Appendix S1
Details on the methods used for ploidy level determination
Because some of the taxa are similar in morphology, flow cytometric analyses were used to confirm the ploidy level of each plant population. Samples were prepared by chopping eight seeds with a sharp razor blade in a few drops of general-purpose buffer following the protocol of Loureiro et al. (2007). After this, 950ȝl of the buffer were added and the mix was incubated for approximately 1 min and then filtered through a 30-ȝm nylon filter. Afterwards, 30 ȝl of a staining solution composed of RNAse (1%) and propidium iodide (1%) was added. The flow cytometric analyses were performed with a Partec Cyflow SL cytometer equipped with a green laser functioning at 532 nm. Seeds from L. vulgare (diploid) and L. ircutianum (tetraploid) were used to adjust the laser. We were not able to accurately estimate the ploidy level of deca- and dodecaploid species with flow cytometry but because of their distinct morphology and geographic distribution we adopted published ploidy levels for these species (Vogt 1991).
Literature cited:
Loureiro, J., E. Rodriguez, J. Doležel, and C. Santos. 2007. Two new nuclear isolation buffers for plant DNA flow cytometry: A test with 37 species. Annals of Botany 100:875-888. Vogt, R. 1991. Die Gattung Leucanthemum Mill. (Compositae-Anthemideae) auf der
Appendix S2
Table S1. P h y lo g en et ic di st ance m at ri x for Le ucanthemum taxa included in this stud
y . adu = L. adustum, at r = L. atratum , cot = L. coronopifolium subsp. te nuifolium, fav = L . f a vargeri , ga l = L . gal la eci cu m, het = L. heterophyllum, ica = L. ircutianum su bsp. cantabricum, ile = L. ircutianum subsp. leucolepis, irc = L. ir cutianum subsp. ircutianum, ma x = L . maximum, mon = L. montserratianum, mop = L. monspeliense, pal = L. pa llens, vir = L. virgatum, vul = L. vulgare subsp. vulgare , vup = L. vulgare su bsp. pujiulae . tax a ad u atr cot fav gal h et ic a il e irc m a x m o n m o p p a l vi r vu l atr 0.764 cot 0.885 0.885 fav 0.734 0.776 0.908 gal 0.825 0.879 0.879 0.864 het 0.783 0.74 0.875 0.667 0.923 ica 0.683 0.81 0.887 0.776 0.829 0.844 ile 0.755 0.756 0.809 0.768 0.871 0.755 0.804 irc 0.500 0.807 0.847 0.734 0.825 0.803 0.537 0.755 ma x 0.761 0.781 0.901 0.718 0.925 0.779 0.746 0.774 0.778 mo n 0.923 0.929 0.929 0.783 0.974 0.879 0.956 0.926 0.923 0.889 mo p 0.800 0.889 0.913 0.875 0.682 0.922 0. 729 0.884 0.776 0.889 0.939 pa l 0.842 0.917 0.886 0.803 0.925 0.878 0.859 0.831 0.842 0.819 0.769 0.906 vi r 0.956 0.972 0.879 0.958 1.000 0.895 1.000 0.906 0.956 0.964 0.946 1.000 0.925 vu l 0.660 0.761 0.925 0.75 0.759 0.784 0.583 0. 75 0.604 0.815 0.946 0.711 0.915 1.000 v up 0.761 0.694 0.907 0.776 0.818 0.762 0.651 0.676 0.705 0.782 0.935 0.655 0.902 1.000 0.516
The ph y lo g enet ic di st anc e m at rix was based on se quence v ariati on at nine nuclear D NA loci (m arkers A39, B12, B20, C12, C20, C33, D18, D23, and D27 of Chapma n et al. (2007) and on five inter g enic spa cer re g
ions of the chloropla
st g enome ( cpDNA: psb A -trnH, trnL-trnF, trnC-petN, petN-psbM, and trnQ-rps16 ), the la tte r be ing c onca te n at ed into a sing le a lig nm en t (see Konow alik et al. (2 015) for details of th e s equencin g pro cedur e in diploid tax a, which wa s a lso applie d to the p o ly ploid ta x a of the p re se n t stud y ). After para me tr ising the s eque n ce substitution mode ls for the te n re g ions, ge ne (alle le ) tre es we re infe rred using B a y esia n (MrBa y es) reconstru ction me thods (Konowalik 2 014). Our ca lc ula tion of p ai rwise ph y lo g en etic dista n ce s am on g ta x a foll owe d the M atrix
Representation with Pars
imon y (MRP) met hod (B aum 1992, Ra g an 199 2, B ininda-Emonds 2004), which is o ften us ed to r econstruct su p ertr ees bas ed on sup ermatri ces r esulting from coding multiple underl y ing ph y lo g enetic tr ees (e .g . J
ohnson et al. (2012)). All ten underl
y in g g en e tr ees (with nodes collapsed a cco rding to a 0.95 posterior prob abilit y criterion) we re
transformed into multilabe
lled trees b y g iving al l accessions
and all alle
les of
a particular
tax
on the same label, c
oding
the gene tre
e topolog ies as 0/1 matrices, and merg in g the matrices into a sing le matrix b y using a sc rip t provided b y J ohnson et al. (2012 ). The resulting 0/1 supermatrix was then used to co mpu te J accard distances amo n g l abels/tax a with the software p ro g ra m MVSP v3.12f (Kovach 200 1). L iteratu re ci ted : B aum, B . R. 1992. Combining trees as a wa y of combining
data sets for ph
y lo g enetic infe re n ce , a nd th e de sirabilit y of co mbining ge ne tre es . Ta x on 41 :3-10. B ininda-Emonds, O. R . 2004. Tre es versus ch ara cters and the su p ertre e/supermatrix "p ar adox ". S y stem atic B iolog y 53 :356-359.
Chapman, M. A., J . Cha n g , D. W eisman, R. V. Kesseli, and J . M. B u rke. 2007. Universal m arkers for com p arat iv e m appi ng and ph y lo g enet ic anal y si s i n t h e Ast era ceae (C om posi tae). Theoret ic al and Applied Genetics 115 :747-755. Johnson, K. A., B . R. H o lland, M. M. Hesle w oo d, and D. M. Cra y n. 20 12. Supermatrices, supert rees and ser endi pi tous scaf fol d in g : In fe rri n g a w el l-resol v ed, g enus -l evel ph y lo g en y o f S ty p h el io id eae ( E ri ca ce ae) despi te m issi ng dat a. M o lecul ar P h y lo g ene ti cs and Evol ut ion 62 :146-158. Konowalik, K. 2014. Reconstructin g reticul ate relationships in the poly p loid complex of Leucanthemum Mill.(Co m posita e, Anthe m ide ae ). Disse rta tion, Univ er sity o f Rege nsburg , German y , av ailable f rom h ttp://epub.uni-regensburg .d e/30913/. Konowalik, K., F . W agner, S. Tomasello, R. Vog t, and C. Ob erpriel er. 2015. D etectin g reticulate rel ationshi ps among diploid Leucanthemum Mill. (Composi
tae, Anthemideae) tax
a
using
multilocus species tree reconstru
ction me
thods and AFL
P fin g erp rinting . Molecular Ph y lo g enetics and Evolution 92 :308-328. Kovach, W
. 2001. MVSP-A multivariate stati
stical Packa g e for W indows, ver. 3.12f, Pentraeth, W ales, UK. Rag an, M. A. 1992. Ph y lo g enetic infer enc e based on matrix repr esentation of tr ees . Molecular Ph y lo g enetics and Evolution 1 :53-58.
Appendix S3
Fig. S1. Weight of Dichrorampha aeratana larvae reared on 15 Leucanthemum taxa varying
in ploidy level. Mean values ± SE are shown for each Leucanthemum taxon and horizontal lines indicate mean values for each ploidy level. vul= L. vulgare subsp. vulgare, vup = L. vulgare subsp. pujiulae, gal = L. gallaecicum, vir = L. virgatum, irc = L. ircutianum subsp. ircutianum, ica = L. ircutianum subsp. cantabricum, ile = L. ircutianum subsp. leucolepis, mop = L. monspeliense, adu = L. adustum, atr = L. atratum, cot = L. coronopifolium subsp. tenuifolium, fav = L. favargeri, het = L. heterophyllum, mon = L. montserratianum, max = L. maximum.
0 5 10 15
vul vup gal vir irc ica ile mop adu atr cot fav het mon max
Lar val w e ight (m g) 12x 10x 8x 6x 4x 2x Leucanthemum taxon
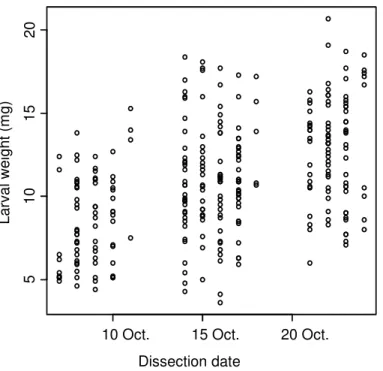
Fig. S2. Correlation between the weight of Dichrorampha aeratana larvae and the date when
its host plant was dissected. Larvae from plants dissected earlier were significantly lighter (t = 7.7, P < 0.0001).
10 Oct. 15 Oct. 20 Oct.
5 10 15 20 Dissection date Lar va l wei ght (m g)
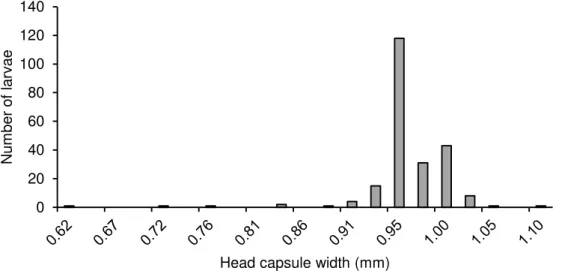
Fig. S3. Head capsule widths of larvae found when dissecting Leucanthemum plants of
different ploidy level in autumn. Larvae with a head capsule width of more than 0.90 mm were attributed to the final larval instar.
0 20 40 60 80 100 120 140 Num ber of lar v ae
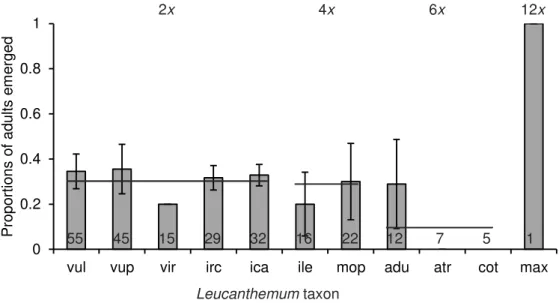
Fig. S4. Mean ± SE proportions of Dichrorampha aeratana adults that emerged per plant in
spring 2014 from larvae transferred in autumn 2013 on 11 Leucanthemum taxa varying in ploidy level. Numbers in bars indicate the number of larvae that had been transferred on each taxon. Each plant had been infested with a maximum of five larvae. Due to a lack of plants available for larval transfer in autumn 2013, no larvae were transferred on L. pallens, L. gallaecicum, L. favargeri, L. heterophyllum and L. montserratianum. vul = L. vulgare subsp. vulgare, vup = L. vulgare subsp. pujiulae, vir = L. virgatum, irc = L. ircutianum subsp. ircutianum, ica = L. ircutianum subsp. cantabricum, ile = L. ircutianum subsp. leucolepis, mop = L. monspeliense, adu = L. adustum, atr = L. atratum, cot = L. coronopifolium subsp. tenuifolium, max = L. maximum.
0 0.2 0.4 0.6 0.8 1
vul vup vir irc ica ile mop adu atr cot max
Pr opor tions of adults em er ged 7 55 45 15 29 32 16 22 12 5 1 12x 6x 4x 2x Leucanthemum taxon