Algorithm engineering for optimal alignment of protein structure distance matrices
Texte intégral
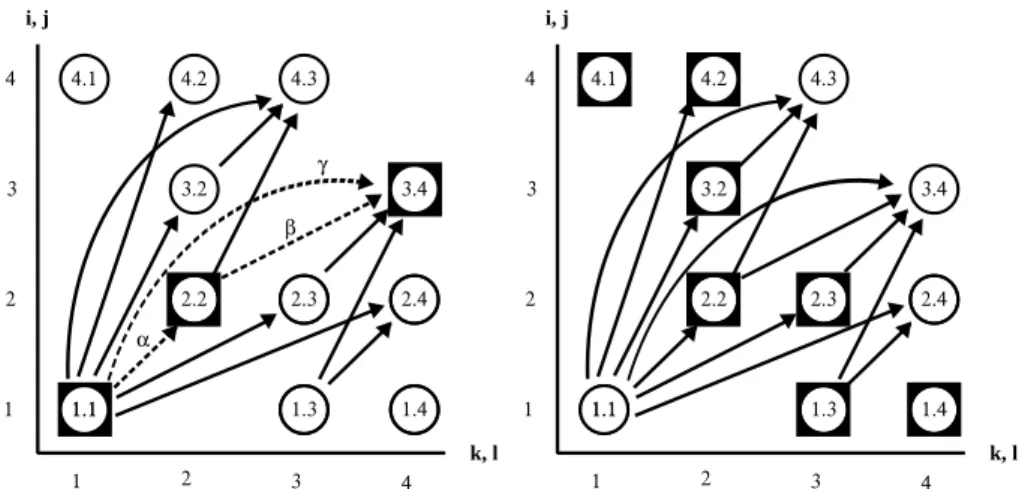
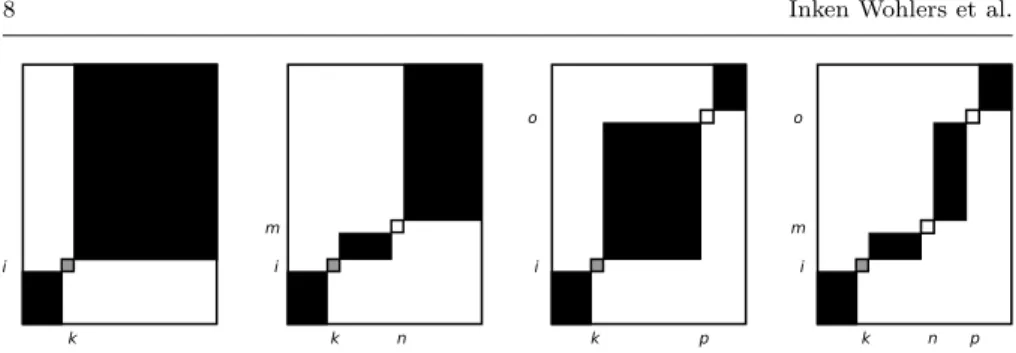
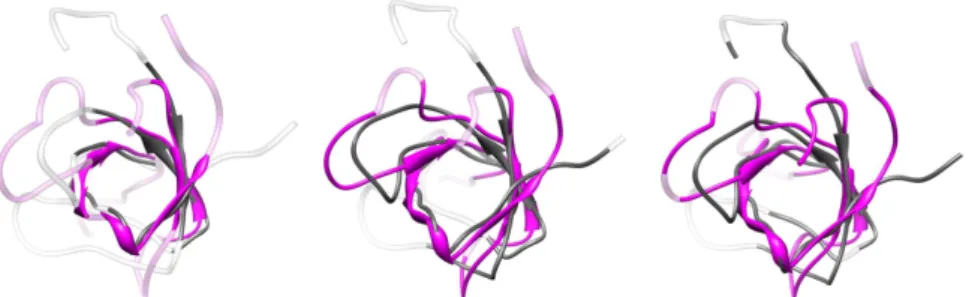
Figure
Documents relatifs
MRFalign showed better alignment precision and recall than HHalign and HMMER on a dataset of 60K non-redundant SCOP70 protein pairs with less than 40% identity with respect to
Abstract—In this paper, we propose a successive convex approximation framework for sparse optimization where the nonsmooth regularization function in the objective function is
Deconvolution in terms of moments turns out to be quite simple if asymptotic freeness holds, and can be performed using the R- and S-transforms [8]. A lthough Gaussian ma- trices
One of the striking consequences of the celebrated Szemer´edi’s Lemma [31] states that an adjacency matrix sampled from a W -random graph model converges to the true graphon W 0 in
OPTIMAL CONTROL OF SOFT MATERIALS USING A HAUSDORFF DISTANCE FUNCTIONAL.. Rogelio Ortigosa, Jesus Martínez-Frutos, Carlos Mora-Corral, Pablo Pedregal,
A comparison between the two latter who generally permit to derive different optimal chain parenthesizations (OP’s), raised the interest of computing the matrix chain product in
Contribution of the paper We present a theorem regarding optimal decomposi- tions of a given matrix I which shows that decompositions which use as factors particular formal
This algorithm, based on circulant, Toeplitz and shift cyclic matrices, does not only reduce the computational complexity of the design process but also decreases the