Biochem. J. (1987) 245, 911-913 (Printedin GreatBritain)
The
crystal
structure
of the /-lactamase
of
Streptomyces
albus
G
at
0.3
nm
resolution
OttoDIDEBERG,* PauletteCHARLIER,* Jean-PierreWERY,* Philippe DEHOTTAY,t Jean DUSART,t ThomasERPICUM,t Jean-Marie FREREt and Jean-MarieGHUYSENt
*LaboratoiredeCristalographie, InstitutdePhysique, B5,andtServicedeMicrobiologie,Institut deChimie, B6, UniversityofLiege, B-4000SartTilman(Liege 1), Belgium
Thecrystal structure of the
fl-lactamase
of Streptomycesalbus Ghas been solved at 0.3 nm resolution by X-ray-diffraction methods. Theenzymeisatypicaltwo-domainprotein.One domain consists of fivea-helices, andthe other isfive-stranded f-sheet witha-helicesonboth sidesof the sheet. The active-siteserineresidue (Ser-48) is within acleft located between thetwo domains.INTRODUCTION
Three classes, A, B and C, of/1-lactamases
(fl-lactam
hydrolases,EC3.5.2.6)have beenidentified onthe basis of their substrate specificity, amino acid sequence and
mechanistic properties. The
fl-lactamases
of class B possess azinc cofactor; those of class A and C operateby an acyl-enzyme mechanism involving an active-site
serine residue. Several
fl-lactamases
of class A and C have been studied by X-ray-crystallographic methods (Knox et al., 1976; Aschaffenburg et al., 1978; Charlier et al., 1983; Dideberg et al., 1985; Moult et al., 1985; Kellyetal., 1986;Samraouietal., 1986). Asreportedby Kelly et al. (1986) and Samraoui etal. (1986), the class Afl-lactamases
of Bacilluslicheniformis
and B. cereus on the one hand and theDD-peptidaseofStreptomycesR61 onthe otheraresimilar intheextentand distribution of secondary structures (though lacking relatedness in primary structure). In no case, however, have these studies disclosed thecompletetracingofthepolypeptidechain.Recently,the geneencodingtheclass
A,f-lactamase
of StreptomycesalbusG hasbeencloned and sequenced,thusgivingaccesstotheprimarystructure of the enzyme (Dehottay et al., 1987), and the active-site serine residue has beenidentified at position48 (i.e. position 89 inthe precursor) after
,l-iodopenicillanate
derivatization of theprotein(De Meester et al., 1987).Inturn, we here present
thethree-dimensionalstructureof thisStreptomyces albus
G
,J-lactamase,
solved at0.3nm resolution.MATERIALS ANDMETHODS
The enzyme (purified to protein homogeneity as
describedbyDuez etal., 1981)wascrystallizedusingthe
vapour-diffusion technique (McPherson, 1976). The
protein[21 mg ml-'in 25mM-sodiumphosphate buffer, pH 7.0, containing 8% (v/v) glycerol and 8% ethylene
glycol] was dialysed against 50mM-Tris/HCl, pH 7.0,
containing 10mM-NaN3. After centrifugation, 10 d
drops ofthe protein solutionwere
(i)
mixed with3,1
ofa
10%
solution of10000-Mrpoly(ethylene
glycol)
madein the above buffer and (ii)
suspended
over 1 ml wells containing thesamepoly(ethylene
glycol)
solution. After1 week ofequilibration at 12°C, well-formed crystals
were obtained whichgrew up to 1.2mm in
length
and0.5mminwidth.When
exposed
tonitrocefin(0.1
mM in the mother liquor), thecrystal
and thesurrounding
mixture became red after 3 min at 18 °C, as aresult of
thecatalysedhydrolysisof the
fl-lactam
amidebond.The space group wasp212121
and the unit cell paramdters were a=4.067 nm, b= 4.353 nm and c= 13.843 nm.Assuming one 28500-Mr protein molecule per asym-metricunit,a
specific
valueof 0.00216 nm3(2.16A3)/Da
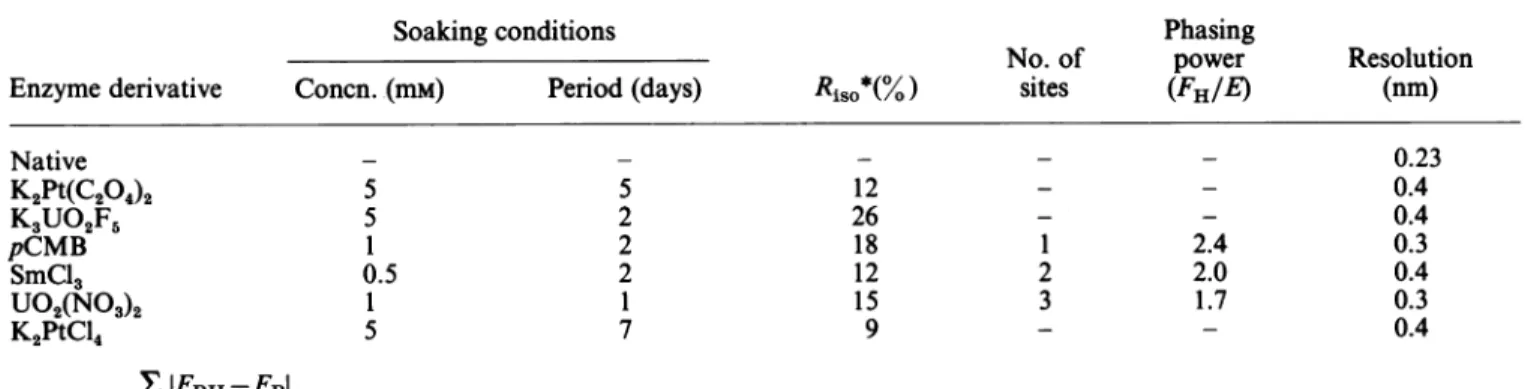
Table1. Data-collectionandphasingparameters Abbreviation:pCMB,p-chloromercuribenzoate.
Soaking conditions Phasing
No. of power Resolution
Enzymederivative Concn. (mM) Period (days) RisO*(%) sites (FH/E) (nm)
Native K2Pt(C204)2 K3UO2F5 pCMB SmCl3 U02(NO3)2 K2PtCl4 XIFPH-FPI *Ri = hkl
klFpl
hkl 5 5 1 0.5 1 5 5 2 2 2 1 7 12 26 18 12 15 9 1 2 3 2.4 2.0 1.7 0.23 0.4 0.4 0.3 0.4 0.3 0.4 Vol. 245 9110. Dideberg and others wascalculated.This value was within the range found for
proteincrystals (Matthews, 1968).Three native datasets werecollectedto aresolution of 0.23 nm.
Six heavy-atom derivatives
[K2Pt(C204)2,
K3UO2F5, SmCl3, U02(NO3)2, K2PtCl4 andp-chloromercuri-benzoate] of the enzyme were studied by collecting the intensity data on a Stoe-Siemens diffractometer with
graphite monochromated CuKa radiation. Two
equi-valentreflections weremeasured with the nativeenzyme and, for the derivatives, the Bijvoet mate of each
reflectionwascollectedat -20. Decaywasmonitoredby using two standard reflectionsfor every 100 reflections. Lorentz,polarization and absorption (Northetal., 1968) corrections were applied. The main data-collection and
phasingparameters are shown in Table 1.
The difference Patterson functions with coefficients
(IFPHI-IFPI)2
or(IFI-_IFi1)2
for thep-chloromercuribenzo-atederivative wereeasily interpretable. The heavy-atom positions and occupancies were refined by using the lowest estimate of theheavy-atom structure
amplitude,
FHLE.Theprotein phases thus derivedwerethen usedto
calculate a cross-phaseddifference Fourier map and to locate the heavy-atom-binding sites of the other derivatives. Each solution was compared with the two Patterson functions. Only two derivatives [SmCl3 and
U02(NO3)2]
gave aconsistentsetofheavy-atom
sites(so that three derivatives were used for thephasing
procedure).
Positional, thermal and occupancy parameters were varied in thephase refinement.Minorsiteswerefoundby inspecting the difference Fourier map. The overall figure-of-meritwas0.79for4462reflections.Amini-map (scale 0.4nm/cm) was calculated on a
grid
of32x32x100using the phase obtainedandtheamplitude weighted by theindividual
figure-of-merit.
Protein and solventregions appeared wellin themap. ApproximateC.,
positions weredirectly read off themapusinga grid chart, and theC.
positions served as guide points toconstruct the polypeptide backbone. The polyalanine modelwasfittedtothe 0.3 nm electron-density map with a PS300 Evans andSutherland colour graphics display system usingthe program FRODO (Jones, 1985).
RESULTS ANDDISCUSSION
The residuenumbering used below refers to the mature enzyme (whose N-terminus is Gly-40 of the precursor
protein). Except for the residues-77-91 amino-acid stretch, which lacked continuous density, thetracing of the chainas derivedfrom theelectron-density map was
unambiguous (Fig. 1). The two-domain protein has an
overall dimension of 5.5 nmx4.3 nm x 3.8nm. One of
thedomainshasacentral structural coreconsisting ofa
five-stranded
f-sheet
withthree a-heliceson oneface andonea-helixonthe other. Thestrands S3,S4 and
S.
forma
fl-meander,
averystablestructurethat is also foundin the active-site serine proteinases of the trypsin family (Schulz & Schirmer, 1979). The strandsS,
and S2 havea hairpin connection, and the
fl-sheet
has the usual left-handedtwist witharather smallQangle. The second domain consists of five a-helices. Stabilizing salt bridgesoccurthroughout thestructure.Inparticular, Glu-16of helixH1 interacts with Arg-39of strand
S2*
The active site (*, Fig. 1), identified as the cleft
possessing theserine residue at position 48 and known to be labelled by
fl-iodopenicillanate
derivatization of theprotein(seetheIntroduction),istopologically defined by the N-terminal portion of helixH3on one side, strand S3 onthe other side,helix H2 at the back, a loop connecting helicesH7andH8onthe top and aloop connecting helices
H5 and H6 atthe bottom (Fig. 2).The active-site Ser-48 is at the N-terminus of helix H2, a position that may
facilitatecatalysis (Hol, 1985). Other residueswhich, in concertwith Ser-48,mayplayimportantroles in catalysis
and/or substrate binding are Lys-51 (on H2), Glu-150
andAsn-1 54(onaloop between
H.
andH,),and Lys-218N
Fig.1. Schematicrepresentation of the three-dimensionalstructureof theStreptomycesalbusG0-lactamase
,f-Strands
(Si)arerepresented byarrows,andhelices(Hi)bycylinders. Nis theN-terminus; Cis the C-terminus.1987 912
Crystal structure of Streptomyces,6-lactamase 913
XA218 A218
CA5i CA5
C A4 A2 20 CA4 CA220
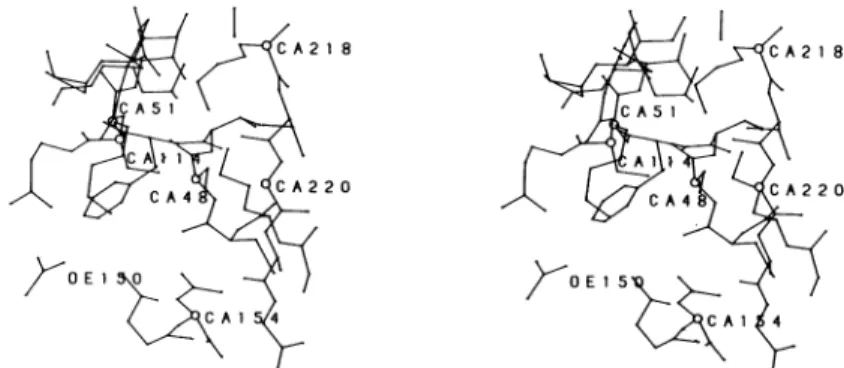
Fig. 2. Astereoview showing potentially important residues in the active site of theStreptomycesalbus GfJ-lactamase
Theorientation of the molecule is similar to that Fig. 1. The labelled residues are: Ser-48, Lys-51, Ser-l14,Glu-150,Asn-154,
Lys-218 andGly-220. 0,C<t(CA) atom.
and Gly-220 (on S3). Lys-51, Lys-218 and Gly-220 are conserved in the /-lactamases of class A and the DD-peptidases/penicillin-binding proteins of known primary structure.
The arrangement of secondary-structure elements in theStreptomyces albus G ,B-lactamase is comparable with those found in B. licheniformis and B. cereus Ienzymes (Kelly et al., 1986; Samraoui et al., 1986), except that one helix is missing in the two latter structures (H8 in B. licheniformisenzymeand H5in the B. cereusenzyme).
The work was supported bythe Fondsde Recherchede la Facultede Medecine, the Fonds de laRecherche Scientifique Medicale, Brussels(contractsno.3.4522.86 and3.4507.83),the Gouvernementbelge (actionconcerteeno. 86/91-90) and the Region wallonne (C2/C16/CONV.246/20428). J.D. is Cher-cheur qualifie of the Fonds National de la Recherche Scientifique(FNRS, Brussels).
REFERENCES
Aschaffenburg, R., Phillips, D. C., Sutton, B. J., Baldwin, G.,
Kiener, P.A. & Waley, S. G. (1978) J. Mol. Biol. 120,
447-449
Charlier,P.,Dideberg, O., Frere,J. M.,Moews, P. C. & Knox, J. R. (1983)J. Mol.Biol. 171, 237-238
Dehottay, P., Dusart, J., De Meester, F., Joris, B., Van Beeumen, J., Erpicum, T., Frere, J. M. & Ghuysen, J. M.
(1987)Eur. J.Biochem.,inthepress
De Meester, F., Joris, B., Lenzini, M.V., Dehottay, P., Erpicum, T., Dusart, J.,Klein,J.M.,Ghuysen,J.M.,Frere, J. M. & Van Beeumen,J.(1987) Biochem.J. 244,427-432 Dideberg, O., Libert, M., Frere, J. M., Charlier, P., Zhao, H.
&Knox, J. R.(1985)J.Mol. Biol. 181, 145-146
Duez, C., Frere, J. M., Klein, D., Noel, M.,Ghuysen, J. M.,
Delcambe, L.&Dierickx,L. (1981)Biochem.J. 193, 75-82
Hol,W.G. J. (1985) Prog. Biophys. Mol.Biol. 45, 149-195 Jones,T.A.(1985) MethodsEnzymol. 115, 157-171
Kelly,J. A.,Dideberg, O., Charlier,P.,Wiry,J.P.,Libert,M.,
Moews, P.C. Knox, J.R., Frere, J. M. & Ghuysen, J. M.
(1986) Science 231, 1429-1431
Knox, J. R.,Kelly,J.A.,Moews, P.C.&Murthy,N.S. (1976) J.Mol. Biol. 104, 865-875
Matthews, B. W.(1968)J.Mol. Biol. 33, 491-497
McPherson, A.(1976) Methods Biochem. Anal. 23, 249-345 Moult, J., Sawyer, L., Hertzberg, O., Jones, C.L., Coulson,
A. F. W., Green, D.W., Harding, M.M. &Ambler, R. P.
(1985) Biochem. J. 225, 167-176
North, A. C. T.,Phillips, D.C. &Matthews,F. S. (1968) Acta
Crystallogr. A24, 351-359
Samraoui, B., Sutton, B.J., Todd, R.J., Artymiuk, P.J.,
Waley, S. G. &PhillipsD.C. (1986)Nature (London) 320, 378-380
Schulz, G. E. &Schirmer, R. M. (1979) Principles ofProtein
Structure (Cantor, C.R., ed.), p. 83, Springer-Verlag, New
York
Received3 May 1987/15 May 1987;accepted27May1987