a larger complex are referred to as subunits. When these complexes need to perform new func-tions, the complex may gain, lose, or exchange subunits. These changes in quaternary structure very often reflect changes in biological function. Once again, chaperones fa-cilitate the assembly, alteration, and deactivation of macromolecular complexes. These multi-faceted processes performed by chaperones are encompassed in the term “remodel-ing.”
Several molecular chaperones participating at initial protein folding stages of the protein life cycle come from the highly conserved heat shock protein (Hsp) family, dis-covered in their roles in heat-shock response (Parsell & Lindquist, 1993). The Hsp60s group includes E.coli GroEL and its co-chaperone GroES (Figure 1-2A). Together they oligomerize into a barrel-shaped structure reminiscent of a basket with a domed lid to provide a defined and isolated environment for the polypeptide to fold and avoid aggre-gation prior to reaching its native state (Braig et al., 1994; Xu et al., 1997; Ellis, 2003). GroEL has a preference for binding stretches of hydrophobic amino acids, an elegant way to corral unfolded or non-native proteins (Fenton et al., 1994; Mayhew et al., 1996). Using non-specific substrate binding strategies and sequestration mechanisms, GroEL/GroES aids in folding of at least half of the proteins in the cell(Houry et al., 1999; Viitanen et al., 1992).
Another group of Hsp, Hsp70s also help proteins reach their native state. In E.coli, the Hsp70 member is DnaK (Figure 1-2B). Working with a co-chaperone,DnaJ, and a nucleotide exchange factor GrpE, the DnaK/ DnaJ/ GrpE chaperone team works at the site of the ribosome, to protect emerging polypeptides and help in folding (Deuerling et al., 1999; Teter et al., 1999). At this stage, these emerging polypeptides may have very
little folded structure, so stretches of exposed hydrophobic amino acids are particularly susceptible to aggregation. DnaK chaperone team binds non-specifically to stretches of hydrophobic residues, shielding them from neighbors. Then, powered by ATP, the chaperone releases the chain when it is ready to fold (Fourie et al., 1994; Flynn et al., 1989). Thus, both Hsp60s and Hsp70s use similar non-specific binding mechanisms to prevent inappropriate interactions and facilitate on-pathway folding.
Although not required for de novo folding of most proteins, the Hsp90s and Hsp100/Clp chaperones use the energy from ATP to aid the final maturation of selected proteins substrates ("clients") that function as a multicomponent or larger oligomeric complex. In most eukaryotes, Hsp90 is an essential chaperone and its clients include the steroid hormone receptors, protein kinases, and transcription factors (Li & Buch-ner, 2013). The growing list of Hsp90-associated co-factors enables the chaperone to have a broad substrate repertoire (Eckl & Richter, 2013). Clp/Hsp100 unfolding chap-erones facilitate protein complex remodeling and disassembly. Similar to Hsp90 group, Clp/Hsp100 chaperones utilize different co-factors to enlarge substrate range. However, many Clp/Hsp100 unfolding chaperones target just one protein complex; often remodel-ing these multiprotein complexes for the goal of recyclremodel-ing the subunits.
In addition to functions during normal growth, chaperones are key protective agents during times of stress. If proteins adopt non-native conformations due to damage or environmental stress, they may form insoluble aggregates which deactivate protein functions and are often toxic to the cell. Unfolding chaperones in the Hsp100/Clp group working as disaggregases can re-solubilize aggregates (Squires et al., 1991). Then the misfolded polypeptide has a chance to refold into its native conformation, often with help from Hsp60 and Hsp70 folding chaperones, or be degraded(McCarthy et al., 1998; Laskowska et al., 1996).
Lastly at the end of a protein’s useful “life”, proteins are degraded into their amino acid building blocks. Whether caused by regulatory responses or quality con-trol mechanisms, proteins which are at the end of their lives are funneled to intracel-lular chaperone-linked proteases also from the Hsp100/Clp group (Figure 1-2C). Unlike Hsp60s and Hsp70s, the Hsp100/Clp group of unfolding chaperones generally use specific
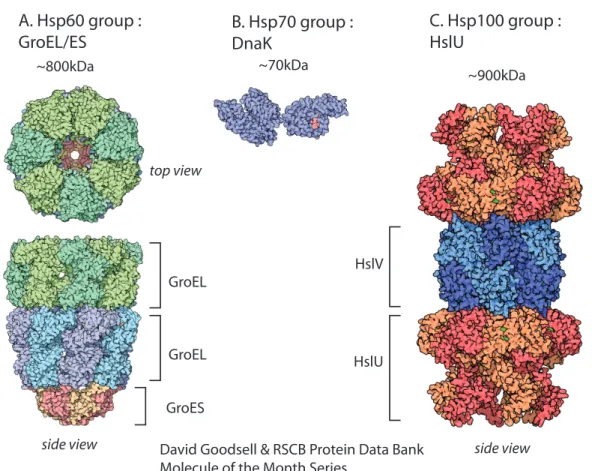
A. Hsp60 group :
GroEL/ES B. Hsp70 group : DnaK
C. Hsp100 group : HslU GroES GroEL HslV HslU GroEL ~800kDa ~70kDa ~900kDa top view
side view David Goodsell & RSCB Protein Data Bank side view
Molecule of the Month Series
Figure 1-2: Atomic structures of members of the Hsp family of chaperones illustrated by David Goodsell. Figures are not to scale.
A. Seven GroEL proteins (Hsp60 chaperone) form a ring. Depending on its nucleotide state, the double stacked GroEL rings are capped by a GroES particle (a 7-mer).
B. DnaK, ATP-binding domain on the left, peptide binding domain on the right with a bound peptide colored in red.
C. HslUV compartmentalized protease. HslU unfoldase forms a hexameric ring. HslV peptidase is a “double donut” of hexameric rings.
substrate selection mechanisms and actively unfold their substrates for two biological outcomes: remodeling and degradation. Because the Hsp100/Clp group deactivate com-plexes and often promote irreversible protein degradation, these destructive powers must be tightly regulated at multiple levels, such as target binding, spatial location, and de-velopmental timing.
1.2
Clp/Hsp100 ATPases
Clp/Hsp100 ATPase family belong to the larger AAA+ (ATPases associated with various cellular activities) superfamily of proteins. These enzymes use ATP hydrolysis to drive repetitive conformational changes that perform mechanical work in the cell (reviewed in Hanson & Whiteheart, 2005). Many AAA+ enzymes function by translocating protein polypeptides or nucleic-acid polymers. Examples include DNA/RNA helicases, protein-secretion translocation machinery, and viral packaging motors.
The Clp/Hsp100 ATPases actively unfold proteins for two biological outcomes: remodeling and degradation. Protein substrates are targeted to Clp ATPases by short peptide sequences, called tags or degrons (discussed in section 1.4.1). Cycles of ATP binding and hydrolysis drive protein unfolding (Figure 1-3A). For the outcome of degra-dation, the Clp/Hsp100 ATPase partners with a peptidase to form a compartmentalized protease (Figure 1-3B). The Clp/Hsp100 ATPases are ring hexamers, containing one or two ATPase modules per polypeptide (Figure 1-4). In E.coli, the first Clp family member identified was ClpA (Caseinolytic protease A) named for its role as the ATP-dependent regulatory subunit which partners with ClpP peptidase to form the ClpAP protease that degrades casein(Katayama et al., 1988). ClpP is unrelated to the AAA+ superfamily. Later a paralog of ClpA, ClpX was discovered which can also partner with ClpP to form ClpXP protease. Similarly, the HslUV protease is comprised of the ATPase Heat shock locus HslU (also known as ClpY) and the the peptidase HslV (also known as ClpQ). Two additional proteases, Lon and FtsH, have the ATPase and peptidase components encoded on a single polypeptide instead of existing as separate subunits (reviewed in Sauer & Baker, 2011). Lon was the first bacterial energy-dependent protease discovered.
A. Unfolding for remodeling/disassembly
tag AAA+ unfoldase ATP ATP AAA+ unfoldase peptidase ATP ATPB. Unfolding for degradation
Figure 1-3: A. A recognition signal (tag) in a native substrate is initially recognized by the AAA+ unfoldase. Repetitive cycles of ATP hydrolysis then power unfolding of substrates and translocation through the enzyme’s central channel. This leads to unfolding and/or remodeling of the protein complex. B. When the AAA+ unfoldase is associated with a compartmental peptidase, translocation of the polypeptide into the degradation chamber leads to protein destruction. (Adapted from Sauer and Baker, 2011)
Lon’s ability to degrade partially folded proteins has led to its designation as the major protease responsible for protein quality control in the cell (Chung & Goldberg, 1981). Although deletion of other AAA+ chaperones and proteases leads to severe pleiotropic phenotypes, FtsH is the only essential protease in E.coli (Ogura et al., 1991). FtsH is anchored to the inner membrane and degrades membrane and cytoplasmic proteins (Tomoyasu et al., 1993).
ClpB cannot partner with ClpP and thus has only chaperone and no coupled pro-tease functions. It is essential for thermotolerance and can solubilize almost any protein that becomes aggregated after severe stress (Squires et al., 1991). ClpB collaborates with the DnaK chaperone system (not members of AAA+ superfamily) to reactive proteins from insoluble aggregates. Studies of these chaperones’ activities in vitro have clarified
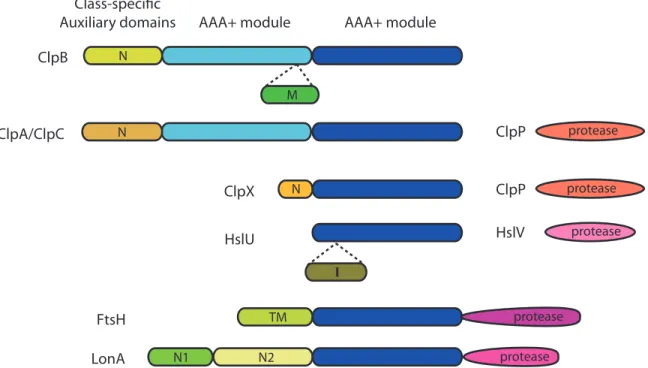
FtsH LonA HslU ClpX ClpA/ClpC ClpB ClpP ClpP HslV AAA+ module AAA+ module
protease protease protease protease protease N M N I N1 N2 TM N Class-specific Auxiliary domains
Figure 1-4: The Clp/Hsp100 family of chaperones and compartmentalized proteases con-tain a conserved ATPase domain, the hallmark feature of the AAA+ superfamily. ClpB, ClpA, and ClpC contain two AAA+ modules. ClpX, HslU, FtsH and Lon contain only one AAA+ module. Each ATPase has class-specific auxiliary domains that are not con-served. Proteases that associate with Clps are shown on the right. ClpP and HslV are separate proteins while FtsH and LonA each contain a protease domain. Adapted from (Sauer and Baker, 2011)
their individual mechanisms but a combined mechanism with wide consensus has not yet been established. One model is that ClpB breaks apart large protein aggregates into smaller ones by extracting and unfolding polypeptides from the aggregate (Weibezahn et al., 2004). The released polypeptide can spontaneously refold or interact with the folding chaperones such as DnaK or GroEL/GroES. DnaK may also act earlier in the ClpB-mediated remodeling reaction by helping ClpB bind to aggregates or regulating ATP hydrolysis by ClpB (Doyle & Wickner, 2009).
The Gram-positive bacterium, Bascillus subtilis, shares many Clp/Hsp100 or-thologs with E.coli. However B.subtilis has a species-specific ClpC ATPase and lacks both ClpA and ClpB. ClpC chaperone can associate with ClpP peptidase to form ClpCP protease (Molière & Turgay, 2009).
1.3
Structural features of Clp/Hsp100 family
1.3.1
ATPase domain
Clp/Hsp100 enzymes are members of the AAA+ superfamily. The hallmark feature of the AAA+ superfamily is a structurally conserved ATPase module, which performs cycles of nucleotide binding, hydrolysis and release to convert chemical energy into mechanical work. The ATPase domain contains a conserved ATP binding pocket. Two conserved sequence motifs which interact with ATP are the Walker A and Walker B motifs, which bind and hydrolyze ATP, respectively (Wendler et al., 2012). Most oligomerize into rings with a central pore.
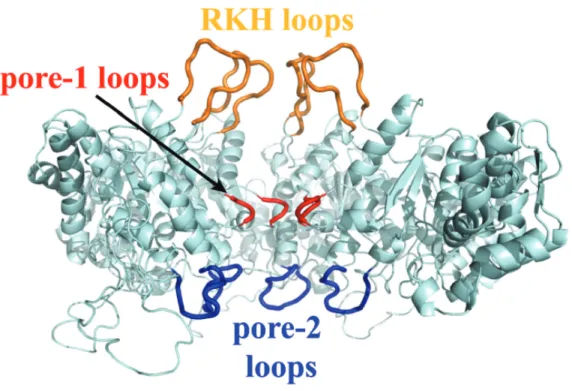
Lining the central pore are flexible sequences referred to as pore loops which bind substrates and translocate the polypeptide through the central channel (Figure 1-5).
Figure 1-5: Cutaway view of ClpX with pore loops highlighted. The RKH loops are colored yellow, pore-1 loops red, and pore-2 loops blue in a model of the ClpX hexamer (based on Kim et al. 2003, Bochtler et al. 2000). Three subunits of the hexamer were removed to allow visualization of the pore loops. Figure taken from Martin et al. 2007.
Located in the center of the channel, the pore-1 loops are the most conserved, with a nearly invariant Aromatic-hydrophic sequence motif, YVG, among all AAA+ ATPases. Mutagenesis studies and structures of ClpX with different nucleotide states argue that the pore-1 loop is key to translocation (Siddiqui et al., 2004; Martin et al., 2008b; Glynn et al., 2009). The RKH and pore-2 loops are conserved among ClpX orthologs and are located at the mouth of the channel and bottom of the channel, respectively. Both RKH and pore-2 loops play a role in initial binding and translocation of substrates (Farrell et al., 2007; Martin et al., 2007, 2008a). Pore-2 loops also mediate ClpP binding and communication (Martin et al., 2007).
Because there are no atomic structures of AAA+ ATPases with bound substrates, the mechanism of coupling ATP hydrolysis, enzyme conformation changes and mechani-cal work is under active investigation. Studies of ClpX have led to a model mechanism in which cycles of ATP hydrolysis are coupled to ClpX subunit conformational changes and thus the orientation of pore loops leading to a net movement of the substrate polypep-tide (Stinson et al., 2013). Crystal structures of HslU and ClpX reveal that the pore-1 loop adopts a dynamic range of confirmations that position the tip of the loop through-out the length of the channel (Bochtler et al., 2000; Sousa et al., 2000; Glynn et al., 2009). Repetitive cycles of ATP hydrolysis by ClpX may be needed to unfold the protein and translocate the polypeptide(Martin et al., 2008c). Translocation of the polypep-tide is proposed to pull the attached folded protein against the entrance to the axial pore, thereby generating a denaturing force because the pore is smaller than the folded protein (Baker & Sauer, 2006). The observable result is that Clp/Hsp100 chaperones and chaperone-linked proteases exert a pulling force on the polypeptide which leads to cooperative unfolding of the target protein (Aubin-Tam et al., 2011).
There appears to be no obligatory directionality to translocation, as ClpXP can degrade substrates starting either from the N-terminus or from the C-terminus (Gottes-man et al., 1998; Gonciarz-Swiatek et al., 1999; Lee et al., 2001; Flynn et al., 2003; Hoskins et al., 2002; Kenniston et al., 2005; Farrell et al., 2007). Once ClpX engages with the protein substrate, the translocation process is processive and remarkably toler-ant to the structural identity of the polypeptide. ClpX exhibits very little preference for
side group size, charge, or chirality and will translocate peptide bonds spaced with long hydrophobic carbon chains (Barkow et al., 2009).
1.3.2
Auxiliary domain
A second feature of Clp/Hsp100 ATPases are auxiliary domains specific to each member. In general these auxiliary domains are not conserved between members, dispensable for ATPase function but play roles in substrate recognition (Sauer & Baker, 2011). Often these member-specific domains provide a docking platform for delivery proteins, called adaptors (discussed in section 1.4.2). ClpA, ClpC, and ClpX all have N-terminal auxiliary domains. However the structure of each auxiliary domain and the connection of each to their respective ATPase module differs among the three Clp ATPases (Zeth et al., 2002; Wang et al., 2011; Park et al., 2007) HslU has an I-domain, which is an insertion of 140 residues in the AAA+ module (Bochtler et al., 2000). The I-domain domain is thought to play a role in substrate binding and allosterically regulates ATPase activity (Sundar et al., 2012). The M-domain (middle domain) of ClpB forms a propeller-shaped coiled-coil and has been proposed to act like a “crowbar” to break apart large aggregates, though the crowbar model has not been experimentally validated (Lee et al., 2003). Interestingly, the M-domain serves as the site for species-specific interaction with the DnaK/DnaJ/GrpE chaperone team (Miot et al., 2011).
1.4
Substrate selection by Clp/Hsp100 ATPases
Clp ATPases employ two modes of substrate recognition. The first is direct substrate recognition by binding to short peptide sequences, known as tags, on the substrate. The tag interacts with the pore of the ATPase and subsequent engagement results in un-folding. The second mode termed “assisted recognition” is when another protein aids the ATPase in recognition of the substrate. Additional mechanisms for substrate selec-tion not discussed below are subcellular relocalizaselec-tion of either substrates or Clp/Hsp100 ATPases, phosphorylation of proteins turning them into substrates, and regulating ex-pression levels of substrates or ATPases by developmental timing.
1.4.1
Direct recognition
Recognition tags are found often at the N- or C-termini of substrates. Tags that bind directly to the unfoldase fall into three classes: intrinsic, latent, and co-translational. intrinsic class
Intrinsic signals are encoded in the primary sequence of a substrate protein. The tags are often present at the N or C-termini but may not be accessible to proteases until a conformational change or loss of a shielding binding partner. Examples of substrates with N-terminal tags include Dps (DNA-binding protein from starved cells), a stationary phase nucleoid protein that sequesters iron and protects DNA from damage (Flynn et al., 2003), and bacteriophage 𝜆 O replication initiator protein when not bound to orilambda DNA (Gonciarz-swiatek et al., 1999). Additionally, 𝜆O may have an internal intrinsic signal. Residues Q49-M67 bind strongly to ClpX N-domain and a peptide containing this sequence can compete with degradation of the full-length protein by ClpXP (Thibault et al., 2006).
Remarkably the stability of a protein can be attributed to a single amino acid at its N-terminus, known as the N-end rule. Present in both prokaryotes and eukaryotes, the N-end rule tags are often the large hydrophobic residues. In E.coli, an N-terminal leucine, phenylalanine, tryptophan or tyrosine directly targets proteins to intracellular proteases (Varshavsky, 1996).
An example of an intrinsic C-terminal tag comes from Supressor of lon (SulA) identified in a screen to suppress the lon- sensitivity to ultraviolet radiation (Gayda
et al., 1976). SulA is an inhibitor of cell division and upregulated during SOS response otherwise during normal growth it is degraded by Lon and HslUV proteases (Gottesman et al., 1981; Mizusawa, 1983; Seong et al., 1999). The last eight C-terminal residues are crucial for recognition by Lon as a truncated SulA variant was stabilized both in vivo ans in vitro. (Higashitani et al., 1997)
A proteomic based screen for in vivo substrates of ClpXP revealed five classes of recognition signals. The screen utilized a tagged and catalytically inactive variant of ClpP (ClpPtrap) to receive and contain proteins translocated by ClpX (Flynn et al.,
2003). Analysis of the >50 captured proteins led to two classes of motifs located at the C-terminus and three classes located at the N-terminus. About one quarter of the trapped proteins contain potential intrinsic ClpX recognition signals at both the terminus and C-terminus. Similarly to phage 𝜆 O replication initiator protein, many N-terminal sequences from the pool of trapped proteins directly bound to ClpX’s N-domain on a peptide array. However with so few examples of confirmed N-domain interacting sequences, it was difficult to establish a consensus motif (Flynn et al., 2003). It is unclear whether both signals are engaged by the ATPase domain of ClpX or contribute to additional enzyme-binding interactions via the auxiliary domain similarly to adaptor proteins (see section 1.4.2).
latent class
Latent signals are also encoded in the primary sequence but require processing of the protein to make the tag accessible. Often an endopeptidic cleavage reveals a new termini containing the tag. Examples include LexA and RseA. LexA is a transcriptional repressor of genes involved in DNA-damage response. During the SOS response to DNA damage, LexA undergoes RecA-stimulated autocleavage between the N-terminal DNA binding domain and the C-terminal dimerization domain. Both fragments are rapidly degraded by Lon and ClpXP (Little, 1983; Neher et al., 2003a). The cleavage reveals a tag at the new C-terminus with residues VAA-CO2 which is similar to the region of the
ssrA tag (LAA-CO2discussed below) recognized by ClpX (Neher et al., 2003a).
RseA is the anti-sigma factor to 𝜎E which activates expression of genes involved
in the bacterial extracytoplasmic stress response. RseA is an inner membrane-spanning protein. The N-terminal cytoplasmic portion binds 𝜎E and sequesters the sigma
fac-tor from its target promoters. In response to extracytoplasmic stress, RseA undergoes sequential cleavage steps by proteases in the periplasm and cytoplasm which results in an N-terminal fragment of RseA and frees 𝜎E (Lima et al., 2013). The released 𝜎E can
activate its regulon, while the N-terminal fragment of RseA, revealing residues VAA at the new C-terminus, is degraded by ClpXP (Flynn et al., 2004).
In contrast to the two previous classes, the co-translation class is not encoded in the primary sequence of the substrate protein. The only member of this class is the ssrA tag which marks proteins for degradation by multiple AAA+ proteases (ClpAP, ClpXP, FtsH, Lon) (Keiler et al., 1996). When ribosomes get stuck on an mRNA, a rescue system of tmRNA (trans-messenger RNA) displaces the offending mRNA. Translation continues on the tmRNA, which encodes an eleven amino acid sequence (AANDENYALAA) and a stop codon. The ssrA tag is appended onto the truncated polypeptide and marks these incomplete translation products for destruction. It is estimated that 0.5% of translation products receive an ssrA tag (Lies & Maurizi, 2008). Thus the ssrA-tagging system rescues stalled ribosomes and destroys potentially dangerous protein fragments (reviewed in Karzai et al., 2000).
1.4.2
Assisted recognition
Clp ATPases also use a second mode termed “assisted recognition” in which accessory proteins called adaptor proteins modulate substrate choice and often give rise to higher-affinity enzyme-adaptor-substrate complexes. Various mechanisms for adaptors have been observed from acting as delivery vehicles to directly affecting chaperone and pro-tease activity (reviewed in Kirstein et al., 2009). Adaptor proteins are themselves not degradation substrates and thus participate in multiple rounds of delivery or modulation of enzyme activity. A general overview of three E.coli adaptors (SspB, RssB, ClpS) and one B.subtilis adaptor (MecA) represents the diversity of mechanisms to deliver target proteins and modulate chaperone-protease activities yet highlights one common theme of docking to the enzyme’s auxiliary domain.
SspB
Stringent starvation protein B (SspB) is the best characterized E.coli adaptor protein. SspB enhances degradation of ssrA-tagged substrates by ClpXP via a tethering mechanism. Here, the adaptor binds to both the substrate and the AAA+ unfoldase to increase the effective concentration of the tag near the enzyme’s active center (Wah et al., 2003). SspB binds to ClpX specifically through the unfoldase’s N-domain (Park
et al., 2007). In E.coli, ClpX and SspB bind distinct portions of the ssrA tag. ClpX recognizes the last three residues and the carboxyl group whereas SspB recognizes the first four and seventh residue (Levchenko et al., 2000; Flynn et al., 2001; Levchenko et al., 2003; Song & Eck, 2003). As mentioned above, ssrA-tagged substrates are also degraded by ClpAP protease. However, the same adaptor SspB inhibits this reaction because ClpA and SspB bind overlapping residues in the ssrA tag (Flynn et al., 2001). Although the biological consequence of this inhibition remains unclear, a plausible outcome is to turn other substrates into high-priority ClpAP targets, leaving ClpXP to clean up ssrA-tagged polypeptides. SspB also recognizes and delivers the N-terminal fragment of RseA to ClpXP for degradation (Flynn et al., 2004).
RssB
Regulator of sigma-S protein B (RssB) also known as stationary-phase regulator (SprE) is essential for turnover of stationary phase sigma factor, 𝜎S (Muffler et al., 1996;
Pratt & Silhavy, 1996). RssB is a ClpXP-specific adaptor and is activated by phospho-rylation. Phospho-RssB protein binds 𝜎S which exposes a latent tag in the N-terminal
region of 𝜎S.
ClpS
ClpS adaptor delivers N-end rule substrates to ClpAP protease for degradation (Dougan et al., 2002). Because both ClpS and ClpA recognize the same N-terminal amino acid, ClpS employs a more involved mechanism than simple tethering. ClpA may recognize additional tag features that are not directly bound to ClpS. ClpA, ClpS and the N-end rule substrate form a high-affinity ternary complex (Román-Hernández et al., 2011). Within this complex, ClpA engages the unstructured N-terminal section of ClpS which causes a conformational change and hand-off of the N-end rule residue from the ClpS binding pocket to the pore of ClpA (personal communication Izarys Rivera-Rivera). ClpS escapes degradation to catalyze another round of delivery because the core substrate binding domain of ClpS carries structural elements that are non-denaturable by ClpA (personal communication Izarys Rivera-Rivera). ClpS interacts with ClpA via the unfoldase’s N-terminal domain. This interaction is necessary for delivery of N-end
rule substrates (Zeth et al., 2002). MecA
Medium-independent expression of competence (MecA), the best characterized adaptor protein of B.subtilis, was identified in a screen for genes involved in regulation of competence (Dubnau & Roggiani, 1990). MecA binds and inhibits the transcriptional activator of competence, ComK. Furthermore MecA targets ComK for degradation by ClpCP. Unlike the previous adaptor examples, MecA has a unique mechanism to modu-late ClpC activity. ClpC is an inactive monomer on it own. MecA triggers the oligomer-ization of ClpC into the active hexameric chaperone which then can associate with ClpP to form an active protease. This adapter-mediated oligomerization requires MecA to bind to the N-domain of ClpC (Kirstein et al., 2006).
1.5
Remodeling enzymes in AAA+ superfamily
In this section, four examples of remodeling enzymes in the AAA+superfamily showcase the breadth of important biological transitions promoted by remodeling. Katanin and Spastin are eukaryotic AAA+ ATPases which remodel microtubules (McNally & Vale, 1993). Microtubules provide support to organelles, shape the cell, and organize into a distinct structure called the spindle that is essential for cell replication and division. They are made up of tubulin subunits, which polymerize into long and branched dynamic polymers. Many factors regulate microtubule assembly and disassembly at the termini of polymers (reviewed in Gardner et al., 2013). However, Katanin and Spastin modulate the dynamics of microtubule from the middle of a polymer. These “microtubule severing enzymes” preferentially unfold and abstract tubulin subunits from the lattice (Roll-Mecak & McNally, 2010). This process of microtubule severing requires the C-terminal tails of tubulin and and the pore loops of Katanin and Spastin (White & Lauring, 2007; Roll-Mecak & Vale, 2008).
Another eukaryotic AAA+ family member, N-ethylmalemide sensitive fusion pro-tein (NSF) is an essential factor in intracellular membrane trafficking. NSF in concert with SNAPs (soluble NSF attachment proteins) disassemble SNARE complexes, thus
freeing SNARE subunits for additional rounds of membrane fusion (Whiteheart et al., 2001).
Although most studied for their functions in intracellular proteolysis, both E.coli ClpA and ClpX have been observed in ClpP-independent remodeling reactions. In vitro, ClpA activates the phage P1 replication initiator protein RepA by remodeling inactive RepA dimers into monomers that are competent to bind DNA (Wickner et al., 1994).
ClpX plays a key role in the phage Mu lytic cycle (Mhammedi-Alaoui et al., 1994; Kruklitis et al., 1996). The biochemical steps in phage Mu transposition and the remodeling of the Mu transposase-DNA complex by ClpX have been extensively studied and are summarized in the next section. This well-characterized protein has provided me an ideal model substrate to address questions of target specificity and design principles of ClpX recognition signals.
1.6
The virus, Bacteriophage Mu
Bacteriophage Mu is a virus that propagates its genome within a bacterial host using a mechanism of replicative transposition. In replicative transposition, the mobile DNA element cuts, copies itself to a new location, and leaves behind the copy at the previous genomic location. This movement occurs using a branched DNA intermediate called a Shapiro structure (Shapiro, 1979). Extensive study of phage Mu has led to a deeper understanding of the molecular mechanism and regulation of transposition by mobile DNA elements. Phage Mu encodes two proteins necessary for in vivo transposition, MuA and MuB. MuA is the transposase and MuB is a regulatory factor. Together they recombine the correct DNA sequences, transpose at the correct time during the phage lifecycle, and avoid disrupting phage Mu’s own genome (Figure 1-6).
Phage Mu regulates this sequence of events by evolving a vectorial process that uses increasingly stable nucleoprotein complexes and co-opting a host chaperone to direct a key transition point. A highly tractable in vitro system was developed early which allowed for biochemical analysis of the transposition reaction. This biochemical system minimally contains the sequences of Mu genomic ends on a supercoiled plasmid, MuA,
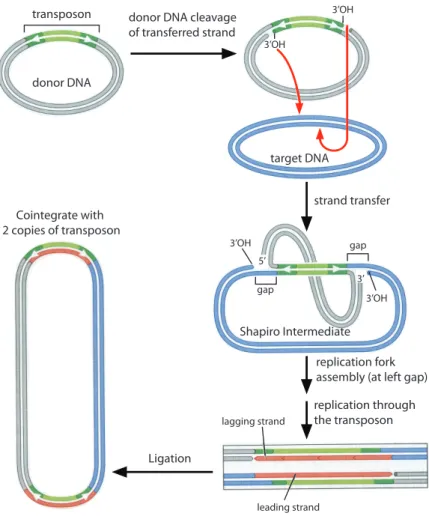
transposon
donor DNA
target DNA
3’OH
3’OH
donor DNA cleavage of transferred strand
strand transfer
replication fork assembly (at left gap) replication through the transposon Ligation Cointegrate with 2 copies of transposon 5’ 3’OH 3’ 3’OH lagging strand leading strand Shapiro Intermediate gap gap
Figure 1-6: The transpososome introduces a single-strand nick at each end of the ends of the transposon DNA (green). The liberated 3’OH groups then attack the target DNA and become joined to the target by DNA strand transfer. At each end of the transposon, only one strand is transferred into the target at this point, resulting in the formation of a doubly-branched DNA structure, the Shapiro intermediate. The replication apparatus assembles at one of these "forks" (the left one in this figure). Replication continues through the transposon sequence. The resulting product, called a cointegrate, has the two starting circular DNA molecules joined by two copies of the transposon. The ssDNA gaps in the branched intermediate give rise to the target site duplications. These duplications are not shown in the cointegrate for clarity. (Adapted from Figure 11-22 of Watson et al. Molecular Biology of the Gene, 6th ed.)
the phage-encoded transposase, host-encoded DNA bending proteins and divalent metal ions (Craigie et al., 1985). Although not required for the core steps of transposition, MuB protein as a key regulator helps ensure transposition into sensible target DNA sites (Reyes et al., 1987).
1.6.1
MuA transposase
MuA transposase is part of the DDE family of recombinase enzymes which include HIV integrase, Tn5 transposase, and RSV integrase (Rice & Baker, 2001). MuA transposase has a mass of 75,000Da and is monomeric in the absence of phage DNA (Baker & Mizu-uchi, 1992). It is organized into three structurally and functionally distinct domains: a DNA binding domain, a catalytic domain, and regulatory domain (Figure 1-7).
Figure 1-7: MuA transposase contains three domains. Domain I binds various phage DNA sequences. Domain II contains the catalytic residues, DDE. Domain III contains interaction sites for MuB, a regulator factor, and ClpX, a host unfolding chaperone.
Domain I is responsible for site-specific binding to repeat DNA sequences at the ends of the Mu genome. The structure of this domain has a winged helix-turn-helix motif(Clubb et al., 1994). Domain II contains the catalytic DDE motif characteristic of the family (Rice & Mizuuchi, 1995). The active site has dual functions of DNA cleavage and DNA rejoining when the protein is assembled into an active tetrameric enzyme as part of a DNA-protein complex (Lavoie et al., 1991; Mizuuchi et al., 1992). Domain III is the site of interaction with allosteric and regulatory factors such as MuB and the host chaperone, ClpX (Baker et al., 1991; Wu & Chaconas, 1994; Levchenko et al., 1997).
1.6.2
Transposition pathway
MuA transposase recombines phage DNA in a reaction pathway characterized by distinct nucleoprotein complexes called transpososomes (Surette et al., 1987; Craigie & Mizuuchi, 1987). MuA binds to DNA attachment sites located at the left and right ends of the phage genome (Craigie et al., 1984). With additional transient binding to an internal enhancer element and help from host histone-like proteins to severely kink the DNA,
MuA subunits bring the DNA ends together to form the stable synaptic complex (SSC, Figure 1-8B)(Surette & Chaconas, 1992; Mizuuchi et al., 1992). The SSC consists of four MuA subunits with extensive interprotein contacts.
To initiate movement to a new host genomic site (target DNA), MuA transposase cleaves the DNA at the junction of the Mu genome and flanking host DNA to generate 3’ hydroxyl groups at nicked ends. This is the next form of transpososome called the cleaved donor complex (CDC, Figure 1-8C)(Craigie & Mizuuchi, 1987; Surette et al., 1987; Lavoie et al., 1991; Yuan et al., 2005). The freed 3’ hydroxyl groups then attack and join opposite strands of target DNA in a step called DNA strand transfer (Mizuuchi & Adzuma, 1991). This generates recombined DNA synapsed with MuA transposase and the next transpososome called the strand transfer complex (Figure 1-8D) (Surette et al., 1987; Mizuuchi & Adzuma, 1991; Montaño et al., 2012). Structures of the CDC and STC reveal greater interprotein contacts are made as the Mu transpososome progresses through the reaction pathway (Figure 1-9).
Selection of the target DNA is regulated by MuB. MuB binds A/T rich target DNA in an ATP-dependent manner and stimulates MuA to catalyze transposition into bound DNA (Baker et al., 1991; Surette et al., 1991; Yamauchi & Baker, 1998). In opposition, MuA stimulates MuB’s ATPase activity. As a result, MuB tends to dissociate near Mu genomic ends where MuA is bound and to associate with DNA far away from the Mu genome (Maxwell et al., 1987; Greene & Mizuuchi, 2002a,b,c). Thus the interactions between MuA and MuB not only promote catalysis but also prevent disruption of its own viral genome, in a phenomenon known as target immunity(Reyes et al., 1987). It is currently unclear what is the molecular mechanism which determines the outcome of the MuA-MuB interaction. In vitro, the minimal system does not require MuB to observe transposition and formation of the STC.
After strand transfer, the enzyme has finished its function of recombination but the STC is so stable that the enzyme does not turn over. In vitro, the hyperstable STC resists temperatures up to 75°C and 6M urea (Surette et al., 1987). The stable STC holds onto the recombined DNA products and blocks replication. In fact, continued presence of MuA transposase on the recombined DNA inhibits recruitment of host DNA replication
Bacterial genome integrated Mu genome
target site DNA MuA
MuB
original Mu genome Polymerase
replicated Mu genome
Stable Synaptic Complex
(SSC)
Cleaved Donor Complex
(CDC)
Strand Transfer Complex
(STC)
remodeled
fragile complex
ClpX remodelling
left end right end
replication replication machinery A B C D E F G H
machinery and thus lytic growth (Nakai & Kruklitis, 1995). At this critical transition, phage Mu switches from its own proteins to ClpX, a host-encoded chaperone, to resolve the replication block (Figure 1-8E).
1.6.3
Transpososome remodeling by ClpX
ClpX remodels the stable STC into a fragile complex which then recruits host replication machinery to complete amplification of the phage genome (Figure 1-8F) (Levchenko et al., 1995; Kruklitis et al., 1996; Jones et al., 1998). Deleting ClpX inhibits phage Mu replication in vivo almost completely, but deleting ClpP has almost no effect (Mhammedi-Alaoui et al., 1994). In fact full length ClpX is required for in vivo Mu replication as ClpX lacking its N-terminal zinc-binding domain could not support phage lytic growth (Wojtyra et al., 2003). Thus, it is the chaperone rather than degradation function that is necessary for the Mu lytic cycle. ClpP is irrelevant for transpososome remodeling in vivo. Purification of a host factor that enabled transpososome remodeling led to a single protein fraction with high-specific activity which turned out to be ClpX (Levchenko et al., 1995; Kruklitis et al., 1996).
Further in vitro studies of purified ClpX enzyme and transpososomes showed that ClpX was sufficient to disassemble the stable STC and that the last eight C-terminal residues comprised an intrinsic recognition tag (Levchenko et al., 1997). Transpososomes biased to have only one subunit with a tag were sufficient to be destabilized by ClpX (Burton et al., 2001). Highlighting that the unfolding process and not degradation of a Figure 1-8 (preceding page): The in vivo replicative transposition of phage Mu begins with an integrated Mu genome and proceeds through multiple nucleoprotein complexes called transpososomes (A). MuA transposase binds to the left and right ends and brings them together to from the SSC while MuB binds to target site DNA (B). MuA trans-posase cleaves the DNA to form the CDC while MuB brings the target site closer (C). Recombination into target DNA occurs to form the STC (D). ClpX remodels the hy-perstable STC into a fragile complex by unfolding a MuA subunit (E). This remodeled fragile complex then recruits host replication machinery and remaining MuA subunits are released (F). Replication of phage Mu DNA (G) results in two copies of the Mu genome integrated into the bacterial chromosome (H) and the transposition-replication cycle repeats.
Cleaved Donor Complex Strand Transfer Complex
Figure 1-9: left: Type 1 transpososome, the CDC (cleaved donor complex). An EM structure with MuA subunits colored, DNA in gray. Figure from Yuan et al. 2005. right: Type 2 transpososome, the STC (strand transfer complex). A crystal structure with MuA subunits colored, DNA in gray. PDB: 4FCY
MuA subunit was key to remodeling, Burton and coworkers showed that STCs assembled from a MuA variant with an alternative tag (ssrA) at the C-terminus were disassembled by ClpX. Even using an alternative chaperone (ClpA), these alternative ssrA-tagged STCs could be destabilized (Burton & Baker, 2003). Since both free MuA monomers and assembled MuA complexes are in the bacterial cytosol, a pertinent question was how ClpX distinguished between the two oligomeric states of MuA and directed its unfolding activity to the biologically relevant target, the STC.
1.7
Motivation for thesis research
AAA+ ATPases use the energy from ATP binding and hydrolysis to drive diverse cellular activities such as DNA replication by helicases and cargo transport along microtubules by dynein. For many of these processes, the ATPase causes key alterations to the struc-ture of the target protein and thus its function. Furthermore, the biologically relevant substrates for some ATPases are large multiprotein structures or a complex in a higher-ordered oligomeric state. As a consequence, the action of unfolding chaperones must be directed away from subunits or monomers because the cellular process requires a change in structure/function of the macromolecular complex. Thus, it’s important to
understand how these enzymes discriminate and prioritize macromolecular complexes over constituent subunits.
Using the extensively characterized Mu transpososome remodeling process, I elu-cidated at the molecular level, the intrinsic recognition signal in transpososomes evolved for disassembly. Then, I uncovered a mechanism used to discriminate between assembled MuA complexes and constituent subunits. Additionally, I articulated the different roles for the multiple intrinsic recognition tags in MuA transposase. Lastly, I engineered MuA variants with different recognition tags to probe the effect of the tags themselves and of the architecture of MuA complex on the holistic “ClpX remodeling” signal. Through this work I present an underlying design framework for how AAA+ enzymes achieve speci-ficity for macromolecular complexes, the biologically relevant targets of remodeling and disassembly reactions.
Chapter 2
Design logic of a multivalent
recognition signal confers preferential
complex disassembly by the AAA+
unfoldase ClpX
This chapter has been written as a manuscript for publication. A draft is currently being reviewed by collaborators. I performed all experiments for all figures except Figure2-10, which was contributed by A. Abdelhakim. I.Levchenko synthesized the peptides used in Figure 2-5. S.P. Montano and P.A. Rice designed and cloned the initial Sin15Mu chimeric protein and Sin-attachment site DNA oligo, both of which I modified for research in this chapter.
2.1
Abstract
AAA+ enzymes are present in all kingdoms and use the chemical energy of ATP to remodel protein complexes and catalyze substrate protein unfolding. How these power-ful enzymes recognize protein complexes and aggregates is poorly understood. Efficient selection of multimeric protein complexes over constituent subunits is key to success-ful disassembly. Here, we use E.coli ClpX, a AAA+ unfoldase, and the tetrameric MuA transpososome, to investigate how preferential specificity for an assembled com-plex is achieved. We demonstrate that the MuA tetramer employs a multivalent set of recognition peptides to ensure that the complex has the tightest affinity for ClpX. The critical recognition components are the weak ClpX pore-binding peptide at the C-terminus of MuA and a second peptide ∼ 40 residues away from the pore signal that binds the ClpX N-terminal domain. By constructing chimeric SinMuA proteins with altered DNA-binding specificity we investigated multiple variant complexes carrying dif-ferent geometries of mutant signals and determined how each subunit contributes to complex-specific recognition. Although individually, the two key recognition peptides bind weakly (70 − 400𝜇M) to ClpX , together within the assembled MuA tetramer they impart an affinity of ∼1𝜇M. All four subunits in the tetramer can donate the ClpX N-domain-binding peptide and optimal recognition is achieved when all four are present. In contrast, only two specifically located subunits can donate the pore-binding signal. Im-portantly, the N-domain-binding peptides become unnecessary for complex recognition when the native weak binding signal is replaced with a much stronger compact pore-binding tag. Thus, we conclude that the design of signals that are specific for assembled complexes depends on collaboration between multiple weak protein-unfoldase-interaction peptides and that strong-binding signals can prevent multimer-specific recognition.
2.2
Introduction
Cells are densely packed with proteins performing structural and/or enzymatic roles es-sential for life. To help respond to environmental changes, manage protein turnover and protein quality control, these cells employ energy-dependent unfoldases/disaggregases and proteases from the AAA+ family (ATPases associated with various cellular activ-ities). Powered by cycles of nucleotide binding, hydrolysis, and release, these ATPases remodel complexes, solubilize aggregates, and degrade proteins (when coupled with part-ner peptidases). E. coli ClpX is arguably the best-characterized AAA+ unfoldase and is known to disassemble complexes and unfolds proteins (Sauer & Baker, 2011). ClpX acts alone as a protein-remodeling enzyme as well as in complex with ClpP peptidase to make the ClpXP protease. In ClpXP, ClpX recognizes and unfolds many substrate proteins and translocates the unfolded chain to ClpP peptidase where it is degraded. Because of its destructive power, ClpX’s selection of substrates must be exquisite. The protein signals (recognition sequences) and design logic governing recognition of different classes of substrates is being actively investigated.
Most bacteria have no ubiquitination system and recognition of protein targets for degradation or disassembly is mediated by a diverse set of short unstructured peptide sequences, or tags. These recognition tags are often located at the termini of otherwise native substrate proteins (Sauer et al., 2004). Examples of substrates with N-terminal recognition tags are 𝜆O, a DNA replication origin-binding protein from phage 𝜆, and UmuD, a subunit of DNA polymerase V, a DNA-repair/tolerance polymerase (Gonciarz-swiatek et al., 1999; Gonzalez et al., 2000) and proteins recognized by the N-end rule pathway (Varshavsky, 1996). The best characterized tag is for one class of substrates: truncated polypeptides from stalled ribosomes. These incompletely translated products are marked at their C-termini with an 11 amino acid sequence called the ssrA tag and targeted for degradation principally by ClpXP (Gottesman et al., 1998). However, some substrates may have more complicated, multicomponent recognition signals. A screen of in vivoClpXP substrates revealed many target proteins carried multiple ClpX-recognition sequences. Furthermore, the screen also indicated that many ClpX substrates ( 60%) are
multimeric or subunits in multiprotein complexes (Flynn et al., 2003). For substrates in multiprotein complexes that are remodeled or disassembled by ClpX, how the enzyme distinguishes between assembled complexes versus subunits is not understood.
To investigate how preference for an assembled complex is achieved over con-stituent subunits, we used the MuA transposase, a natural disassembly substrate of ClpX. Phage Mu duplicates its genome by replicative transposition. During transposi-tion, MuA binds DNA sites located at the ends of the Mu genome, forms a tetramer that brings the two ends of the DNA together, and catalyzes the DNA cleavage and join-ing reactions core to transposition (Craigie et al., 1984; Kuo et al., 1991; Lavoie et al., 1991). This recombination phase of transposition does not require external energy (such as ATP) and is driven forward by proceeding through a series of nucleoprotein complexes (transpososomes) that increase in stability to a final hyperstable DNA product-bound transpososome (Surette et al., 1987). Remodeling converts the hyperstable transposo-some (MuA complex) into a fragile complex, which facilitates both disassembly and the recruitment of DNA-replication machinery (Levchenko et al., 1995; Nakai & Kruklitis, 1995; Jones et al., 1998). Completion of a reaction cycle therefore requires that the stable MuA complex be remodeled by ClpX (Mhammedi-Alaoui et al., 1994).
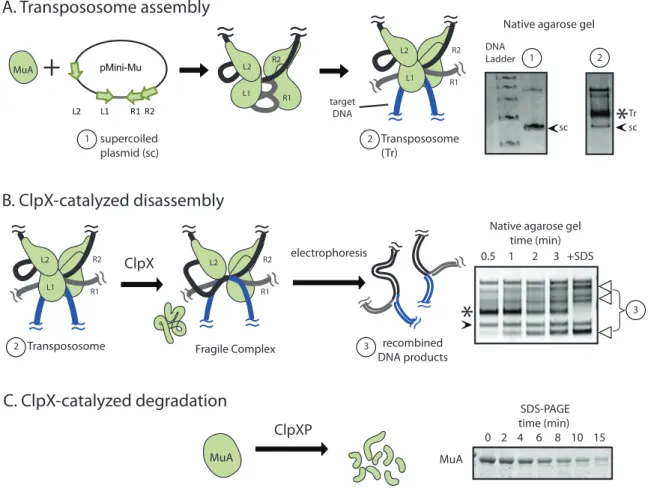
MuA transpososome assembly, recombination, and ClpX remodeling have all been reconstituted in vitro. On a native agarose gel, the stable transpososome is observed as a slower migrating band (asterisk, lane 2) as compared to supercoiled substrate plasmid (arrow, lane 1) (Figure 2-1A). In contrast, the fragile complex is unstable to gel elec-trophoresis and the liberated DNA transposition products are visible as a characteristic series of topoisomerases (Figure 2-1B, bracket). As both monomeric and DNA-bound tetrameric MuA are present in the cytoplasm we sought a molecular understanding to explain how ClpX recognized the transpososome as a high-priority target.
Previous analysis revealed that there was information throughout MuA protein that guided ClpX recognition and that information in domain III is most central. MuA contains a C-terminal sequence (RRKKAI) that is necessary for ClpX recognition of both monomeric MuA and assembled transpososomes (Levchenko et al., 1995, 1997; Abdel-hakim et al., 2008). It is also established that interaction between the transpososome
C. ClpX-catalyzed degradation MuA ClpXP SDS-PAGE time (min) 0 2 4 6 8 10 15 MuA MuA L2 L1 R1 R2 pMini-Mu 1 supercoiled plasmid (sc)
+
Transpososome (Tr) 2 A. Transpososome assembly≈
≈
R1 L1 L2 R2≈
≈
R1 R2≈
≈
≈ ≈
L1 L2 target DNA B. ClpX-catalyzed disassemblyFragile Complex recombined DNA products 3 ClpX electrophoresis Transpososome 2
≈
≈
R1 R2≈
≈
≈ ≈
L2≈
≈
≈
≈
≈
≈
≈
≈
R1 R2≈
≈
≈ ≈
L1 L2 DNA Ladder*
1 2*
3Native agarose gel time (min) 0.5 1 2 3 +SDS
sc sc Tr
Native agarose gel
Figure 2-1: In vitro assays for Mu complex assembly and recognition by ClpX
A. MuA transposase monomers and host protein HU are incubated with a plasmid sub-strate (“pMini-Mu”) containing “left” and “right” phage Mu attachment sites ( L1, L2, R1, R2). Mu catalyzes DNA cleavage and recombination with target DNA to form the trans-pososome, a stable complex. When visualized on a native agarose gel, the transpososome appears as a band (“Tr”, asterisk) that migrates slower than supercoiled plasmid alone (“sc”, black arrow)
B. ClpX remodels the transpososome (MuA complex) by unfolding a subunit bound to L1 or R1 attachment site to produce the fragile complex. The fragile complex falls apart during gel electrophoresis and produces a stereotypical series of recombined topoisomers (white arrows). Addition of SDS disrupts all inter-protein and protein-DNA interactions within the MuA complex and serves as the “100% disassembly” control. Rates of MuA complex disassembly by ClpX were assayed by measuring the rate of appearance of the lowermost disassembly DNA product on a native agarose gel.
C. Schematic of monomeric MuA degradation by ClpXP protease. Rates of protein degradation were assayed by measuring the rate of disappearance of MuA on SDS-PAGE.
and the unfoldase is not simply due to avidity contributed by the four C-terminal tags (Mu pore-binding tag). This conclusion is supported by the findings that additional mu-tations in MuA domain III antagonize only MuA transpososome disassembly by ClpX and not degradation (Abdelhakim et al., 2008). Furthermore, the N-domain of ClpX is exceedingly important for transpososome remodeling but has little role in MuA monomer degradation (Abdelhakim et al., 2008).
As previous studies established that ClpX recognizes short peptide-like signals, we characterized whether the remodeling-specific protein-protein contacts were also peptide-like or not. Second, we sought to understand at the molecular level how MuA assembly into a DNA-bound tetramer modulates recognition by ClpX. Because recognition de-pends on architectural features of the complex, we designed a hybrid Mu protein with novel DNA-binding specificity to assist in specifically placing subunits with altered recog-nition tags within assembled complexes. Lastly, we modulated the binding affinity of the C-terminal tag to understand the extent of cooperation among the different recognition peptides that together comprise the Mu transpososome remodeling signal “recognon”. Thus, here we establish molecular interactions between MuA and ClpX that enable ClpX to preferentially target the assembled Mu tetramer. This work also elucidates principles attractive to the general problem of designing recognition mechanisms that favor assem-bled, multimeric protein complexes.
2.3
Results
2.3.1
Identification of a region critical for enhanced recognition
of transpososomes by ClpX
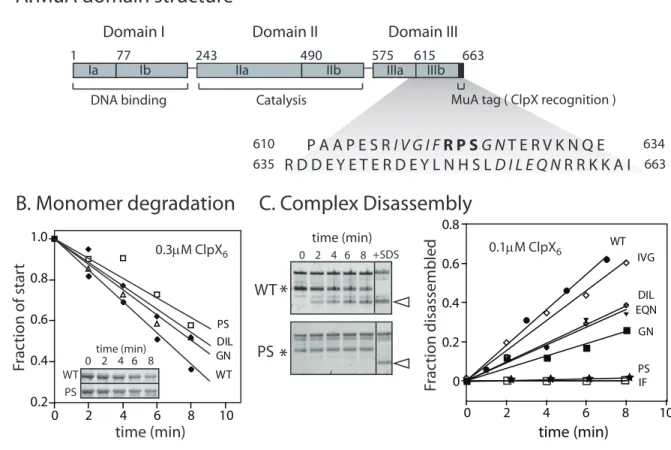
MuA consists of three domains and belongs to the DDE family of recombinases (reviewed in Rice & Baker, 2001). Previous analysis revealed that Domain III contributes most of the information recognized by ClpX (Abdelhakim et al., 2008). Although the majority of the structure of MuA is known, including the architecture of the transpososome (Clubb et al., 1994, 1997; Schumacher et al., 1997; Montaño et al., 2012), there is essentially no structural data of Domain III to guide our analysis. Therefore, we tested the primary sequence surrounding three arginines that had been identified previously as participat-ing in transpososome-specific contacts (Figure 2-2A). Selected residues were mutated to aspartic acid as acidic amino acids disrupt ClpX contacts within other recognition tags (Flynn et al., 2003).
MuA variants were purified, shown to assemble into stable transpososomes, and assayed for monomer degradation by ClpXP (Figure 2-2B) and for complex disassembly by ClpX (Figure 2-2C) at a substrate concentration significantly below the KMfor
trans-pososomes. Most substitution variants had small to modest defects on degradation rates by ClpXP (within 75% of wild-type rate) indicating that the interaction between these altered monomer variants and ClpX was similar to that of wild-type MuA. These same mutations in MuA complexes were also modestly slower (at most 2-fold) in disassembly reactions (Figure 2-3). The exception was that the I620D/F621D variant reduced both degradation of monomers and disassembly of complexes rates to a significant extent. The reason for this dual effect was not analyzed in detail, but it appeared the introduction of negative charges at this position was broadly deleterious. Because the goal of our mutation-based search was to uncover residues specific to the ClpX and transpososome interaction, we did not continue analysis of IF/DD. We categorized these residues as not contributing to ClpX’s distinction between the monomeric and tetrameric states of MuA. However, P623D and S624D displayed similar characteristics as the previously
1 77 243 490 575 615 663 Ia Ib IIa IIb IIIa IIIb
Domain I Domain II Domain III
DNA binding Catalysis MuA tag ( ClpX recognition )
P A A P E S R I V G I F R P S G N T E R V K N Q E R D D E Y E T E R D E Y L N H S L D I L E Q N R R K K A I
610
635 663
634
A. MuA domain structure
C. Complex Disassembly
WT PS*
time (min) 0 2 4 6 8*
+SDS 0 0.2 0.4 0.6 0.8 0 2 4 6 8 10 WT IVG IF PS GN DIL EQN time (min) Fr ac tion disassembled 0.1µM ClpX6 0.2 0.4 0.6 0.8 1.0 0 2 4 6 8 10 WT GNDIL PS time (min) Fr ac tion of star t WT PS 0 2 4 6 8time (min)B. Monomer degradation
0.3µM ClpX6E.
Figure 2-2: Mutation of a sequence region R622-S624 reduces disassembly and degrada-tion rates
A. MuA transposase is a 75kDa protein comprised of three domains. Domain III con-tains the C-terminal Mu pore-binding tag comprised of the last eight residues, which is recognized by ClpX
B. Degradation of wild-type MuA and MuA “aspartate” variants by ClpXP protease at sub-saturating enzyme concentrations. All “aspartate” variants are labeled with the endogenous residues that were targeted for aspartate substitution. Substrate concen-tration was 1uM. Inset shows a representative SDS-PAGE gel of wild-type MuA and MuA(P623D, S624D) monomer.
C. Disassembly of complexes assembled from wild-type MuA and MuA “DD” variants by ClpX unfoldase. Transposososmes are marked by asterisks. Initial transpososome concentration was 100nM. DNA disassembly product used for quantification is marked by white arrow. The “+SDS” lane shows the pattern of topoisomer migration upon complete disassembly. Two representative native agarose gels of wild-type MuA complexes and mutant Mu(P623D, S624D) complexes. Quantification of DNA disassembly product appearance.
0 20 40 60 80 100
WT Δ8 IVG617 620IF 622R 623PS GN625 DIL653 EQN656
DD DD DD DDD A DDD DDD Ra te r ela tiv e t o W T (%) Monomer degradation Complex disassembly
Figure 2-3: Comparison of all reaction rates for Mu aspartate variants
Quantification of differences in degradation and disassembly rates of MuA variants whose indicated sequences were mutated to alanine or aspartic acid relative to wild-type MuA. Reactions with error bars were performed in triplicate. Error bars are the standard error of the mean.
identified transpososome-specific contact residue R622 in which substitution specifically slowed disassembly rates by as much as 10-fold at the sub-saturating protein concentra-tions (Figure 2-3).
We determined the functional interaction of ClpX with double mutant (P623D S624D) variant complexes during disassembly by measuring the concentration of enzyme for half-maximal velocity (KM). Because it is difficult to obtain transpososomes at high
concentration, we started with a fixed substrate concentration, varied the concentration of ClpX, measured the rate of appearance of DNA transposition product released by disassembly, and analyzed these data as previously described to obtain apparent KM
values (Pyle & Green, 1994; Abdelhakim et al., 2008). For many ClpX substrates, the KM is nearly equivalent to the KD because catalysis is relatively slower that the binding
reactions. Therefore the apparent KM is a measure of the functional affinity of ClpX for
transpososomes. The apparent KMfor disassembly of PS→DD double-mutant complexes
0 0.5 1 1.5 2 2.5 0 5 10 15 20 25
WT
PS
ClpX (µΜ)
6Reac
tion R
at
e (min
-1)
K
M appV
app max ( µM ) (min )10.6 2.7
±
1.4 0.2
±
2.7 0.1
±
0.6 0.1
±
WT
PS
−1Figure 2-4: Residues P623 S624 form a critical interaction between MuA complex and ClpX
Half-maximal velocity determination for ClpX-mediated disassembly of wild-type com-plexes and MuA(P623D,S624D) mutant comcom-plexes. Curves were repeated in triplicate. Error bars are the standard deviation of the average.
the Vmax was 5-fold slower compared to that of wild-type MuA complex. These data
indicate that the PS/DD mutations impact both recognition and initial post-recognition steps of disassembly (see discussion). Furthermore, these residues are not major contrib-utors to monomer recognition but specific for transpososome recognition by ClpX.
A boundary defined by the sharp drop in disassembly rates of the mutant vari-ants spans MuA residues 622-624. This critical region for enhanced recognition of Mu transpososomes behaves in contrast the C-terminal Mu pore-binding tag. Truncations of the C-terminus established that MuA monomers and MuA complexes both rely on the Mu pore-binding tag for ClpX specificity (Figure 2-3D and Levchenko et al. 1995). Point mutations of tag residues also led to a significant decrease of both degradation and disassembly rates (Abdelhakim et al., 2008). Because residues 622-624 appeared to be functionally important to ClpX interaction exclusively in the context of the assembled MuA complex, we hypothesized that this critical region may function as a peptide-signal key for transpososome-specific recognition by ClpX.