HAL Id: hal-01898765
https://hal.archives-ouvertes.fr/hal-01898765
Submitted on 18 Oct 2018HAL is a multi-disciplinary open access archive for the deposit and dissemination of sci-entific research documents, whether they are pub-lished or not. The documents may come from teaching and research institutions in France or abroad, or from public or private research centers.
L’archive ouverte pluridisciplinaire HAL, est destinée au dépôt et à la diffusion de documents scientifiques de niveau recherche, publiés ou non, émanant des établissements d’enseignement et de recherche français ou étrangers, des laboratoires publics ou privés.
Protein Lii Methylation (prmA) and Pantothenate
Transport (panF) in Escherichia coli K-12 Downloaded
from
Anne Vanet, Jacqueline Plumbridge, Jean-Herve Alix
To cite this version:
Anne Vanet, Jacqueline Plumbridge, Jean-Herve Alix. Cotranscription of Two Genes Necessary for Ribosomal Protein Lii Methylation (prmA) and Pantothenate Transport (panF) in Escherichia coli K-12 Downloaded from. Journal of Bacteriology, American Society for Microbiology, 1993, 175 (22), pp.7178 - 7188. �10.1128/jb.175.22.7178-7188.1993�. �hal-01898765�
Vol. 175,No.22 JOURNALOFBACTERIOLOGY, Nov.1993, p.7178-7188
0021-9193/93/227178-11$02.00/0
Copyright © 1993, American Society forMicrobiology
Cotranscription
of
Two
Genes
Necessary
for
Ribosomal
Protein
Lii
Methylation
(prmA) and
Pantothenate Transport
(panF)
in
Escherichia coli
K-12
ANNEVANET,* JACQUELINEA. PLUMBRIDGE, ANDJEAN-HERVE ALIX
Institut de BiologiePhysico-Chimique, (URA 1139), Centre National de la RechercheScientifique,
13, ruePierreetMarie Curie, 75005 Paris, France
Received 1 June1993/Accepted 17 September1993
Genetic complementation andenzymeassayshave shown thatthe DNAregion betweenpanF,whichencodes pantothenatepermease,and
orfl,
thefirstgeneofthefisoperon,encodesprmA,thegenetic determinantfortheribosomal protein Lll methyltransferase. Sequencing ofthis region identified one long openreading frame thatencodesaprotein of31,830 Da and correspondstotheprmAgene.Wefound,both in vivo and invitro, that prmAis expressed frompromoterslocatedupstreamofpanF and thus that the panFandprmAgenesconstitute
abifunctionaloperon.Welocated the major 3' end ofprmA transcripts 90 nucleotides downstreamofthestop codon of prmA in the DNA region upstream of thefis operon, a region implicated in the control of the expression ofthefis operon.Although nopromoteractivity wasdetected immediately upstream ofprmA, Si
mapping detected5' ends of mRNA in this region, implying that somemRNAprocessing occurswithin the bicistronicpanF-prmA mRNA.
Posttranslational modification of several ribosomal proteins is a general phenomenon observed in both prokaryotes and eukaryotes (2). Although much information ontheexpression of the genes for bacterial rRNAs and ribosomal proteins is available(29), little attention has been giventotheir
posttrans-lational modifications, which are actually quite numerous. In
Escherichia coli, proteins suchasS5, S18, L12, andEF-Tu are
acetylated at their N-terminal residues (L7 is the acetylated form ofL12). Others modifications include addition ofamino
acids to the polypeptide chain, e.g., addition ofone to four glutamic acid residues to the C terminal ofribosomal protein
S6 (25, 30). However, the most frequent modification is methylation. Several ribosomal proteins, e.g.,
Sli,
L3, L7/L12,Lii, L16, and L33, as well as EF-Tu and IF3, have
N-methylated amino acids at specific positions (reviewed in reference 2). Lii is the most heavily methylated ribosomal protein, with three trimethylated amino acid residues: two
Ns-trimethyllysines at positions 3 and 39 (19) and an amino-terminalNox-trimethylalanine (18, 35). Methylation of
riboso-mal proteinL1I requiresS-adenosyl-L-methionine as amethyl
group donor(4) in a co- or posttranslational event.
Mutations calledprmA, which result in an unmethylated form of
L1i,
have beenisolatedindifferent laboratoriesandbyindependent procedures (15-17, 27). The prmAl locus was
mapped by Colson et al. (14) to 71 min on the E. coli chromosome. This methylation might be expectedtoaffect the function of L1I. However, theprmA mutant strains were all
phenotypically indiscernible from their wild-type parents,and
therefore the function, if any, of this energetically costly
methylation of protein
L1i
is unknown.L1i
has beenimpli-cated in several aspects of ribosome function and assembly, namely, the stringent response in vivo (6, 43-45) and invitro,
ribosomal subunit association (21,31, 40), thebinding domain
of the antibiotic thiostrepton (60), ribosomal protein L16
assembly during 50S subunit reconstitution (9, 23), protein
*Correspondingauthor.
synthesis
termination (7, 58, 59), and the ribosomal GTPasedomain
(20,
48). Despite all of these suggestive results, theprecise role for Lii in the ribosome is obscure.
Moreover,
mutantsapparently lacking Lii areviable(56,
57).
The L1I molecules isolated from prmA strains are severely undermethylated. In vitro methylation experimentswith
ribo-somes from theprmA strains and with extracts fromprmA+
bacteria and radioactively labeled
S-adenosyl-L-methionine
show that theprmA mutations result in loss ofmethyl groups
from both the N-terminal alanine and the internal lysine
residues
(5, 14,
15). However,someresidualLiimethyltrans-feraseactivitycanbedetected inextractsfrom strainscarrying prmAl orprmA3mutations(2,15).This residualactivitycould
accountfor thelack ofaphenotype associatedwith theprmA
mutations. Since single mutations apparently result in the
absence of both internal and N-terminal methylgroups from
Li1,itseemspossible that thesame enzymeisresponsiblefor
themethylationof allthreeamino acids. If this is the case, then the distinctive trimethylation of two different amino acids at threedifferentloci when all of the otherlysine residuesof the
protein are unmodified poses an interesting problem of en-zymespecificity. We cannot, however,rule out thepossibility
thattwoormoreenzymes are necessaryandthatprmA isthe
genetic determinant of thelimitingstepwhich prevents subse-quentmethylations.
Tostudythe role ofmethylation onLii function,we have
undertaken a systematic study of the prmA gene.
Chromo-somalDNAfragmentscarrying theprmAgene wereisolatedin a series of X transducing phages by complementation of a
thermosensitive mutation in the nearby genefabE (now re-named accB; encodes the biotin carboxyl carrier protein, a subunit ofacetylcoenzymeAcarboxylase)(3).ThepanFgene
(encodes pantothenate permease) has been mapped to the
samechromosome region (62).All of the accB+ Xtransducing
phages which carried panF were also found to be prmiA+,
showingthat genes prmnA andpanF arecloselylinked(3).The nucleotide sequence of panF has been determined (28). We presentheretheDNAsequenceof prmA and ananalysisofits
transcription. 7178
on December 7, 2016 by INIST-CNRS
http://jb.asm.org/
TABLE 1. Bacterialstrains, plasmids, and phages used in this study
Strain,plasmid, Genotype Referenceororigin
orphage
E.coli strains
IBPC5321 thi-1argG6 argE3 his-4xyl-5 mtl-l tsx-29 rpsL AlacX74 46
MB984 srl::TnJO recA prmA3 IBPC101(47) x MB1984(3)
byP1 transduction
JM109 recAI endAIgyrA96 thihsdRI7supE44relAI A-A(lacproAB)F' (traD36proABlacIq 65
lacZAM15)
MB1541 thr-I leu-6 trp his-4 argHlthyAl thi-1xyl-7 tonA2 supE44rpsL9ficTs' prmA3 3
Plasmids
pBR322 amp tet 10
pFA accBC panF amp 3
pAF2 panFprmAorfl amp Thiswork
pAF3 panFprmA amp This work
pRS415 amp 'lacZ 54
pAO1 panFprmA-lacZamp This work
pAO2 panF-lacZamp This work
pROl 'panFprmA-lacZamp This work
pRO2 'panFprmA-lacZamp This work
pUC(PrmA) placprmA This work
Phages
X6G3 accBC panFprmA orflfis 33
XFA6 accBCpanFprmA 3
XRS45 'lacZYA 54
XRS/A01 panFprmA-lacZ This work
XRS/A02 panF-lacZ This work
XRS/RO1 'panFprmA-lacZ This work
XRS/RO2 'panFprmA-lacZ Thiswork
a Aprime denotes thatthegeneistruncatedonthat sidesothat thepromoter regionisabsent.
MATERIALS AND METHODS
Strains and media. Thebacterial strains andplasmids used
in this study are listed in Table 1. All strainswere routinely
grown in Luria-Bertani (LB) medium. Synthetic MOPS
me-dium(41) containing0.2%glucose and 50jigeach of arginine
andhistidinepermlat30°Cwasusedinbacterial cultures for
measurements of P-galactosidase activities as previously
de-scribed (39). Plasmidswere propagated inE. coli JM109 or
IBPC5321. X6G3 is from the E. coli library of Kohara etal. (33). Plasmid pUC(PrmA) carries the prnzA structural gene,
between oligonucleotides CHIEN and IFRO (see Fig. 2),
cloned downstream ofthe lac promoter. Itsconstruction will
bedetailed elsewhere (64).
DNAmanipulationsandsequencing. Restriction
endonucle-asesand others enzymeswereobtained commercially. Radio-chemicalswerepurchasedfrom Amersham. Large-scale
prep-arations of plasmid DNA, digestions, and cloning were
performed by standard procedures (50). DNA restriction fragments and polymerase chain reaction (PCR)-generated
fragmentswerepurifiedfromagarosegels bythe freezephenol method(53)orwithGene Clean(Bio 101). ThePCR(50 ,ul)
contained 20pmolofeacholigonucleotide, 10ngofaplasmid
[pAO2, pUC(PrmA), or pAF3] template, 2 U of Taq
poly-merase, 0.2 mM each deoxynucleoside triphosphate, 10 mM
Tris-HCl (pH 8.3), 50 mM KCl, 2.5 mM MgCl2, and 0.01%
gelatin.PCRsweregenerallycarriedoutbythefollowingthree
steps repeated 20 times: denaturation at 92°C for 1.5 min, annealingat55°Cfor 1.5min,andextensionat72°Cfor2min. Double-stranded DNA of recombinant plasmid pAF3 was
directly sequenced on both strands as previously described
(51). For the oligonucleotides used as sequencing primers
(three foreachstrand), see Fig. 2. Dideoxy-sequencing
reac-tions using T7 DNApolymerase and a-35S-dATP were
ana-lyzed
on 0.4-mm-thick 8% acrylamide gels containing 7 Murea.
mRNA preparation. Total RNAwas prepared from strain
JM109 or IBPC5321 grown in LB medium. Strains carrying plasmidpAF3 or pFA were grown in LB containing500 ,ug of
ampicillin per ml. Cells were harvested at an A650 of 0.7, washed with 10
mM
MgCl2, andresuspendedin 20 mMsodium acetate (pH4.7)-1%
sodium dodecyl sulfate, and RNA wasextracted bythe hotphenolmethod (49).
S1 nucleasemapping. Tomapthe 5' ends ofpanF
andprmA
transcripts,probes uniquelylabeledatone5'endwere synthe-sized by PCR. A 20-pmol sample of one
oligonucleotide
(NAPCO
for panF and ARTforprmA)was5'end labeled with[y-32P]ATP
(specific activity, 3,000Ci/mmol)
andpolynucle-otidekinase. The second
oligonucleotide
was complementarytoplasmidvectorsequences upstreamof the
panFprmA
regioninserted in theplasmidusedasthetemplatefor the PCR.The reverse sequencing primer (REV17) was used with
plasmid
pUC(PrmA)
for theprnmA
probe (see Fig.6)
andanoligonu-cleotidehomologoustothe amp gene ofpRS415
(RBP22)
withpAO2for panF (see Fig. 4). The
SI
probes thussynthesized
carried a nonhybridizing DNA extension at the unlabeled
oligonucleotide
end whichpermits
differentiation betweenreadthrough transcription andreannealing of the probe. The labeledfragmentswereseparated from
unpolymerized
oligo-nucleotides on a 1% agarose gel andpurified by
the freezephenolmethod (53).
To mapthe 3' ends of
prmA
transcripts, aDNAfragment,
RUE-IFRO (see Fig. 2), was
synthesized
by PCR. Thisfrag-ment was digested by
HindIll.
TheHindlIl-IFROfragment
waspurifiedfroma1% agarose
gel
with GeneClean(Bio
101),
and the
Hindlll
endwas labeled with[ot-32P]dATP
with the Klenow fragment of DNApolymerasein the presence of 0.2on December 7, 2016 by INIST-CNRS
http://jb.asm.org/
7180 VANET ET AL.
mMeach
dCTP,
dGTP, and dTTP. This 3' end-labeledDNAfragment
was purified by elution from a 5% polyacrylamidegel.
The end-labeled DNA probes, approximately 50,000 cpm
per
hybridization experiment,
wereincubated with 10to30p.gofE. coli total RNA in 80% formamide-40 mM
piperazine-N,N'-bis(2-ethanesulfonic acid) (PIPES;
pH6.4)-I
mMEDTA-400 mM NaCl
(final
volume,
50Rl).
After denatur-ation at85°C
for 10 min, hybridization was carried outovernight
at56°C
(5'-end
mRNAmapping)
or 52°C (3'-endmRNA
mapping)
and stopped bytransferring
the hybridizedmixtureto 0.4 mlof coldSI nuclease
digestion
buffer(30
mMsodium acetate
[pH 4.6],
250 mM NaCl, 1 mM ZnSO4, 5%glycerol) containing
100 UofSI nuclease(Boehringer GmbH,
Mannheim, Germany).
Thesamples
were incubated at 37°Cfor 45 min
(5'-end mapping)
or at25°C
for 1.5 h (3'-endmapping).
SI nuclease-resistant DNAfragments
wereana-lyzed
onpolyacrylamide
gels 1mm thickcontaining7 Murea.The
gels
werefixed, dried,
andanalyzedwithaPhospholmager
(Molecular Dynamics),
which permitted quantitation ofS1-protected
bands,
orautoradiographed
with Cronex Hi-Plusamplifying
screens.Reversetranscription.
Synthetic
oligonucleotides
wereusedfor
primer
extensionexperiments
with avian myeloblastosisvirusreverse
transcriptase
(Boehringer).
The oligonucleotideswere 5' end labeled with
[_y-32P]ATP
andpolynucleotide
kinase and separated from the labeled nucleotide
by
electro-phoresis
ondenaturing
20%polyacrylamide gels. Typically,
15or 30 jig ofE. coli total RNA and 0.5 pmol of the labeled
oligonucleotide
wereannealed inavolume of5Rl.
Thereversetranscription
reaction wasperformed
aspreviously
described(61).
Theprimer
extensionproducts
wereanalyzed
ondena-turing polyacrylamide gels
as described above.In vitro transcription.
Templates
were madeby
PCRwith suitableoligonucleotides.
The reaction mixture(20
il)
con-tained40 mMTris-HCl
(pH
8.0),
10 mMMgCl2,
100 mMKCl,
1 mM
EDTA,
100pLg
ofbovine serumalbuminperml,
50ngof
template
DNA(e.g.,
theAGFIN-NAPCOPCR-generated
fragment
formapping
of thepanFpromoter),
and0.3to 1 UoftheE. coli RNA
polymerase
holoenzyme
(Boehringer).
After15min of
preincubation
at37°C,
atranscription
wasinitiatedby
additionof 400 FMeachATP, CTP,
and GTP and 100 FM[0-32P]UTP.
After 15 min, thesamples
wereprecipitated
with ethanol andanalyzed
ondenaturing polyacrylamide
gels asdescribed above.
Construction of
panF-lacZ, prnzA-lacZ,
andpanFprmA-lacZtranscriptional
fusions. All fusionswere constructedonplas-mids in vitro and transferred to X
by
in vivo recombination. PlasmidpAOI(panF
prmA-lacZ)
wasconstructedbyinserting
the 2.4-kb BamHI
fragment
frompAF3, carrying
a smallfragment
ofpBR322,
the end of accC(fabG),
theentirepanF
gene,and thefirst 10nucleotides of
prmA,
into theBamHI siteof
pRS415
(54).
Plasmid pAO2(panF-lacZ)
was constructedsimilarly,
by
inserting
the 1.95-kbBamHI-BglII
fragmentfrompAF3
(the BglII
site is locatedatthe end ofpanF[see Fig. 1])into the BamHI site ofpRS415. Plasmid
pROl
was derived frompAOI
by
deletion ofthe DNAbetween the EcoRIandBglII (at
theend ofpanF)sites(see Fig. 1).Plasmid pRO2wasmade
by
inserting
the 1,530-bpEcoRV-to-BamHI
fragmentfrom pAF3 into pRS415 digested by
SmaI
andBamHI.
Thefusions carried
by
pAOI,
pAO2, pROI,
and pRO2 weretransferredtoXinvivo
by using XRS45
aspreviously
described(54)
togive
XRS/AO1, XRS/AO2, XRS/ROI,
andXRS/RO2.
Two
independent
plasmid
constructs for each fusion were transferredtoXRS45,
and two recombinants from eachplas-mid were
picked, purified,
and used to make lysates tolysogenize
strain IBPC5321.Monolysogens
were determinedand
,3-galactosidase
activities were measured aspreviously
described
(39).
Lll
methyltransferase
activity
measurements. Ribosomalprotein
L1I methyltransferase
activity
wasmeasuredbasically
as
previously
described(15).
To measure thespecific
Lii
methyltransferase
activity
ofaparticular
crudeextract,bacte-riafroman
overnight
culture ofE.colicells in LBmedium with500 jig of
ampicillin
per ml(for
plasmid-carrying strains)werepelleted, resuspended
in anappropriate
volume of a buffercontaining
10mMTris-HCl(pH 7.6),
10 mMMgCL2,
60 mMNH4Cl,
and 6 mMf-mercaptoethanol,
andbrokenby
sonica-tion.The25-jl
methylation
assaycontained 1A260
unit of50S ribosomalsubunits ofE.coli MB1541(prmA3)
asthesubstrate(methyl
group acceptor), 25 jiMS-adenosyl-L-[methyl-3H]-methionine
(3 Ci/mmol; Amersham)
as the methyl groupdonor
(a
concentrationeightfold
over the reported Km[3.2
,uM]
of the enzyme for S-adenosyl-L-methionine)(13),
and various amounts of the bacterial extracts as a source of Li1 methyltransferase. Reaction mixtures were usually incubated at 30°C for 1 to 30min,
and the hot 5% trichloroacetic acid-insolubleradioactivity
was collected on Millipore filters and counted. Proteinconcentrations in the crudeextractwereestimated
(ii),
andthefinal resultwasexpressedinpicomoles
of
methyl
groupsincorporated
per minute of incubation at30°C per
milligram
ofprotein
in the crude extract. RESULTSSubcloning
and identificationofprmA
asthegeneticdeter-minant for L 11methyltransferase.TheprinAgenewasfound
to be
closely
linked tothe accB (formerly fabE; encodes thebiotin
carboxyl
carrier protein subunit ofacetyl-coenzyme Acarboxylase) and
panF
(encodes pantothenatepermease)
genes at 71 min onthe E. coli chromosome (3). Of the 75 X
bacteriophages
isolated as complementing a thermosensitivemutation inaccB, the 58 thatwerepanF were alsoprmiA
+,
showing thatpanF andprmiA are likely to be adjacent. The gene
fis,
which encodes the factor for inversion stimulation(Fis),has beenmappedto the sameregion (32).
Comparison
oftherestriction maps of the
accB,panF,prmA,
andfisregions
showed that these fourgenesarecarriedon Kohara
bacterio-phage
X6G3(33) (Fig. 1).
The accB, accC (formerlyfabG;
encodes thebiotin
carboxylase
subunit ofacetyl coenzymeAcarboxylase),andpanFgeneshave beensequenced(3, 28,
34,
38),
as have fis (32) and a gene (orfl) with an unknown function positioned immediately upstream offis andtran-scribed from the same promoter (8, 42). Located between
panF and orfi is a region of about 1.0 kb which was not
sequenced
(Fig. 1).
This region is alsopresent on the accB+panF+ prmiA+transducingphagesof Alix(3),while it isabsent
from plasmid pFA, which complements the accB and
panF
mutationsbutnot aprmnA mutation (3). Thus, theprmnA gene should lie within theunsequenced DNA
region.
Wetherefore cloneda4.2-kbPvuIIfragment,
encompassing
thisregionfrom X6G3into theEcoRVsite ofpBR322to
give
plasmidpAF2.This fragmentcarries the end(last 136
codons)
of the accC gene, panF, and the orfi gene, as well as the
unsequenced DNA. A slightly smaller plasmid, pAF3, was
madeby deleting 1,060 bp which includedorfi (Fig. 1). This
plasmid was introduced into the prmA3 mutant strain (MB984),andLII methyltransferase activitywasmeasuredon crude extracts. MB984 (prmA3) showed about 2.5% of the
activity ofwild-type E. coli IBPC5321 (Table 2). Introduction
ofplasmid pAF3 ledto aconsiderable increase ofLii
meth-yltransferase activity, showing that plasmid pAF3
comple-J. BACTERIOL.
on December 7, 2016 by INIST-CNRS
http://jb.asm.org/
B
B
XFA6 -\. v//
B B
accBC p&nF pnnA r
X6G3 xY
Z//
// ,xx x xy/ccBC ~~~~p&nFprmAorf t
BEV/P EV B
tet' 'accC panF prmA or11 amp
B EV/P EV Bg B H E
1-k,V
:tet' 'accC panF prmA amp
ESB EV/P EV Bg B
amp accC panF,prmA-lacZYA tet
ESB EV/P amp 'accC amp EV Bg panF-lacZYA tet E/Bg B
amp 'panF, prmA-lacZYA tet
E Bg B
li
X>tIj
'panF, prmA-lacZYA tet
1kb
oligonucleotides
'accC panF prmA orfl'
_GF
__I
- -_-_F
AGFIN BGFIN FNAP NAPCO CHIEN AMFIP VERSE ART RUE AL IFRO
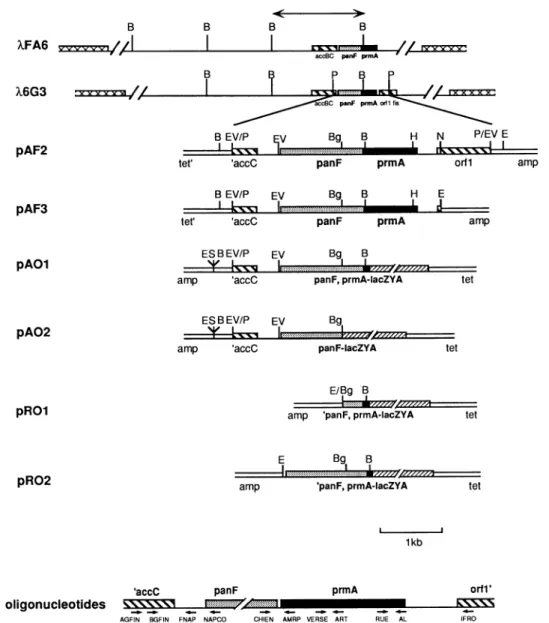
FIG. 1. Subcloning of the panF prmA region of theE.coli chromosome.RestrictionmapsofXFA6andX6G3are shownwith thelocations of
theaccBC,panF,andprmA genes.The4.2-kbPvuIIfragment of X6G3wascloned intotheEcoRV siteof pBR322togive pAF2.Deletion of the
NcoI-EcoRI fragment(by fillinginof thesiteswith the Klenowenzyme and ligation,which recreatesthe EcoRIsite)gavepAF3.The different
DNAfragments insertedinto pRS415toproduceoperonfusions with lacZ, pAO1,pAO2, pROl, and pRO2areshown. _,prmA; Eli,panF;
=,othergenes; ,lacZ; VA, phageX; II1,plasmidvector.Abbreviations:B,BamHI;P,PvuIl; EV, EcoRV; Bg,BglII;E,EcoRI;H,HindIII;
N, NcoI; S, SmaI; E/Bg, hybridsite fromEcoRI and BglII;amp,ampicillin resistance gene; tet, tetracycline resistance gene. <-*, 6-kb BamHI
fragment of XFA6 inserted into the BamHIsiteofpBR322togiveplasmid pFA (3).
TABLE 2. Specificactivities ofL1i methyltransferasein
plasmid-carryingstrains
prmAalleleon Enzymeactivity"
chromosome Specific Relative
MB984(pBR322) prmA3 0.01 0.02
MB984(pAF3) prmA3 6.04 13.72
IBPC5321(pBR322) prmA+ 0.44 1.00
IBPC5321(pAF3) prmA+ 7.34 16.68
'Specificactivitiesareexpressedinpicomolesofmethylgroupsincorporated
permilligramofproteinin thebacterialextractperminuteat30°C.
mented theprmA3 mutation (Table 2, line 2). PlasmidpAF3
gave a similar level ofLII methyltransferase activity in
wild-type strain IBPC5321, resulting in about 15-fold overproduc-tioncomparedwith thesamestraincarryingpBR322 (Table 2,
lines3 and4). This level ofoverproduction, comparabletothe
pBR322copynumber, is consistentwith the idea that theprmA locus encodes the structural genefor LII methyltransferase.
Sequenceof the prmA gene. The DNA sequence of pAF3
betweenpanF and orfi was determined (Fig. 2). One long
open reading frame (ORF) was found in the region
corre-spondingtotheprmA gene.The first ATG codonof this ORF islocated 11 nucleotidesdownstream of the terminationcodon
ofpanF.TheORFterminatesataTAA codon328 nucleotides
upstream of the orfi gene. The prmA gene codes for a
polypeptide of 293 amino acids, giving a protein with a
pAF3
pAO1
pAO2
pROl
pRO2 on December 7, 2016 by INIST-CNRS
http://jb.asm.org/
7182 VANET ET AL.
1
'panF CHIEN
GAATATTCAGTACCTGGGC ITCCACCCTATCGTGCCTCGTI. ATACTA
F3 N I Q Y L G F H P I V P S L L L S L L A F2 0 0 61 TTTCCTGGTCGGAAACCGTTTCGGTACATCCGTCCCGCAAGCTACCGTTTTGACTACTGA F L V G N R F G T S V P Q A T V L T T D 0 *F1 prmA BamHI 121 TAAATAAAGAGTTTTGCCATGCCTTGGATCCAACTGAAACTGAACACCACCGGCGCGAAC 1 K * M P W I Q L K L N T T G A N BglII AAGTCCTA 181 GCGGAAGATCTTAGCGATGCGCTGATGGAAGCGGGTGCCGTTTCTATCACTTTTCAGGAT 15 A E D L S D A L M E A G A V S I T F Q D AMRP TGGGTGCTATGCGGT 241 ACCCACGATACGCCAGTATTTGAACCGCTGCCGGGCGAAACGCGCCTGTGGGGCGACACC 35 T H D T P V F E P L P G E T R L W G D T 301 GATGTGATTGGTCTGTTCGACGCTGAAACCGATATGAACGACGTGGTGGCGATTCTGGAA 55 D V I G L F D A E T D M N D V V A I L E VERSE 361 AACCATCCGTCGCTCGGCGCAGGCTTCGCGCATAAA CGAACAACTAGAAGATAAAGAC 75 N H P S L G A G F A H K I E Q L E D K D 421 ;AGCGCGAATGGATGGATAATTTCCACCCGATGCGCTTTGGTGAACGACTGTGGATC 95 W E R E W M D N F H P M R F G E R L W I ART ACCGCACTACACGGCCTGCTTTTGl 481 TGCCCTAGCTGGCGTGATGT[CCGGACGAAACGCCGTCAACGTGATGTTAGATCCAGGG 115 C P S W R D V P D E N A V N V M L D P G KpnI 541 CTGGCGTTTGGTACGGGTACCCATCCAACCACCTCTCTGTGCCTGCAATGGCTCGACAGC 135 L A F G T G T H P T T S L C L Q W L D S 601 CTCGATTTAACCCGGTAAAACAGTCATCGACTTTGGCTGTGGTTCCGGCATTCTGGCGATC 155 L D L T G K T V I D F G C G S G I L A I EcoRV 661 GCGGCGCTGAAACTGGGTGCAGCAAAAGCCATTGGTATTGATATCGATCCGCAGGCGATT 175 A A L K L G A A K A I G I D I D P Q A I 721 CAGGCCCAGCCGCGATAACGCCGAACGTAATGGCGTTTCTGACCGTCTGGAACTCTACTTA 195 Q A S R D N A E R N G V S D R L E L Y L 781 CCGAAAGATCAGCCAGAAGAAATGAAAGCCGACGTGGTGGTCGCTAACATCCTTGCAGGC 215 P K D Q P E E M K A D V V V A N I L A G RUE 841 CCATTACGTGAACTGGCACCGT ATCAGCGTCCTGCCGGTTTCAGGCGGTTTGCTGGGC 235 P L R E L A P L I S V L P V S G G L L G HindIII 901 CTTTCCGGTATTCTGGCAAGCCAGGCAGAGAGCGTTTGTGAAGCTTATGCCGATAGCTTC 255 L S G I L A S Q A E S V C E A Y A D S F AL LACCTTTTTCTTCTCACCACGGCAT 961 GCACTGGACCCGGTCGTGGAAAAAGAAGAGTGGTGCCGTATTACCGGTCGTAAGAATTAA 275 A L D P V V E K E E W C R I T G R K N * 1021 CCTTCGCATCGCCGTAGGGTGACGCGGGGGCAAGTGCGAGCAAGCTCACAAAAGGCACGT
1081 AAATTTGCCGATTATTTACG;AAATTz5CGTGCCAAAATTTTCATTCATAAAGAAAAATT 1141 GAGAACTTACTCAAATTTCTTTGAGTGTAAATTTTAGTCACTATTTTCTAATATGATGAT 1201 TTTTATGAGTAATTATCGCACCACGCTCAATTTAAATGCAATTCTTTGATCCATCTCAGA 1261 GGATTGGTCAAAGTTTGGCCTTTCATCTCGTGCAAAAAATGCGTAATATACGCCGCCTTG orfI' 1321 CAGTCACAGTATGGTCATTTCTTAACTCATGCGCATCGGACAATATCAGCTCAGAAATCG 1 M R I G Q Y Q L R L R NcoI IFRO CGGTACCGACCGTAATGTCTGTCTj 1381 CCTGATCGCAGCGCCCATGGCTGGCATTACAGACAGACCTTTT 12 L I A A P M A G I T D R P F
FIG. 2. Nucleotide and deduced amino acid sequences of the prmA gene. Theboxed sequences show theoligonucleotidesused asprimers fordeterminationof the nucleotide sequence: CHIEN, VERSE, and RUEon one strand and AMRP,ART, and AL on the other. The sequence presented includes the3' end of panF(28),prmA, andthe
intergenicprmA-orflregion(8,42). Relevantrestriction endonuclease
sites are indicated. The putative ribosome-binding site ofprmA is
shaded. StarsindicatetheterminatiQncodons of panF andprmA. The black circles indicate the 5' mRNA ends mapped immediately
up-streamofprmA byS1nuclease and reversetranscriptasemapping,and
theopencircles indicate the5' ends detectedbyreversetrancriptase.
molecular mass of
31,830
Da. This is in agreementwith theprevious
biochemical data for the molecular mass of themethyltransferase
of30,000
to40,000 Da,
obtainedby
sucrosegradient centrifugation (4)
andSephadex gel
chromatography
(13). Analysis
oftheprmA coding
sequence shows no signifi-canthomology
with otherproteins
in theDNA-protein
data bases.However,
anonapeptide
motifcharacteristic ofmethyl-transferases,
DXGXGXGXL(where
X is any aminoacid),
asdescribed
by
Ingrosso
et al.(26),
is found inprmnA
between amino acids 164 and 172. Thispatternof amino acids has been observed inalarge
number ofmethyltransferases specific
forawide range of
substrates-proteins,
nucleicacids,
and small molecules. Thenonapeptide
motifwas found associated with two otherweakly
conservedmotifs, primarily
in themethyl-transferases
specific
forsimple
biochemicalsubstrates,
e.g., amino acids(26).
The othertwomotifsarenotpresentin the PrmAprotein
sequence. The presence of thenonapeptide
motif is consistent with the idea thatprmA is the structural gene for the
methyltransferase
rather thanaregulatory
protein
necessary for its
activity.
The
prmA
genedoes not show the codon usage character-istic of astrongly
expressed protein (22). Upstream
of theATGinitiation
codon,
there isnocomplementarity,except forGAG,
with the 3' end of 16S RNA(52).
However, somehomology
with the "downstream box"(55)
associated withhigh-level
expression
of certain genes is apparent, 9 of 14 nucleotidesatpositions
143to159within theprmA
gene(Fig.
2).
The
panF-prmA intergenic distance,
11 nucleotides, is notsufficient foraconventionalRNA
polymerase binding
site,andinspection
ofthisregion
showed no obviousfactor-indepen-dent
transcriptional
terminatororhomology
with theconsen-sus -35 and -10 promotersequences,
suggesting
thatthese twogenesarecotranscribed(see below).
The shortintergenic
distance between
panF
andprmAmight
indicate that the twogenes are
translationally
coupled.
Expression
of theprmA andpanFgenesin vivo. The lack ofobvious
transcriptional
signals
ontheDNA sequence betweenpanF
andprmA ledustotrytolocalize the prmA promoterby
using
gene fusions: fourfragments
were cloned into operonfusionvector
pRS415
togive plasmids
pAOI,
pAO2,pRO1,
and
pRO2 (Fig.
1). pAO2
carriesDNA from the end of accC tothe middle ofpanF
(BglII site; Fig. 1)
togive
apanF-lacZ
fusion; pAO1
starts at the same point but extends into thebeginning
ofprmA (BamHI site; Fig. 1)
toproduce
aprmrA-lacZ
fusion,
whilepROl
andpRO2
carry thesameprmA-lacZ
fusion and all(pRO2)
or part(pRO1)
ofpanF
but lack thesequence upstream ofthe
panF
structuralgene. The operon fusions carriedby
theseplasmids
weretransferred toXRS45,
giving
risetoXRS/AO1, XRS/AO2, XRS/RO1, andXRS/RO2,
which were used to
lysogenize
IBPC5321. Lysogens ofXRS/ AO1 andXRS/AO2give
moderate 3-galactosidaseactivities,
while those of XRS/RO1 and XRS/RO2 are near the
back-ground
value(Table 3).
Thisshows thatmostprmAexpression
is
dependent
uponapromoterupstream ofpanF,which couldThesquaresshow the 3'mRNA endsmapped downstreamofprmAby
S1 nuclease, and the major3' end is indicated bythe larger, open square.Thenonapeptidemotifcharacteristic ofmethyltransferasesis inboldface. Thesequence determinedinthis work wasfrom nucleoti-des 60to1080. Theremaining sequencewastaken fromreferences8, 28, and 42 and is included forclarity. The 3' end of the sequence determinedin this workoverlapsthatpreviously published (8, 42).Our
sequenceis inagreementwith that of Ball et al.(8), except foraG-C-G changetoG-G-C at nucleotidepositions1049 to 1051.
J.BAcrERIOL.
on December 7, 2016 by INIST-CNRS
http://jb.asm.org/
TABLE 3.
,3-Galactosidase
activities instrainIBPC5321 carryingdifferentpanF-lacZorprmA-lacZfusions on A lysogens
Lysogen Insert' Activity" ± SD
XRS/AO1 panFprmzA-lacZ 40 ± 2
XRS/AO2 panF-lacZ 156 ± 10
XRS/R01 'panFprmA-lacZ 1.2 ± 0.2
XRS/RO2 'panFprmA-lacZ 3.5 ± 0.3
XRS45 'lacZ 1.0 ±0.2
aAprimeindicates that the gene is truncated so that the promoter is absent.
"Activities are expressed in the units defined by Miller (39). Means of four
independentculturesareshown.
be thepanF promoter itself. The low level ofXRS/RO2 activity couldindicate that there is a weak promoter activity in the 5' region of the panF structural gene. However, it can account for only a minor amount (less than 10%) of prmA expression compared with that from the panF promoter region.
Localization of the panF promoter. The intergenic region between accC and panF is shown in Fig. 3. We used a combination of techniques, Si nuclease protection, primer extension, and in vitro transcription, to localize the panF promoter within this sequence.
The probe for SI nuclease protection experiments as the probe, PCR-generated fragment RBP22-NAPCO, which was synthesized with plasmid pAO2 DNA as the template (Fig. 4C). The NAPCO oligonucleotide was 5' end labeled with
[y-32P]ATP
andpolynucleotide kinase before polymerization.With this probe and total E. coli RNA extracted from a
wild-type strain carrying or not carrying plasmid pFA, we
detected amajorSi-resistantDNAfragment about 340 nucle-otides long (Fig. 4A). This located the 5' end of the panF
mRNAto theseries ofTresiduesaround nucleotide 263 (Bi
of the sequence in Fig. 3). A weaker band of about 395 nucleotides which corresponds to a5' endaround nucleotide 213 (B2 in Fig. 3) was also detected. Primer extension using
B2 B1 pB32 F i'~ panF S1 mlappirgprobe RBP22 NAPCo' B2 S1resistart D tragmenis E
FIG. 4. Mapping of the 5' RNA ends upstream of panF. (A) S1 analysis.Theprobewasthe1,463-bp RBP22-NAPCOPCR-generated
fragment 32p labeledatNAPCO. Itwashybridized with RNA made
from JM109 grown in LB (lanes 2, 15 jLg; lane 3, 30 jig) or
JM109(pFA)grownin LBwith 500jigofampicillinperml(lane 4,15
jig; lane5, 30 jig)orwith30 ,ug of tRNA (lane 1).TheSI-resistant productswereelectrophoresedon a5%acrylamide-7 Mureagel.The
positions oflabeled DNA molecular size markers (inbasepairs) are
shownontheleft, and the locations of the majorprotectedbandsBI, B2, D, and E are noted. (B) Primer extension. Total RNA isolated
from either IBPC5321 (lane 1, 30 jigofRNA) orIBPC5321(pAF3)
(lane 2, 30 jigof RNA; lane 3, 15 jigofRNA) wasused for primer
extension with the 5'-end 32P-labeled NAPCO oligonucleotide. The
twomajor extension products, 340and 395nucleotides, B1andB2,are
indicated. Lanes T, G, C, and Aaresequencing reactions with pAF3
DNAand theNAPCOoligonucleotideasthe primer andrepresentthe
sequenceofthe noncoding strand. Thesequencearound themajor5' mRNAend, shown onthe right, corresponds tothesequence of the codingstrand.The5' endsofthe transcripts within thissequenceare
shownby dots. The numbers refertothenucleotidepositionsofFig. 3. (C) Diagramatic interpretationofthecombinedmapping results.
AGFIN BGFIN TGAAAACCGTGACGTGGCGATTGCCCATGTGCGCTGCAGGAGCTGCATATCGACGGTATCAAAACCAACGTTGATCTGCAGCAGATCCGCATCATGAATGACGAGAACTTCCA 10 20 30 40 50 60 70 80 90 100 110 120 ACTTTTGGCACTGCACCGCTAACGGGCGTACTTCTTACGCGACGTCCTCGACTAGTAGCTGCCATAGTTTTGGTTGCAACTAGACGTCGTCTAGGCGTAGTACTTACTGCTCTTGAAGGT E N R D V A I A R M K N A L Q E L I I D G I K T N V D L Q Q I R I M N D E N F Q accc B2 GcATGGTGGCACTAACATccAcTATCTGGAGAAAAAAcTcGGTCTTCAGGAAAAATAAGAcTGCTAAAGcGTcAAAAGGCCGGAT=ccGGCCTTTTTTATTAcTGGGGATCGACAACC 130 140 150 160 170 180 190 200 210 220 230 240 CGTACCACCGTGATTGTAGGTGATAGACCTCTT=TGAGCCAGAAGTCCTTTTTATTCTGACGATTTCGCAGT=CCGGCCTAAAAGGCCGGAAAAAATAATGACCCCTAGCTGTTGG H G G T N I H Y L E K K L G L Q E K CCCATAAGGTACAATCCCCGCTTTCTTCACCCATCAGGGACAAAAAATGGACACTCG'ITTGTTCAGGCCCATAAAGAGGCGCGCTGGGCGCTGGGGCTGACCC=TGTATCTGGCAGT 250 260 270 280 290 300 310 320 330 340 350 360 GGGTATTCCATGTTAGGGGCGAAAGAAGTGGGTAGTCCCTG=TTTACCTGTGAGCA,A*CA-AGTC-CGG-GTATTTCTCCGCGCtACCCGCGACCCCGACTGGGAAAACATAGACCGTCA FNAP TTGGTTAGTAGCCGCTTACTTATCTGGCGTTGCCCCGGTTTTACCGGCTTTCCGCGCTGGTTTGAGATGGCCTGCATCCTGACGCCGCTGCTGTTrTATTGGACTGTGCTGGGCGATGGTG 370 380 390 400 410 420 430 440 450 460 470 480 AACCAATCATCGGCGAATGAATAGACCGCAACGGGGCCAAAATGGCCGAAAGGCGCGACCAAACTCTACCGGACGTAGGACTGCGGCGACGACAAATAACCTGACACGACCCGCTACCAC AAATTTATCTATCGCGATATCCCACTGGAGGATGACGATGCAGCTTGAAGTAATTCTACOGCGGTCGCCTATCTGGTGGTGGTGTTCGGTATCTCGGTTTATGCGATGCGTAAACGGAG 490 500 T0 520 530 540 550 560 570 580 590 600 TTTAAATAGATAGCGCTATAGGGTGACCTCCTACTGCTACGTCGAACTTCATTAAGATGTrGACCAGCGGATAGACCACCACCACAACCCATAGAGCCAP TACGCTACGCACTTTGCcrc M Q L E V I L P L V A Y L V V V F G I S V Y A M R K R S panF--. NAPCO CACC 000 T
FIG. 3. ThepromoterregionofpanF. The sequence shows the 3' end of accC and thebeginningofpanFtaken from references 28 and 34. The sequences of theoligonucleotidesused in thepromoter-mappingexperimentsareboxed. Thelocations of the mRNA 5' ends(B1andB2)upstream
of panFare shownbytheblack dots. Thepossible -10 boxof promoterB1 isunderlined. A stem-loopstructure after accC is indicated by convergentarrows.Apotential ribosome-bindingsite forpanFis shaded.
1636 -1018 -517 506 396 344 298 on December 7, 2016 by INIST-CNRS http://jb.asm.org/ Downloaded from
7184 VANET ET AL.
reversetranscriptase and the5'-end-labeled NAPCO
oligonu-cleotideontotal E.coli RNAextractsdetected 5' mRNA ends at the same positior. as found by SI protection experiments
(Fig. 4B, lanes I to3).
The 1,463-bp PCR-synthesizedprobe used in these
experi-mentscarried plasmid-derived DNAsequences at the RBP22
oligonucleotide end, which are not homologous to mRNA transcripts in this region. Protection of fragment D, with a
length of 1,028 nucleotides and corresponding to the accC panF region of the labeled probe, demonstrated thatthere is
somereadthrough from the upstream (accBC) operon.
Quan-titation ofthese bands in mRNA from a plasmid-free strain
suggests that readthrough from accBCcan account for 10to
20% of panFtranscription.
Inspection of the sequence of Fig. 3 shows that the 5' end
localized to nucleotide 263 (position Bi) is preceded by a
possible -10consenstssequence,TACAAT(nucleotides 250
to 255), but no sequei 2e homologous to the -35 consensus
sequence is located at a reasonable distance (16 to 19 bp)
upstream. The second 5' mRNA end corresponding to the
longer transcriptwaslocatedatnucleotide 213 (position B2).It fallsnear a series of T's at the base ofastem-loop structure which could be a transcription terminator for the upstream accC gene. Reverse transcriptase, notoriously sensitive to
secondarystructureswithin an mRNA, might stop atthis site and produce an artifactual 5' end. The fact that the same 5'
endwas detectedby theS1 protectiontechnique,which is less
sensitive to secondary-structure artifacts, suggests that the 395-nucleotide band detected does, indeed, correspond to a
discrete mRNA species present in vivo. Since we found readthrough transcripts from accBC into panF, it is possible that the B2 RNA is derived from these longer transcripts rather thandue to adenovo transcription start.
Toresolve thisproblem,weperformed in vitro transcription
with [ot-32P]UTP onthe unlabeled AGFIN-NAPCO fragment
(Fig. 3). Thetranscription products observedareshown inFig.
5A, lane 1: the strongest transcript (band A) is about 220 nucleotides long, theone nearthetopof the gel (bandC) is a
copy of the full-length template, and two or three RNA
transcripts of 350 to 420 nucleotides (position B) are also
evident.
The 220-nucleotide RNA (A) is a transcript made in the
opposite direction (compared with the direction of transcrip-tion ofaccC,panF, andprmA). Thiswasdemonstrated by using
another PCR-generated fragment between oligonucleotides
BGFIN and NAPCO asthetemplate for in vitro transcription
(Fig. 3). Band Awasreplaced byaband 34 nucleotides shorter.
Theclusterof RNA transcripts ofabout350to420nucleotides
(B) was unaffected, while the full-length template transcript
(C) became slightly shorter (data not shown). These
experi-ments localized the 5' end of the backward transcript to
aroundnucleotide 220onthesequenceof Fig. 3,i.e., within the
putative transcription terminator for accC. We are currently investigating whether this transcript exists in vivo.
Transcripts in the size range of350 to 420nucleotides (B)
areconsistent with the size of theS1-protected fragments and
primer extension products (340 and 395 nucleotides) detected with in vivo mRNA. To confirm that the in vitro transcripts found in position BonFig.5A, lane 1,really correspondtothe 5' ends mapped in Fig. 4,we performed reverse transcriptase
mapping ontheunlabeled products of in vitro transcription of
the AGFIN-NAPCO template. The primer used was the 32P-labeledFNAPoligonucleotide, which hybridizesnearer to the panF promoter region than does NAPCO (Fig. 3). The primer extension productsobtained areshown on Fig. 5, lane
2. Two major bands about 64 and 112nucleotides long, which
C -> A -> 1 p:: 147 = 622 527 404 309 242 238 -217 201 190 180 160 147 123 = 110 90= 76 = 67-_ 123 110 34=I =90 2 a sC 4:i IEit I B2 <- Bl accC (?F F panF ---4-- DNAtemplate
AGFIN FNAP NAPCO
A _2 RNAsynthesized
~~~~~~~~&C invitro
FIG. 5. Determination of the promoter upstream ofpanFby in
vitro transcription. (A) Lane 1, in vitro transcription on the
PCR-generated NAPCO-AGFIN fragment with[a-32P]UTP. The originof thethreemajor transcripts (A, B, and C)isdiscussed in thetext.Lane
2,unlabeledinvitrotranscriptionproductsfromtheNAPCO-AGFIN fragmentwere subjected toprimer extension performed with5'-end
32P-labeled oligonucleotideFNAPasthe primer. The extension prod-ucts(BI andB2)areindicated. Themolecular sizes (in basepairs)of labeled DNAmarkers(pBR322 digested with MspI)areindicated.(B) Diagramatic representation ofthe transcription results.
correspond to start sites around nucleotides 261 and 212 on Fig. 3,werefound. Thesetermini agreeextremelywell withthe location of the 5' ends,Bi and B2, detected previouslyon in vivo-synthesized mRNA. These data are consistent with the existence oftwopromoterregions for theexpression ofpanF. Thestronger startsite in vivo and invitro, BI,coincideswith a sequence showing homologyto the -10 consensus sequence
and seemslikelytocorrespond to arealpromoter.
Our datado notallowus tobe categorical in ourexplanation
of the B2 transcript. The lack of any promoter consensus sequLence and its location within a stem-loop structure makes
us suspect that B2 is derived from the accBC readthrough transcripts, either as a result ofendonucleolytic processingor as an experimental artifact due to thesecondary structure in the region. Such a structure could well cause reverse
tran-scriptasetoabort prematurely. This isequallypossible onthe
in vivo- and the invitro-synthesized transcriptsusedhere.The
in vitro transcription experiment produced an appreciable
amount of a full length, end-to-end transcript (C in Fig. 5A,
J. BACTERIOL.
on December 7, 2016 by INIST-CNRS
http://jb.asm.org/
12 3 4
-i
IJ
I
536 H 4-490 G 4-460 F3*4 F3205 +4- 405F2 380 F1 iThT-]]
D)UG18Fp_anF
IprmApm. - - Simappingprobe
REV17 ART'
4-MP reversetranscriptase primer AMRP*
F2 S1resistant
F3 fragments H I
FIG. 6. Mapping of the 5' mRNA ends upstream of prmA. (A)SI analysis.Theprobe used for S1mapping was the 536-bp REVI 7-ART PCR-generated fragment 32p labeled at the ART 5' end. It was hybridized with RNA extracted from strain JM109 (lane 2, 30 ,ug) or
JMI09(pAF3)(lane 3, 15jig;lane4, 30jig)orwith tRNA (lane 1, 30
jig).
TheSI-resistantfragments were electrophoresed on a 6%poly-acrylamide-7Mureagel. Thepositionsoflabeled DNA molecular size markers(in base pairs) are shown on the left. The lengths of protected DNAfragments Fl,F2, F3, G, and H are indicated on the right. (B) Primerextension. RNA (15 jiginlane 1 and 30 jLgin lane2) isolated from JM109(pAF3)wasused for primer extension analysis with 5'-end 32P-labeledoligonucleotideAMRP astheprimer. Lanes A, C, G, and
T are sequencing reactions with pAF3 DNA and oligonucleotide AMRP as the primer; the sequence corresponds to the noncoding strand. Molecular size markers (pBR322 digested with MspI) are
shownontheright in base pairs, and the sizes of the major extension products areindicatedontheleft. (C) Diagramaticrepresentation of the combined results.
lane 1) in which the stem-loop structurecould also form and block reverse transcriptase. Whatever itsorigin, B2isaminor componentcomparedwith the Bi transcript.
The prmA transcript. The data from the fusions panF
prmA-lacZ,panF-lacZ, andprmA-lacZ suggest that panF and prmA are cotranscribed. We looked for the polycistronic
mRNAbySI mapping withalongprobe covering bothgenes. We actually detectedverylittlebicistronic panF-prmA mRNA
but did find evidencefor mRNA5' endsimmediatelyupstream
of prmA.Toconfirm theexistenceoftranscriptstraversing the panF-prmA junction and localize the 5' mRNA ends
immedi-atelyupstreamofprmA,anS1mappingstrategysimilartothat
employed to identify the panF mRNAs was used. SI probe
REV17-ART, labeled at the ART oligonucleotide, carries
plasmid-derived sequences at the REV17 end which do not hybridize to transcripts from the panFprmA region of the chromosome(Fig.6C).Useof thisprobeandmRNAextracted
from JM109carryingor not carryingplasmid pAF3 produced
aseries ofprotectedfragmentswithlengthsof about 380
(Fl),
405(F2), 460(F3), and 490(G) nucleotides, aswellas abandcorresponding to the reannealing of the
probe (H).
Someadditional, shorterbandswerealsoobservedwith mRNA from
apAF3-carrying strain (Fig. 6A).The G bandcorresponds to
protection of the entirepanFprmA region of the probe and is
therefore equivalent tothebicistronicpanF-prmA transcript.
Reverse transcriptase mapping with the 5'-end-labeled
AMRPoligonucleotideas theprimer (Fig. 6C)produced four transcripts in the sizerangeof 125to150nucleotides (approx-imate sizes: 125, 130, 140, and 145 nucleotides) plus a longer transcript of205nucleotides(Fig.6B, lanes 1and2).These5' endswere precisely located on the sequenceofFig. 2.
Com-parison with the results of theSI experiment(Fig. 6A) showed that
SI-protected
fragment Fl corresponds to the 125-nucle-otide reverse transcriptase product, the 145-nucleotide bandcorrespondstoF2, and the 205-nucleotide bandcorrespondsto F3. The 130- and 140-nucleotide transcripts were always detected by primer extension, but their relative intensities varied between experiments, suggesting that they could be artifacts duetosecondarystructure.Todetermine whetherany
of these 5' ends correspond to promoters, we performed in
vitrotranscriptiononpurified DNAfragment CHIEN-AMRP.
We were not able to detect any discrete band, except one
corresponding to the full length of the template (data not shown). Since no significantpromoteractivitywasdetected in vitro or in vivoin this region,we conclude thatthe 5' ends of
mRNAs
Fl,
F2, and F3are generated byinvivoprocessingofthepanFprmA transcript.
Comparison of the intensities of the three
SI-protected
fragments,
Fl,
F2, and F3, with that of the Gband,represent-ingthebicistronicpanFprmAmRNA,suggeststhat75% of the
chromosome-derived transcripts are processed (Fig. 6A, lane 2). In pAF3-derived transcripts (Fig. 6, lanes 3 and 4) the
processed-to-bicistronic
transcript ratio decreased to 50%,possibly because of limiting amounts of the processing
en-zymes.
3' mRNA extremities downstream of prmA. The DNA sequence downstream of
prmA
(Fig. 2) isvery AT rich(72%
AT if considered from the stop codon ofprmA to the first codon of orfl, of which the central 220 bases are 78% AT) compared with theprmA structural gene
(54%
GC).There is no strong hairpinstructure followed by a series ofTresidues characteristic of prokaryotic terminators in the intergenicprmA-orfl
region.Twopromoter sequences,atleastsixstrongfactor for inversion stimulation (Fis)-binding sites, and a cAMP receptor protein
(CRP)-binding
site have beenlocal-ized in this AT-rich region, and the DNA exhibits inherent
curvature (8, 42).
We used
SI
mapping to determine the fate ofprmAtran-scripts after the structural gene. The fragment used covered the 3' end ofprmA andwaslabeledatthe
IHindIII
site(Fig. 2).
The other, unlabeled end was located within the
orfi
gene(corresponding
tooligonucleotide IFRO; Fig. 2).
WhenmRNA froma
plasmid-bearing
strain[IBPC5321(pAF3)]
wasused to enhance the amount ofprmA
transcripts,
thisprobe
detectedamajorbandofabout 185nucleotides andaseries of longertranscripts (Fig. 7A, lanes4 and 5). The same 3' ends weredetectablein RNAextracts
prepared
fromaplasmid-free
strain
(Fig.
7A,lane 6).The major3' end(open
squareinFig.
2)is located100nucleotides downstream of theprmAgeneand
mapstothe distal side ofa small
palindromic
sequencewhich isnot,however, followedbyaseries ofTresidues(nucleotides
1098 to 1111 of the sequence in
Fig.
2).
The ladder of bandscorresponding tolonger
transcripts
of200to 250 nucleotides showsaperiodicity
ofabout 10nucleotides.Thus,
these 3'ends onthe mRNAcorrespond
topositions
onthe DNAtemplate
separated byaboutone turnofthe B DNAhelix
(10.5
bp).
Thisspacing
suggests that 3' endsaregenerated
by
periodic
termi-nation of
transcription
atin-phase
positions
ontheDNA.Sincethis
region
isinherently
bent because of the presence ofon December 7, 2016 by INIST-CNRS
http://jb.asm.org/
7186 VANET ET AL.
1 2 3 4 5
HindlII
E . ',~~~~~~~~~~~~~~~~~~~~~
6 M transcribe through the intergenic prmA-orfl region into the
orfl-fis
operon.However,
the contribution ofpanF-prmA
transcription to orfl-fis expression in vivo is not likely to be verysignificant invivo.
517 506 396 344 298 220 201 154 134
|
panF IprmA|
-
OS
|
V CRP V IVIII Hindlill 4. Simappin( I IFRO probe S1 resistant fragments i-I -I IIFIG. 7. SI analysis ofthe 3' RNA endsofprmA. (A) The probe usedwasthe477-bp HindIll-IFRO fragment3' 32Pendlabeledatthe
HindIll end. Theprobewashybridizedwith 15or30 jLgof RNAfrom
IBPC5321(pBR322) (lanes 2 and 3), 15 or 30 jLg of RNA from
IBPC5321(pAF3) (lanes4and5),50jLgof RNA fromIBPC5321(lane
6),or30 jLgof tRNA(lane 1).TheSI-resistantproductswereanalyzed
on a5%polyacrylamide-7Mureagel.Thepositions ofmolecular size
markers are indicatedon the right inbase pairs,and the sizeof the
major SI-resistant fragment is given on the left. (B) Diagrammatic
representation of theexperiment.CRP,cAMPreceptorprotein
bind-ing site; ItoVI,Fisbinding sites.
poly(A) tracts and has several Fis-binding sites and a cAMP
receptor protein-binding site (Fig. 7B), which are known to
affectDNAstructure(8),wesuspectthat the 3' endsobserved hereare related tothe topology of the region rather than due to classical transcription termination. Analysis of the mRNA
fromthepAF3plasmid-carryingstrain revealed protection ofa
fragment just shorter than thefull-length probe (Fig. 7, lanes 4 and 5). This isbecause the insert in pAF3, which endsat the NcoI site, is slightly shorter than the SI probe which was
synthesized with oligonucleotide IFRO (Fig. 2). The
protec-tion of thisshorterfragment shows thatacertainpercentageof RNApolymerase moleculesontheplasmid DNA templatecan
DISCUSSION
Althoughcotranscription isaconvenient strategy for coregu-lation of genes with a common function, there are acertain number ofexamplesofcotranscription of genes withunrelated
functions,
e.g., the rpsU dnaG rpoD operon that encodesribosomal protein S21, DNA primase, and RNA polymerase
subunit sigma 70 (12). In thiswork, we show that the gene necessary for posttranslational modification of a ribosomal
protein-the methyltransferase ofLi1-is cotranscribed with the gene for pantothenate permease. Pantothenate is a
pre-cursorfor coenzyme A. The genepanFisnecessaryonlywhen
pantothenatemustbetaken up from theenvironment,because
wild-type E. coli can synthesize pantothenate from 1-alanine and pantoateby usingthe enzyme encodedby panC (see Fig.
I inreference 62). Weare currently investigatingthe
implica-tions of coexpression of these two genes, panF and prmA,
which have verydifferent functions.
prmA is the first gene that controls the methylation of a
ribosomal protein to be sequenced. A mutation (prmB) that affects themethylationofL3 has beenisolated and mappedto
50minonthe E.coli chromosome (14, 37).We are unaware of
information, either genetic or biochemical, concerning the other ribosomal protein methyltransferases. The genedosage
effect on L1I methyltransferase activity observed with the
prmA gene carried on amulticopy plasmid, the fact that the size ofthe protein encoded by prmA agrees with themolecular weight of the partially purified enzyme (4, 13), and the presence ofashort amino acid motif found in many methyl-transferases ofdifferent specificities (26) argue thatprmA is the structural gene for LII methyltransferase. A detailed
analysisofmutations within the prmA gene and their effecton
LII methylation also support this conclusion (64).
The prmA ORF appears to start at the ATG codon I1
nucleotides downstream of thepanFtermination codon. Pro-teinfusionswith a truncated lacZ gene were constructed at the
BamHI site 11 nucleotides downstream of this ATG. The
hybrid gene which carries the first four codons of prmA in phase with the lacZ gene, when present on a plasmid or bacteriophage A vector, produces blue colonies on 5-bromo-4-chloro-3-indolyl-3-D-galactopyranoside (X-Gal) plates,
show-ingthat the first ATG doesfunction as an initiation codon(63).
A second ATGcodon, 22 amino acidsdownstream, shows no
betterribosome-bindingsite. The short distance betweenprmA andpanF, together with the lack of any recognizable Shine-Dalgarno sequence, exceptGAG, suggests thatthe twogenes aretranslationallycoupled. Efficienttranslationalcouplinghas been observed in various bacterial systems in which the termination and initiation codons canoverlapor beseparated bytensof nucleotides (1, 24,36).
The datapresentedhereshowthat, bothinvivoandinvitro, genes panF and prmA are cotranscribed from a promoter locatedupstream of the panFstructural gene.Thebest candi-date for the panF promoter isthatwhich gives risetotranscript BI in Fig. 3. Transcripts from BI carry a 255-nucleotides
untranslatedleaderupstreamof the panFstructuralgene. This long mRNA leadermight be involvedin regulationof expres-sion of theoperon,althoughwehavenotidentifiedany obvious
regulatory elements in thisregion.
We alsodetected, by different techniques, a5' mRNA end located at the base of astem-loop structure (B2inFig. 3). Our 185 J. BAC-FERIOL. lr ,., ig on December 7, 2016 by INIST-CNRS http://jb.asm.org/ Downloaded from
present data cannot unambiguously assign this 5' end to a promoteractivity. If it doesnotcorrespondto apromoter,then it derives from posttranscriptional processing of the longer transcript whichwas detected in small amounts. These longer transcripts presumably initiate from the upstream accBC operonandmust readthrough theputative terminator located afteraccC (nucleotides200 to 220 inFig. 3), which in thiscase
could serve as an mRNA-processingsite.
Another potential regulatory device is suggested by the detection of a promoter activity in vitro thatgives rise tothe synthesis ofadivergentmRNAinthissameregion. The length of thetranscript detected (AinFig.5A, lane 1)locates itsstart
pointvery near to B2(Fig. 3) butontheopposite strand.There are no sequences strongly homologous to the consensus -35 and -10 promoter sequences in thisregion.We are currently investigating whether this promoter is functional in vivo and what the length of the putative antisense mRNA is. It is interesting that there is a long ORF of 314 amino acids downstream ofthis promoter,completely contained within the accCstructural gene but expressed from theopposite strand.
Despitethelack ofanypromoteractivity, in vivoorinvitro,
expressing a monocistronic prmA transcript, we did locate several mRNA 5' ends upstream of theprmA structural gene
which lie mostly within the 3' end of the panF structuralgene
transcript. The existence of a discrete prmA transcript was suggested by Northern (RNA) analysis (63). The use of a probe internaltotheprmA genedetectedbasically asmearof hybridizing material with lengths ofup to 1,800 nucleotides. There was also a rather diffuse band of about 1,200 nucleo-tides. This is sufficient to cover the entireprmA gene but not bothpanF andprmA. The use ofapanF-specificprobe failed
to detect this or any other discrete bands. The simplest
interpretation of these results is that thebicistronicpanF-prmA
transcript is subject to posttranscriptional processing which generates an mRNAwhichjustcoverstheprmA gene. Weare currentlyinvestigatingthishypothesis. Itisinterestingthatany
endonucleolytic processing within the 3' end of the panF
mRNA will generate a truncated panF mRNA incapable of directing the synthesis of a full-size PanF protein. The
n-ga-lactosidase activities of the operon fusions (Table
2) provide
some indication of the relative expression levels of the two genes. The lower level of ,B-galactosidase
activity
of theprmA-lacZoperonfusion than of thepanF-lacZoperonfusion
suggeststhatsomedown regulationof
expression
occurs.This effect could be relatedtotheputative processingeventsthatwehave detected in thepanF-prmA mRNA.
ACKNOWLEDGMENTS
We thank Marie-France Guerin for performing the methylation tests,RichardBuckinghamandMathiasSpringerforcriticalreadingof themanuscript, MarianneGrunberg-Managoforconstantinterest,P.
LeChien for encouragement,SylvieBlanda forpatience,andareferee forconstructivecriticism.
This workwassupported bygrantsfrom theCNRS, INSERM,Paris 7 University, the A.R.C., and the Fondation pour la Recherche
Medicale.A.V. wassupported byagrantfrom theMRT.
REFERENCES
1. Aksoy, S., C. L. Squires, and C. Squires. 1984. Translational
coupling of the trpB and trpA genes in the Escherichia coli
tryptophanoperon.J.Bacteriol. 157:363-367.
2. Alix, J. H. 1988. Post-translational methylations of ribosomal
proteins.Adv. Exp.Med. Biol.231:371-385.
3. Alix, J. H. 1989. A rapid procedure for cloning genes from A librariesby complementation of E. coli defectivemutants:
appli-cation to the fabE region of the E. coli chromosome. DNA 8:779-789.
4. Alix, J. H.,and D.Hayes.1974.Propertiesof ribosomes and RNA synthesized byEscherichiacoligrown in the presenceofethionine. III. Methylatedproteinsin50Sribosomes of E. coliEA2. J. Mol. Biol. 86:139-159.
5. Alix, J. H.,D.Hayes, J.F.Lontie,C.Colson,A.Glatigny,and F.
Lederer. 1979.Methylatedamino acids inribosomalproteinsfrom Escherichiacolitreated with ethionine and from amutantlacking
methylationofprotein Lii. Biochimie 61:671-679.
6. Ambulos,N.P.,E.J.Rogers,Z.Alexieva,and P. S.Lovett.1988. Induction of cat-86bychloramphenicoland amino acid starvation in relaxedmutantsofBacillussubtilis.J.Bacteriol. 170:5642-5646. 7. Armstrong, I. L., and W. P. Tate. 1978. Requirement for the Escherichia coliribosomal proteinLii inpeptidechain termina-tion. J. Mol. Biol. 120:155-166.
8. Ball,C.A., R.Osuna,K.C.Ferguson,and R. C.Johnson. 1992. Dramaticchangesin Fis levels upon nutrientupshiftinEscherichia coli. J.Bacteriol. 174:8043-8056.
9. Baxter, R.M.,and N.Zahid. 1986. L16,abifunctionalribosomal
proteinand theenhancingeffect of L6 andLI1.Eur. J. Biochem. 155:273-277.
10. Bolivar, F., R. L. Rodriguez, P. J. Greene, M. C. Betlach, H. L.
Heyneker, H. W. Boyer, J. H. Crosa, and S. Falkow. 1977.
Construction and characterization ofnew cloningvehicles. II. A
multipurposecloningsystem. Gene 2:95-113.
11. Bradford, M. M. 1976. A rapid and sensitive method for the
quantitation of microgram quantities of protein
utilizing
the principleofprotein-dyebinding.Anal. Biochem.72:248-254. 12. Burton, Z. F., C.A. Gross,K. K.Watanabe,and R. R.Burgess.1983. The operon that encodes the sigma subunit of RNA polymerasealsoencodes ribosomalproteinS21and DNA
primase
in E. coli K12. Cell32:335-349.
13. Chang,F.N.,L. B.Cohen,I.J.Navickas,and C. N.Chang. 1975. Purification andpropertiesofaribosomalproteinmethylasefrom EscherichiacoliQ13. Biochemistry14:4994-4998.
14. Colson,C.,J.Lhoest,and C.Urlings.1979. Genetics ofribosomal protein methylation inEscherichiacoli. III. Map position oftwo
genes,prmAandprmB,governing
methylation
ofproteins
LI1and L3. Mol. Gen. Genet. 169:245-250.15. Colson, C.,and H.0.Smith. 1977.Genetics of ribosomal
protein
methylation inEscherichiacoli. I.Amutantdeficient in
methyla-tion ofproteinLii. Mol. Gen. Genet. 154:167-173.16. Dabbs, E. R. 1978. Mutational alterations in 50
proteins
of the Escherichiacoli ribosome. Mol. Gen. Genet. 165:73-78.17. Dabbs, E. R. 1980. The ribosomal components
responsible
for kasugamycin dependence, and its suppression, in a mutant of Escherichiacoli. Mol. Gen. Genet. 177:271-276.18. Dognin,M.J.,and B.Wittmann-Liebold. 1980. Identification of
methylated amino acidsduringsequence
analysis.
Application
tothe Escherichia coli ribosomal
protein
LI1.Hoppe-Seyler's
Z. Physiol. Chem. 361:1697-1705.19. Dognin,M. J.,and B. Wittmann-Liebold. 1980. Purification and
primarystructure determination of the N-terminal blocked
pro-tein, LI1, from Escherichia coli ribosomes. Eur. J. Biochem. 112:131-151.
20. Egebjerg, J.,S. R.Douthwaite,A.Liljas,andR. A.Garrett. 1990.
Characterization of the binding sites of
protein
L1I and theL10.(L12)4 pentameric complex in the GTPase domain of 23S ribosomal RNA from Escherichiacoli. J. Mol. Biol.213:275-288. 21. Gotz,F.,C.Fleischer,C.L.Pon,andC.0.Gualerzi. 1989.Subunit
association defects in Escherichia coli ribosome mutants
lacking
proteins S20 and LI1. Eur. J.Biochem. 183:19-24.
22. Grosjean, H., and W. Fiers. 1982. Preferential codon usage in
prokaryotic genes: the
optimal
codon-anticodon interactionen-ergy and theselective codon usage in
efficiently
expressed
genes. Gene 18:199-209.23. Hampl, H., H. Schulze, and K. H. Nierhaus. 1981. Ribosomal
components from Escherichia coli 50S subunits involved in the reconstitution of
peptidyltransferase
activity.
J. Biol. Chem.256:2284-2288.
on December 7, 2016 by INIST-CNRS
http://jb.asm.org/
7188 VANET ET AL.
24. Hellmuth, K., G. Rex,B.Surin,R.Zinck,andJ.E.G.McCarthy. 1991.Translationalcoupling varying inefficiencybetween different pairs of genes in the central region of the atp operon of Esche-richia coli. Mol. Microbiol. 5:813-824.
25. Hitz, H., D. Schafer, and B. Wittmann-Liebold. 1975. Primary
structureofribosomalprotein S6 from the wild type anda mutant
of Escherichia coli. FEBS Lett. 56:259-262.
26. Ingrosso, D., A. V. Fowler, J. Bleibaum, and S. Clarke. 1989.
Sequence of theD-aspartyl/L-isoaspartyl protein methyltransferase
from humanerythrocytes. J. Biol. Chem. 264:20131-20139. 27. Isono, K., J. Krauss, and Y. Hirota. 1976. Isolation and
charac-terization of temperature-sensitive mutants of Escherichia coli with altered ribosomalproteins. Mol. Gen. Genet. 149:297-302. 28. Jackowski, S., and J. H. Alix. 1990. Cloning, sequence, and
expression of the pantothenatepermease (panF) gene of Esche-richia coli. J. Bacteriol. 172:3842-3848.
29. Jinks-Robertson, S., and M. Nomura. 1987. Ribosomes and tRNA, p. 1358-1385. In F. C. Neidhardt, J. L. Ingraham, K. B. Low, B. Magasanik, M. Schaechter, and H. E. Umbarger (ed.),
Escherichia coli andSalmonella typhimurium:cellular and molec-ularbiology, vol. 2. AmericanSocietyforMicrobiology,
Washing-ton, D.C.
30. Kang,W.K.,T.Icho, S. Isono,M.Kitakawa,and K. Isono. 1989. Characterization of the gene rimKresponsiblefor theaddition of glutamicacid residuestotheC-terminus of ribosomalproteinS6 in Escherichiacoli K12. Mol. Gen. Genet. 217:281-288.
31. Kazemie, M. 1975. TheimportanceofEscherichiacoliribosomal proteinsLI,LII and L16 for the association of ribosomal subunits and the formation of the 70-S initiationcomplex.Eur. J.Biochem. 58:501-510.
32. Koch,C., J. Vandekerckhove,and R. Kahmann.1988.Escherichia coli host factor for site-specific DNA inversion: cloning and characterization of the fis gene. Proc. Natl. Acad. Sci. USA 85:4237-4241.
33. Kohara, Y., K. Akiyama,and K. Isono. 1987.Thephysicalmapof the wholeE. colichromosome: application ofa newstrategyfor
rapid analysisandsortingofalargegenomic library.Cell 50:495-508.
34. Kondo, H.,K.Shiratsuchi,T.Yoshimoto, T. Masuda,A.Kitazono,
D.Tsuru,M.Anai,M.Sekiguchi,and T. Tanabe.1991.Acetyl-coA carboxylase from Escherichia coli: gene organization and nucle-otide sequence of thebiotincarboxylasesubunit. Proc. Natl. Acad. Sci. USA 88:9730-9733.
35. Lederer, F., J.H.Alix,and D.Hayes. 1977.N-trimethylalanine,a
novel blocking group found in E. coli ribosomal protein L1. Biochem.Biophys. Res.Commun. 77:470-480.
36. Lesage,P.,C.Chiaruttini,M.Graffe, J. Dondon,M.Milet, andM.
Springer. 1992.MessengerRNAsecondarystructureand transla-tionalcouplingin the Escherichiacoli operonencodingtranslation initiation factor IF3 and the ribosomalproteins, L35 and L20.J.
Mol.Biol. 228:366-386.
37. Lhoest, J.,andC. Colson.1981.Cold-sensitive ribosomeassembly
in an Escherichia coli mutant lacking a single methyl group in ribosomalprotein L3. Eur. J.Biochem. 121:33-37.
38. Li, S.-J.,andJ.E.J. Cronan. 1992. The gene encodingthebiotin carboxylase subunit of Escherichia coli acetyl-coA carboxylase. J. Biol. Chem. 267:855-863.
39. Miller, J.H. 1972.Experiments in molecular genetics, p. 352-355. ColdSpring Harbor Laboratory Press, Cold Spring Harbor,N.Y.
40. Naaktgeboren, N.,P.Schrier,W.Moller,andH.0.Voorma. 1976.
Theinvolvement ofproteinL1iin thejoining of the 30-S initiation
complextothe 50-Ssubunit. Eur. J. Biochem. 62:117-123. 41. Neidhardt, F. C., P. L. Bloch, and D. F. Smith. 1974. Culture
medium for enterobacteria. J. Bacteriol. 119:736-747.
42. Ninnemann, O.,C.Koch, and R. Kahmann. 1992. The E. coli fis promoter is subject to stringent control and autoregulation. EMBOJ.11:1075-1083.
43. Ochi, K. 1990. Arelaxed (rel) mutant of Streptomyces coelicolor
A3(2) with a missing ribosomal protein lacks the ability to
accumulate ppGpp,A-factor andprodigiosin.J. Gen. Microbiol. 136:2405-2412.
44. Ochi,K.1990.StreptomycesrelCmutantswithanaltered ribosomal
protein ST-L1IandgeneticanalysisofaStreptomyces griseus relC
mutant.J. Bacteriol. 172:4008-4016.
45. Parker, J.,R. J. Watson,J.D. Friesen, and N.P. Fiil. 1976. A relaxedmutantwith analtered ribosomalprotein LI1.Mol. Gen. Genet. 144:111-114.
46. Plumbridge, J. A., J. Dondon,Y. Nakamura, and M.
Grunberg-Manago. 1985. Effect of NusAproteinonexpressionof thenusA, infBoperon inE. coli.Nucleic Acids Res. 13:3371-3388. 47. Plumbridge, J. A., and M. Springer. 1983. Organization of the
Escherichia coli chromosome around the gene for translation initiation factor IF2(infB)andatranscription termination factor
(nusA).J.Mol. Biol. 167:227-243.
48. Ryan,P.C.,and D. E. Draper. 1991. Detectionofakeytertiary interaction in thehighlyconserved GTPasecenteroflargesubunit ribosomal RNA. Proc. Natl. Acad. Sci. USA 88:6308-6312. 49. Salser, W.,R. F.Gesteland,and A. Bolle.1967.In vitrosynthesisof
bacteriophage lysozyme.Nature(London)215:588-591. 50. Sambrook,J., E. F. Fritsch, and T. Maniatis. 1989. Molecular
cloning: alaboratorymanual,2nd ed. ColdSpringHarbor Labo-ratoryPress, Cold Spring Harbor,N.Y.
51. Seto, D. 1990. An improved method for sequencing double strandedplasmid DNAfrom minipreps usingDMSOand modi-fiedtemplatepreparation. Nucleic Acids Res. 18:5905-5906. 52. Shine, J., andL.Dalgarno. 1974.The 3' terminal sequence of E.
coli 16S ribosomal RNA: complementarity to nonsense triplets
and ribosomebindingsites. Proc. Natl. Acad. Sci. USA 71:1342-1346.
53. Silhavy,T.J.,M. L. Berman, and L.W. Enquist. 1984.
Experi-mentswith genefusions, p. 162-165. ColdSpringHarbor
Labo-ratoryPress, ColdSpringHarbor,N.Y.
54. Simons, R. W., F. Houman, and N. Kleckner. 1987. Improved single and multicopy lac-based cloning vectors for protein and operonfusions. Gene 53:85-96.
55. Sprengart, M. L., H. P. Fatscher, and E. Fuchs. 1990. The initiation of translation inE. coli: apparent basepairingbetween the 16S rRNA and downstream sequences of the mRNA.Nucleic Acids Res. 18:1719-1723.
56. Stoffler, G., E.Cundliffe,M. Stofller-Meilicke, and E. R. Dabbs.
1980. Mutantsof Escherichia colilackingribosomalprotein LI1.J. Biol. Chem. 255:10517-10522.
57. Stoffler, G.,R.Hasenbank,andE. R. Dabbs. 1981.Expressionof the L1-LI operon in mutants of Escherichia coli lacking the ribosomalproteins Li andLi1. Mol. Gen. Genet. 181:164-168. 58. Tate,W. P.,M.J. Dognin,M.Noah,M. St8ffler-Meilicke,andG.
Stoffler. 1984. The NH,-terminal domain of Escherichia coli ribosomalprotein LI1:its three-dimensional location and its role in thebindingof release factors 1 and 2.J.Biol. Chem. 259:7317-7324.
59. Tate,W.P.,H.Schulze,andK. H.Nierhaus.1983. The Escherichia coli ribosomal protein LI1 suppresses release factor 2 but
pro-motesthe release factorI activities inpeptidechaintermination.
J. Biol. Chem.258:12816-12820.
60. Thompson, J., E. Cundliffe, and M. Stark. 1979. Binding of thiostrepton to acomplexof 23-S rRNA with ribosomal protein LI1. Eur. J. Biochem. 98:261-265.
61. Uzan, M., R. Favre, and E. Brody. 1988. A nuclease that cuts specificallyin theribosomebindingsiteofsomeT4 mRNAs. Proc. Natl. Acad. Sci. USA 85:8895-8899.
62. Vallari, D. S., andC. 0. Rock. 1985. Pantothenate transport in Escherichia coli. J. Bacteriol. 162:1156-1161.
63. Vanet,A.Unpublishedresults.
64. Vanet, A., J.A.Plumbridge,M.-F.Guerin,andJ.-H. Alix.
Unpub-lished data.
65. Yanisch-Perron, C.,J. Vieira, and J. Messing. 1985. Improved M13phage cloningvectorsandhost strains: nucleotide sequences of theMl3mpl8andpUC19vectors.Gene33:103-119.
J. BACTERIOL.
on December 7, 2016 by INIST-CNRS
http://jb.asm.org/



![FIG. 5. Determination of the promoter upstream of panF by in vitro transcription. (A) Lane 1, in vitro transcription on the PCR-generated NAPCO-AGFIN fragment with [a-32P]UTP](https://thumb-eu.123doks.com/thumbv2/123doknet/14601817.544076/8.918.508.769.101.618/determination-promoter-upstream-transcription-transcription-generated-napco-fragment.webp)

